Project Page
Team Pink's Final Project
| General Overview | |
| Our project is based on the general design of an AND logic gate using an alternative splicing RNA aptamer construct as the functional mechanism. Our project overview is to apply this two-step AND gate to a different gene of interest than the original researchers but to still retain the ability to preferentially target cells with a specific activated pathway and to change that cell’s phenotype non-invasively with that sensing construct | |
| Background Info | |
This construct scheme is designed off the work of Culler et al. from Stanford University as highlighted in her November 2010 Science article . This AND gate takes in a user provided small molecule and an endogenous protein as inputs. The endogenous protein of interest interacts with the RNA construct and activates the production of a downstream enzyme. This enzyme acts on (or in conjunction with) the user provided compound. Together those two produce the desired output phenotype. (Reprogramming Cellular Behavior with RNA Controllers Responsive to Endogenous Proteins. Stephanie J. Culler, Kevin G. Hoff, and Christina D. Smolke. Science 26 November 2010: 330 (6008), 1251-1255. )
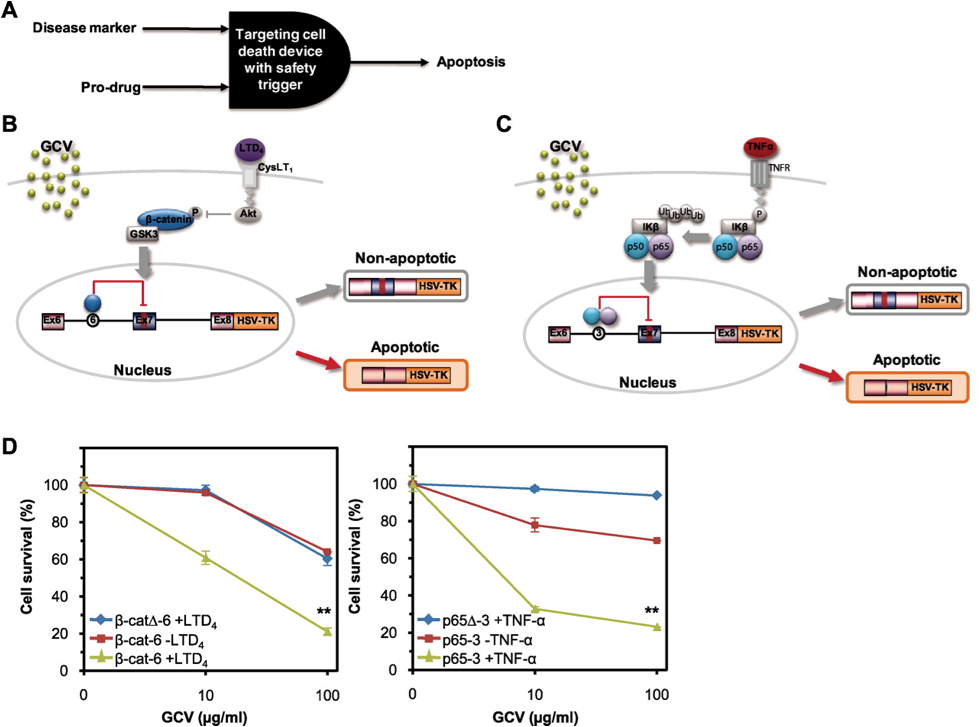 | |
| Figure 1 Preferential apoptosis of cells with LTD4 and TNF-α activated pathways. The RNA construct is localized in the nucleus and its activity is controlled by alternative splicing. The Ex7 sequence contains a stop codon and if included in the post-processed RNA greatly diminishes the product of the downstream gene of interest (in this case HSV-TK). If Ex7 is repressed then the splicing modification removes that segment and the subsequent RNA product is highly transcribed. In this case HSV-TK, the gene of interest, increases the likelihood of apoptosis when GCV is also present. This way, in two distinct steps an AND gate is created where both GCV and the activated pathway are needed.]] | |
| Research Problem and Goals | |
| Our research problem will be to test this RNA construct in new environments to begin to assess its viability as a general cross-platform motif for pathway sensing and controlled phenotypic response. Three main areas of investigation present themselves. The first is the possibility the test for a different cell type (A different prokaryote, eukaryote, mammalian etc). Unlike other synthetic circuits we have seen that have been relatively independent of the cell type, this construct’s strength comes from directly sensing specific pathway activations that might be different across cell lines, kingdoms etc. The other two areas involve the two steps of the AND gate implementation. We could keep the same cell line but look at a different pathway to sense. Along the same lines we could change the gene of interest and its associated researcher added compound (GCV and HSV-TK in the first example) to modify the expected phenotypic output response. | |
| Sources and Summaries | |
| Escape from HER-family tyrosine kinase inhibitor therapy by the kinase-inactive HER3
Nature 445, 437-441 (25 January 2007) | doi:10.1038/nature05474; [1] Sergina et al article from 2006 that described the interrelation of the HER-family of tyrosine kinases and their ability to maintain breast cancer growth. Interesting because this has become one of the most successful applications of genetic screening of in clinical breast cancers because the over activation of HER2 leads to a much more aggressive cancer. This paper highlights another specific pathway that is over active in cancerous cells and again gives plenty of molecular pathway components that could be targeted with our construct for phenotypic change. | |
| Suppression of lung adenocarcinoma progression by Nkx2-1 Winslow et al. Koch institute
Nature (2011) doi:10.1038/nature09881 [2] Interesting article coming out of Tyler Jacks lab at the Koch Institute (Completely unrelated to the UROP Brett does at the Manalis Lab at the same place). This article goes into the in-depth discovery of the mechanism for lung adenocarcinoma metastatic progression and provides many high quality pathways that we could target with our RNA-based sensing construct. |