Oneclickfeature
<html><head> <style type="text/css"> .main { margin: 0 auto; width: 804px;border-style:solid; border-width:5px; }
- logo{ margin:10px 0 0 52px; display:block; background:url('http://openwetware.org/images/7/74/Banner20.png') 0 0 no-repeat; width:700px; height:200px; text-indent:-9999px;}
.navbox { position: relative; float: left; }
- menu{ background:url('http://openwetware.org/images/3/3f/Menu_bg.png') 0 0 no-repeat; width:643px; height:100px; margin:51px 0 0 78px; padding-top:100px}
- menu li{ float:left;}
- menu a{ font-size:30px; color:#000000; line-height:1.2em; text-decoration:none; letter-spacing:-1px;}
.nav1{ padding:26px 0 0 37px;} .nav2{ padding:16px 0 0 30px;} .nav3{ padding:36px 0 0 17px;} .nav4{ padding:16px 0 0 19px;} .nav5{ padding:26px 0 0 30px;}
- menu .nav1 a:hover{ color:#bb0e0e}
- menu .nav2 a:hover{ color:#ca6509}
- menu .nav3 a:hover{ color:#3f9711}
- menu .nav4 a:hover{ color:#0ca0ce}
- menu .nav5 a:hover{ color:#8606c5}
</style> </head>
<header> <a href="index.html" id="logo">DNAmazing asfsaf. Smart. Effective</a> <nav>
</nav> </header>
</html>
"One-click" feature to get the results
The DNAmazing program successfully
- Perform the automatic generation of possible scaffold folding paths from users' input and crossover positions for 2D DNA Origami structures
- Generate possible sticky end sequences
- Generate staple sequences with sticky ends which are necessary to conduct wet lab experiments for 2D DNA Origami structures.
With the "one-click" feature, the processing of DNA Origami design can be accelerated as users no longer have to do the raster filling (the folding path) manually which is definitely tedious for large and complex structure.
Automatic generation of possible folding paths and crossover positions
The traditional algorithm of Hamiltonian circuit/circle was modified and adjusted for the finding of folding pathway in DNAmazing.
DNAmazing has been tested for various 2D structures with arbitrary shapes and holes. The positions of crossover are implicitly determined during the process of folding the scaffold strand and not shown here. However, the console version of DNAmazing can provide the information about the crossover positions.
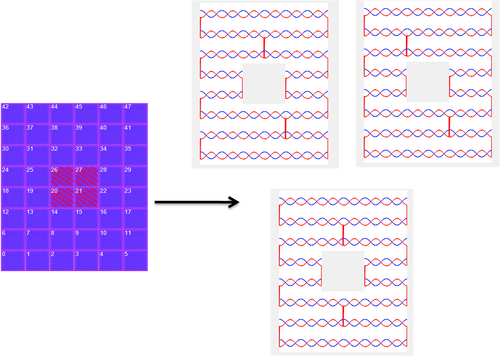
Below are some typical results of DNAmazing.These results can be easily reproduced by using the DNAmazing program in this report. The left images are the input from the users and the right images are the scaffold folding ways produced by DNAmazing. DNAmazing uses a filtering process to select the most suitable candidates. These results can be produced in very short time. In fact, if we compare DNAmazing to other CAD program which users have to design the scaffold way by themselves, DNAmazing is extremely fast to get the right results.
The generation of a 1 hole structure
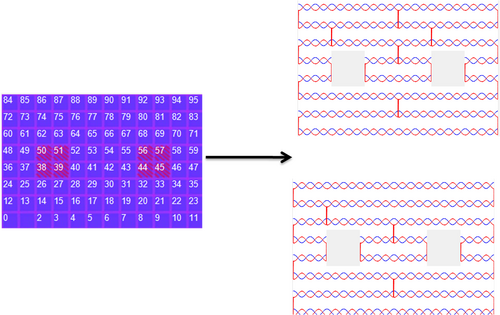
The generation of a 2 hole structure
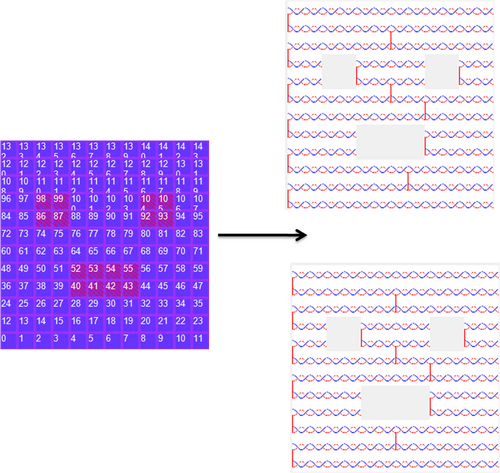
The generation of a "smiling face" structure
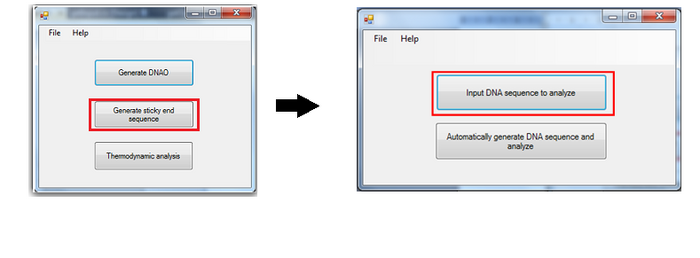
Generate sticky end sequence
To support generation of sticky end, as well as, to ensure that the sticky end will not affect the scaffold folding, an additional component is provided. User can choose to manually input a DNA sequence, and the program can help to check for the most stabilizing binding position in the scaffold. The binding energy is also calculated for users’ reference.
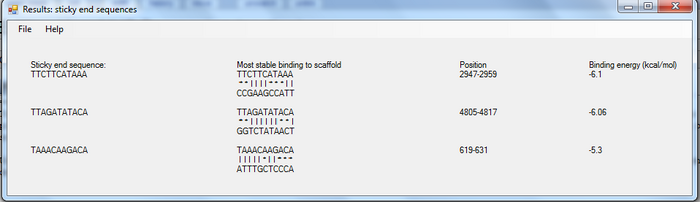
User can also ask the program to generate the sticky end sequence with the defined length. DNA sequences with binding energy higher than a limit defined are given. The below image illustrates the output of sticky ends' sequence generation.
Generation of staple sequences
As the folding paths, positions of crossover and sticky end sequences are determined in above steps, the sequence of staple strands are easily found by using the complementary base pairs rules:A-T, G-C. DNAmazing produced a list of staples sequences at the final step. These information can be used to create DNA Origami structures in wet lab experiments.
Because there are not strict rules about the merging process of DNA staples, each CAD programs may provide different staples sequences and only experiments can verify the results. However, due to the limitation of time and the cost of experiments, we did not have chance to test the result of DNAmazing.