Protein Quantification Using ImageJ
Determining the concentration of protein in SDS-PAGE gel bands using ImageJ
- To determine protein concentration you will need to have a standard curve to compare your samples to
- - For 5GB1, BSA works great as a protein standard, and a range of 0.025 μg/μL to 5.0 μg/μL works well as a range for the standard curve
- After running and destaining the gel, take a picture and save it as a .tif and as a .jpg (in case the tif file can’t be opened—an issue I am experiencing at the other lab).
- - Make sure you save your gel images as the same type of image (either .jpg or .tif) each time!
1. Download the ImageJ software: http://rsbweb.nih.gov/ij/download.html
2. Open ImageJ
3. Go to File→Open→(your image)
- Does your image look too dark or too light?
- - Image→Adjust→Brightness/contrast
- - I recommend saving the image with an updated name at this point so that you have it to go back to
- If image looks good as is...
4. Find the lane with the lowest concentration of BSA
5. Select the rectangle tool, and draw a box around the lane, making sure to include some of the empty gel between lanes and white space outside of the band
6. Go to Analyze→Gels→Select first lane
7. Make sure your cursor shows as an arrow, grab the rectangle you just made, and drag it to the next lane
- - DO NOT DRAW A NEW RECTANGLE! You must drag the same rectangle you just made
- - The point here is to compare the band in each subsequent lane using the exact same size/white space/noise as the originally defined area in Lane 1
8. Go to Analyze→Gels→Select next lane
9. Repeat steps 7-8 until all lanes have been selected and numbered
- - I think at this stage it's easiest to use key command "Ctrl+2" to continue numbering the subsequent lanes (less of a chance to mess up!)

- - I do not know how to correct the inevitable mistake boxes that you are going to make by accident and that cannot be undone...I'm sorry! Deleting them will cause useless white space where the rectangle was previously. My best advice if you find yourself in that predicament is to close the file, reopen it and start again. You will get really good at marking the lanes, I promise. If you are some kind of wizard who knows how to fix these mistake boxes without having to start over again, you should let me know!
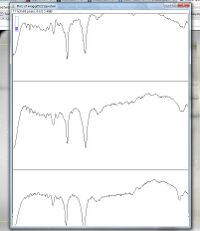
10. Once all lanes are defined, go to Analyze→Gels→Plot lanes (or use "Ctrl+3") to generate histograms of each lane
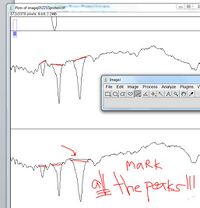
- - The peaks (or valleys if your image is inverted like the one in this example) in each grid correspond to the intensity of the bands in the lane
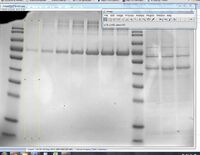
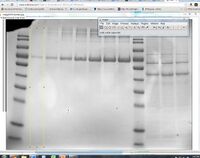
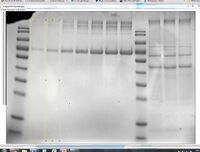
- The standards lanes should only show one peak, while the lanes with protein you want to quantify will probably show multiple bands, as in the image below
- - Now is a good time to save! If the plot window is active, when you go to File→Save through the ImageJ interface, it will be trying to save your histograms; if the image window is active, the image will saved. Save both, smartypants!
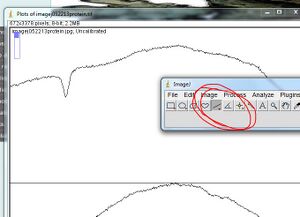
11. On the ImageJ interface, select the "line" button
- - Draw a line at the bottom of the peak that represents the first standard to define the area of the curve
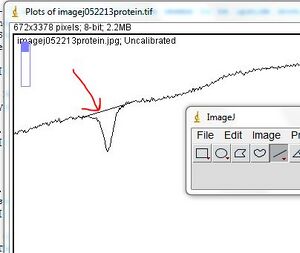
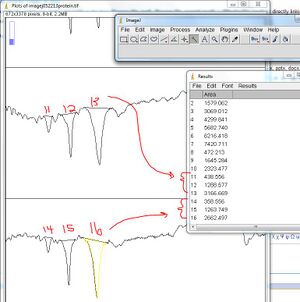
- - Keep drawing all the single lines to define the curves in your standard lanes, and draw multiple lines in your lanes with your protein of interest. In this example, we know that the protein consists of 3 subunits, and they represent the majority of the protein sample. Therefore, three curves have been established.
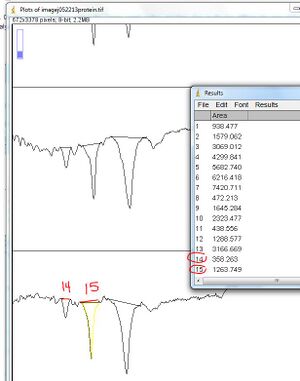
12. On the ImageJ interface, select the "magic wand" button and then click on the line defining the area of the curve of the first standard, and the areas of the curves in your protein analysis lanes
- - Continue selecting the area outlines of the remaining lanes
- The measurement of the areas will be bumped to a "Results" window
- Be aware that ImageJ isn't "smart" -- The order in which you click corresponds to the numbers in the Results window; numbering does NOT correspond to the lanes you defined!
- If you click on the wrong part of the histogram, you can find the bad measurement in the Results window, highlight it, go to Edit→cut and remove the bad entry. You should fix this as soon as you click on the wrong part of the histogram to keep your numbering correct.
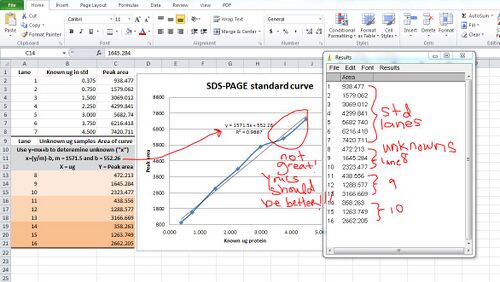
13. Save your Results window so that you can transfer the measurements to excel to generate a standard curve
- Use the trendline formula to solve for your unknown proteins
- Your standards should be linear...you can go back to the histogram and try to draw better lines to define the standard peaks and that might clean up ugly data a bit (but in this case it would require new screenshots, and ain't nobody got time for that)