Registry of Standard Biological Models/Basic Component Models/RBS mRNA
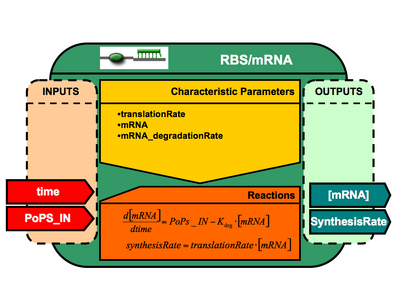
Ribosome Binding Site + mRNA architecture
CellML structure (CellML 1.1 spec)
- Component: RBS_mRNA
- Units:
- Imported from Environment component
- Variables:
- time (public interface = in )
- translationRate (public interface = none / init value = XXX)
- K_deg (mRNADegradationRate) (public interface = none / init value = XXX)
- mRNA (public interface = none / init value = XXX)
- synthesisRate (public interface = out )
- MathML
- <amsmath>\frac{d[mRNA]}{d time} = PoPsIN - K_{deg}*[mRNA]</amsmath>
- <amsmath> synthesisRate = translationRate*[mRNA] </amsmath>
CellML File
<syntax type = 'xml'> <?xml version="1.0"?>
<model xmlns="http://www.cellml.org/cellml/1.0#"
xmlns:cmeta="http://www.cellml.org/metadata/1.0#" xml:base="file:///C:/CellML_models/RBS+mRNA.cml" cmeta:id="RBS+mRNA" name="RBS+mRNA">
<component name="RBS+mRNA">
<variable name="mRNAsynthesisRate" initial_value="" public_interface="in" units="moles_per_second"/> <variable name="nb_mRNA" initial_value="0" public_interface="none" units="mole"/> <variable name="mRNAdegradationRate" initial_value="1" public_interface="none" units="per_second"/> <variable name="proteinSynthesisRatePermRNA" initial_value="1" public_interface="none" units="per_second"/> <variable name="proteinSynthesisRate" initial_value="" public_interface="out" units="moles_per_second"/> <variable name="time" initial_value="" public_interface="in" units="second"/>
[math]\displaystyle{
<apply id="proteinSynthesisRate">
<eq/>
<ci>proteinSynthesisRate</ci>
<apply>
<times/>
<ci>proteinSynthesisRatePermRNA</ci>
<ci>nb_mRNA</ci>
</apply>
</apply>
<apply id="rate of change of mRNA">
<eq/>
<apply>
<diff/>
<bvar>
<ci>time</ci>
</bvar>
<ci>nb_mRNA</ci>
</apply>
<apply>
<minus/>
<ci>mRNAsynthesisRate</ci>
<apply>
<times/>
<ci>mRNAdegradationRate</ci>
<ci>nb_mRNA</ci>
</apply>
</apply>
</apply>
}[/math]
</component>
<import xmlns:xlink="http://www.w3.org/1999/xlink"
xlink:href="file://C:\CellML_models\units.cml">
<units name="moles_per_second" units_ref="moles_per_second"/>
<units name="per_second" units_ref="per_second"/>
</import>
</model>
</syntax>
Comments
- trying to unified the interface between the quasi steady state approximation on the mRNA level and its fully dynamic simulation
- I try to encapsulate the mRNA dynamic inside the RBS component so it can deliver a 'synthesis-rate' flow to the downstream protein element. This way it would be the same interface a the Promoter_RBS_Coupled element.
- Don't know if it is possible to define a system of ODEs in the MathML description part of the component ???