OpenSourceTB:OSTB Series 2
Current Activity
The data on this page are now (March 2016) out of date, as is the project status, but all the raw structures and potencies are shown here. Updates will trigger alerts to Twitter, or please email opensourcetb@gmail.com for more.
Starting Point
TCMDC143693 identified by GSK in a high throughput screen (to be published shortly).

Properties
Inherited data from GSK for TCMDC-143693: MIC vs. H37Rv = 10 μM, active against non-replicating TB (>50% inhibition) in 10 experiments out of 13) and with mean PXC50 of 5.2 μM. Toxicity vs. HepG2 was >100 μM. cLogP is 3.1 and (presumably) measured solubility is 72 μg/mL. Chrom LogD (pH 7.4) is 5.21, and molecular weight 385.
Known occurrences
TCMDC143693 was, when this page was created, absent from Google. Pubchem search leads to a patent.
ChEMBL flags up that a search on SureChEMBL reveals that the compound has appeared (along with nearest neighbours) on 2 patents of the same patent family: https://www.surechembl.org/document/US-20090176778-A1/ and https://www.surechembl.org/document/US-20120121540-A1/
Related Structures
Discovery of Q203, a potent clinical candidate for the treatment of tuberculosis
Lead Optimization of a Novel Series of Imidazo(1,2-a)pyridine Amides Leading to a Clinical Candidate (Q203) as a Multi- and Extensively-Drug-Resistant Anti-tuberculosis Agent
Identification of Novel Imidazo(1,2-a)pyridine Inhibitors Targeting M. tuberculosis QcrB
Bactericidal Activity of an Imidazo(1, 2-a)pyridine Using a Mouse M. tuberculosis Infection Model
Predicted Target
A prediction of the target was carried out by Marc Marti-Renom and ChEMBL as part of the original HTS paper identifying the compound.
The compound (TCMDC143693) has four predictions, three from Marc's approach and one from the ChEMBL’s approach. (The details on how the predictions were identified are in the forthcoming manuscript).
Marc's predictions:
1) O06266 (an epoxide hydrolase). Link: TCMDC143693 —> 3TH-S38-3ans_A —> S38-O06266 (i.e. the GSK compound is similar to 3TH in PDB, which was co-crystallized with 3ans (a Human soluble epoxide hydrolase in complex with a synthetic inhibitor). In turn, the model of the Tuberculosis epoxide hydrolase (O06266) was based on 3ans template and the binding site was conserved. Thus, the path links TCMDC143693 to O06266.
2) A2VJ47 (an epoxide hydrolase ephB), TCMDC143693 —> 3TH-S38-3ans_A —> S38-A2VJ47
3) P64411 (Heat shock protein 90, TCMDC143693 —> 3TH-PFT-3k99_A —> PFT-P64411
The EBI prediction:
P96222 (Mtb HTH-type transcriptional regulator Eth) now http://www.uniprot.org/uniprot/P9WMC1 and https://www.ebi.ac.uk/chembl/target/inspect/CHEMBL1772929. The compound shares a lot of structural features found in known actives (activity <=10 μM) for this target in ChEMBL. These features include the thiophene ring, -CF3, pyrrole ring and nitrile.
Current question: are there known inhibitors of any of these targets that show inhibition of Mtb? There is a paper identifying hits vs. this target, but the effect of inhibition is to increase the potency of another compound - the compounds are not themselves antitubercular. Thus if TCMDC143693 has potency on its own (which it does), perhaps there is another (different or additional) MoA.
Synthetic Chemistry
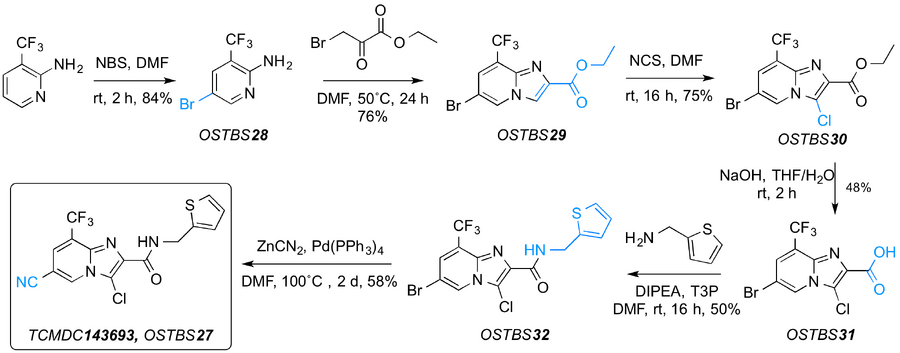
Initial efforts to explore the series were focused on the resynthesis of the hit compound TCMDC-143693. The synthesis of this compound has already been described, therefore the same sequence of reactions was performed. The synthesis started with bromination at the 5-position of the 2-amino-3-(trifluoromethyl) pyridine (JBG28-1) followed by cyclisation with ethyl bromopyruvate to generate the imidazopyridine ring (JBG43-1), then chlorination at the 3-position with N-chlorosuccinimide (JBG45-1). The resulting ester was hydrolysed with NaOH (JBG47-1) to generate the corresponding acid, which could easily be converted to the amide by treatment with 2-(aminomethyl)thiophene and a coupling agent such as T3P (JBG48-1). The last step, the palladium cyanylation reaction, was modified from the procedure described in the literature, with a longer reaction time and conventional heating instead of microwave irradiation, improving the yield of the final compound (JBG50-1), also known as OSTBS27.
Data
All OSTB molecules are listed in the OSTB Database. Chemdraw/png files will be on Github under Series 2. Wiki files are here.
The hit: TCMDC143693 ClC1=C(C(NCC2=CC=CS2)=O)N=C3C(C(F)(F)F)=CC(C#N)=CN31 InChI=1S/C15H8ClF3N4OS/c16-12-11(14(24)21-6-9-2-1-3-25-9)22-13-10(15(17,18)19)4-8(5-20)7-23(12)13/h1-4,7H,6H2,(H,21,24) LWIBMEYBVJUXDE-UHFFFAOYSA-N
Licence
Content is CC-BY-4.0.
Contact
To discuss this project or page, see the contact info on the OSTB Main Page
