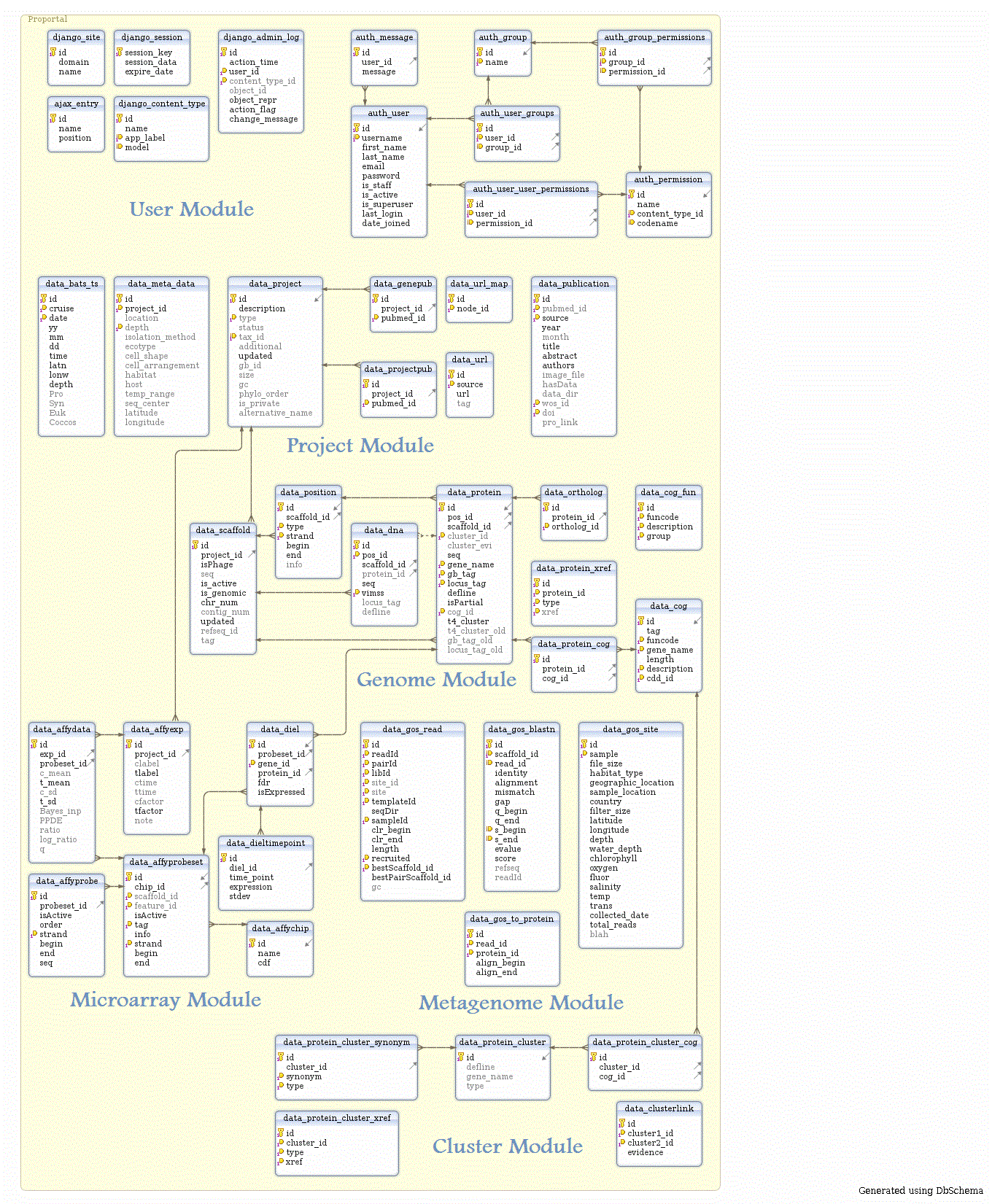
Proportal DBSchema
Proportal DB Schema
User Module
Project Module
Table: data_project
A list of projects
* 72 projects, as of 07-21-2011 (To be updated: 58 in PRO DB while 72 in DEV DB) * Last updated: 2010-12-10 * No foreign key
Notes The field "type" is the genome type for this project. This information is used to assign the proteins into two types of either PhCOG or CyCOG in the ortholog clustering. It has the following distinct types, which may be moved into a separated table for a clear definition,
* cpm, Cyanophage genomes part 1 (To be updated: 18 records in PRO DB, 28 in DEV DB), also for 'PhCOG' cluster. * cpp, Cyanophage genomes part 2 (To be upadted: 8 records in PRO DB, 11 in DEV DB), also for 'PhCOG' cluster. * cps, Cyanophage genomes part 3 (2 records in both DBs), also for 'PhCOG' cluster. * ma, physiology experiments (4 records in both DBs): Light Sensing, Nitrogen Availability, Phage Infection, and Phosphate Starvation) * mt, expression experiment (1 record in both DBs): Microbial community gene expression in ocean surface waters * p, Prochlorococcus genomes(13 genomes in both DBs), also for 'CyCOG' cluster. * pb, Prochlorococcus Publications (1 record in both DBs) * s, Synechococcus genomes(11 genomes in both DBs), also for 'CyCOG' cluster.
The link for "tax_id" is defined in data_url table.
* type_id = 59919 * source = tax * url = http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?id=59919
Table: data_projectpub
A list of publications from various projects.
* 32 publications as of 07-21-2011
* Foreign keys:
o project_id = data_project.id
o pubmed_id = data_publication.id
Notes
This table is used for mapping projects with related publications, which is displayed on the strain page in the "Genomes" section, for instance: MED4.
Table: data_genepub
This table is empty. Consider to use data_publication table instead?
* Foreign key: data_project
Table: data_publication
A list of publications related to Prochlorococcus, Cyanophage, and Synechococcus genomes.
* 2528 publications listed as of 08-1-2011 (both DEV and PRO DBs are updated) * pubmed_id can be used as a foreign key. * "year": last updated 2010
The primary key in this table is referred by data_projectpub table as pubmed_id.
Table: data_url_map
This table is empty.
Table: data_url
The list of data links or data folders.
Meta Data Module
Table data_bats_ts
Information about field investigation.
* No foreign key
Table: data_meta_data
Information about field investigation for each project
* 66 meta data sets, as of 07-22-2011 * Foreign key: data_project.id, to be fixed.
Error
* One project_id 26 is missing in data_project table. * Six projects defined in data_project table (all in year 2008) do not have meta data defined in this table.
Genome Data Module
Table: data_scaffold
A list of strains/genomes used in various projects. Some strains defined in data_project table have multiple contigs for the same genome. For instance, strain Syn_BL107 (project id 31) has six contigs sequenced thus there are six records in data_scaffold table for the same project id 31.
* Last updated: 12-10-2010 * 213 strains, as of 07-22-2011 * Foreigh key: data_project.id
Questions
* "refseg_id" not defined * "seq" field can be removed because its content is further defined in data_dna and data_protein tables.
Table: data_position
List of start and end positions of gene/DNA for each strain defined in Table data_scaffold. Its primary key is used in data_protein table as the foreign key pos_id.
* 67516 pair of positions, as of 07-22-2011 * 9 types of sequences are defined: 16s, 23s, 5s, as, m, n, orf, ps, t * Foreign key: data_scaffold.id.
Table: data_dna
A list of DNA sequesnces in correspondence to sequence postion information defined in data_position table.
* 67516 pieces of DNA sequences stored, as of 07-22-2011 * Three foreign keys: data_position.id, data_scaffold.id and data_protein.id
Error
* Foreign key pos_id has error:
o Two position ids in data_position table: 37163 and 46814 are missing in this table
o Two pos_id: 36978 and 37113 do not exist in data_position table.
Table: data_protein
Unique proteins from various genomes, which are shown on protein pages, for instance, http://proportal.mit.edu/protein/70464/0/.
* 65909 proteins defined, as of 07-22-2011 (1607 DNA sequences are not present in this table)
* Two foreign keys:
o pos_id = data_pos.id, the foreign key to the table data_position, in which the start and end of positions if protein sequences are stored.
o scaffold_id = data_scaffold.id
* Relation to ortholog clusters
o cluster_id: id for the cluster into which this protein falls. This
o cluster_evi: evidence for the cluster (ortholog, hmmscan, singleton, uclust)
Notes
1. "cluster_id" and other ortholog cluster related fields should be removed from this table and defined in a separated table.
2. "cog_id" should be removed from this table, which is redundant with "cog_id" defined in data_protein_cog table, i.e., data_protein.cog_id equals data_protein_cog.cog_id when data_protein.id=data_protein_cog.protein_id.
Table: data_protein_xref
Definition: ?
* 36774 records stored, as of 07-22-2011 * Foreign key: data_protein.id, to be fixed,
Error
* Two records have missing protein_id: 36950 and 45482 in data_protein table
Affychip Expression Module
Table: data_affychip
Information about each affychip used.
* 1 chip defined, as of 07-22-2011 (same for DEV and PRO DBs) * No foreign key
Table: data_affyexp
A list of affychip expression experiments displayed on Proportal expression page, for instance, http://proportal.mit.edu/expression/17/,
* 20 affychip experiments, as of 07-22-2011 (same for DEV and PRO DBs) * Foreign key: project_id, only three projects involved affychip experiments.
Table: data_affyprobeset
A list of probe sets for various affychip experiments.
* 9966 records, as of 07-22-2011
* Three foreign keys:
o chip_id:
o scaffold_id: has missing keys
o feature_id: not defined
Notes
* feature_id not defined, but is actually in one to one correspondence to gene_id in data_diel table, which is further linked to data_protein table using protein_id. * Use "begin" and "end" to match DNA\gene\protein?
Table: data_affyprobe
A list of probes for various affychip experiments.
* 89749 records, as of 07-22-2011 * Foreign key: probeset_id
Table: data_affydata
The expression results of Affychip experiments.
* 110848 records, as of 07-22-2011
* Foreign keys,
o exp_id
o probeset_id
Notes
* No DNA\gene\protein info, use probeset_id?
Table: data_diel
The mapping between gene id and protein id, which is used in Affychip expression experiments.
* 1695 records, as of 07-22-2011
* Foreign keys,
o probeset_id: foreign key to data_affyprobeset table.
o protein_id: foreign key to data_protein table.
o gene_id: not defined, foreign key to protein table.
Notes
* gene_id not defined, but is actually in one to one correspondence to feature_id in data_probeset
Table: data_dieltimepoint
Time courses of Affychip experiemnts.
* 42375 records, as of 07-22-2011 * Foreign key: diel_id
Cog Module
Table: data_cog_fun
A list of Cog gene functions.
* 24 funtion categoriess, as of 07-22-2011 * No foreign key
Table: data_cog
A list of Cog genome annotations
* 4874 records, as of 07-22-2011 * Foreign key: data_cog_fun.funcode ?
Notes
* data_cog_fun.funcode can't be regarded as a foreign key becuase some of funcodes in this table are missing in data_cog_fun table.
Table: data_protein_cog
The mapping between Cog genome and proteins.
* 18498 records, as of 07-22-2011 * Foreign keys: data_protein.id and data_cog.id
Notes
The mapping between "protein_id" and "cog_id" has already been defined in data_protein table. This table is actually part of data_protein table.
Microarray Module
Table: data_gos_site
A list of Gos field experiments, such as sites of experiments etc.
* 78 records, as of 07-22-2011 * No foreign key
Table: data_gos_read
A list of field reads for various Gos experiments.
* 9893120 records, as of 07-22-2011 * Foreign key: data_gos_site.id, no error
Table: data_gos_to_protein
The mapping between Gos genomes and proteins.
* 926072 records, as of 07-22-2011
* Foreign keys:
o data_protein.id, has error, to be fixed
o data_gos_read.id, has error, to be fixed
Error
* The foreign key: read_id=0 is not defined in data_gos_read table for id=1 and id=705172 in this table * The foreign key: protein_id=0 is not defined in data_protein table for id=1 and id=705172 in this table
Table: data_gos_blastn
A list of sequences from Gos experiments.
* 8666847 records, as of 07-22-2011
* Foreign keys:
o data_scaffold.id, has error, to be fixed
o data_gos_read.id, has error, to be fixed
Error
* The foreign key: scaffold_id=0 is not defined in data_gos_read table for 211 records in this table * The foreign key: read_id=0 is not defined in data_gos_read table for 56438 records in this table
Cluster Module
Table: data_ortholog
This table is used to store protein orthologous pairs that are identified from the cluster pipeline, in which all protein sequences have been blasted all-to-all for the identification of orthologs.
* 830944 orthology pairs defined, as of 07-22-2011
* Foreign keys:
o protein_id: first protein id in the orthologous pair
o ortholog_id: second protein id in the orthologous pair
Notes
This table gets updated after assignOrthologs.py parses the BLASTP m9 results and assigns orthologs.
Table: data_protein_cluster
A list of protein clusters.
* 5597 records in DEV DB and 16013 in PRO DB, as of 07-22-2011 * No foreign key
Notes
* Two distinct "type":
o PyCOG: for all Cyanophages
o CyGOC: for all Sy and Pro-chlorococcus (Not sure why this naming convention)
* "gene_name" not in use
This table is supposed to store the information about COG clusters in the format of cluster id, gene name, gene description and cluster type. However, the current implementation of this table has the following problems: (a) most of gene_names have null values because no unique gene id or gene name is defined/used in the DB; (b) the gene description is incomplete and outdated.
Actually, the data stored in this table can be generated dynamically from data_protein table. The final COG cluster information can be retrieved from the join of data_protein_cluster and data_protein tables as follows,
select * from data_protein_cluster c left join data_protein p on c.id = p.cluster_id;
Obviously, gene_name and define in data_protein_cluster table are redundant with data_protein table. Both fields should be part of data_protein table but not of data_protein_cluster.
Another issue is: data_protein table should be maintained for defining a list of unique proteins from various genomes. The cluster information should be separated from data_protein table and defined elsewhere in a downstream table in the hierarchy. In addition, different versions of COG clusters should also be defined and stored. To be compatible with previous versions of cluster IDs used in the published papers, a mapping table is necessary to store individual cluster IDs for different versions of clusters, as well as their mapping to the master cluster IDs, which are conserved and compatible.
Therefore, the ultimate solution to the COG clustering pipeline is to add the data_protein_cluster_mapping table in such a format,
* id, the primary key * protein id, the foreign key to data_protein table * master cluster id, this is the conserved and compatible unique IDs of COG clusters * cluster type, either CyCOG or PhCOG * version #, version number * version-ed id, the cluster ID in the version.
The master cluster IDs are similar to unique COG IDs. The individual protein constitutes in a COG cluster may change in different versions. However, the majority should remain the same for an unique master cluster ID.
Table: data_protein_cluster_xref
This table is used to stored the information for the mapping between a cluster id and a COG id. Such information is displayed on the protein page, for instance, http://proportal.mit.edu/genome/id=1/proteins/p1/.
* 1100 records, as of 07-22-2011 * Foreign key: data_protein_cluster.id, has error, to be fixed
Notes
* Only one "type": c * "xref": COG external reference id, which may correspond to multiple cluster ids
Error
Some of the foreign key: data_protein_cluster.id are not defined in data_protein_cluster table for about 880 records. The problem was caused by different versions of ortholog clustering. Contains cluster ids in the range 18, 2580. Not sure what the refs are x-referencing. These cluster ids need to be off-limits during reassignment.
Table: data_clusterlink
A list of pairs of clusters.
* 71 records in DEV DB and 219 records in PRO DB, as of 07-22-2011 * Foreign key: data_protein_cluster.id,has error, to be fixed
Notes
* "evidence" is not in use
Error
* The foreign key: cluster_id=0 is not defined in data_protein_cluster table.
Table: data_protein_cluster_synonym
The table is empty.
Table: data_protein_cluster_cog
This table is empty.