McClean: designing primers
Overview
Primers are short segments of DNA which can be used for PCR. Designing them isn't tricky. Keep in mind that DNA is polymerized in the 5' -> 3' direction. Thus, for a given 5' -> 3' strand to be amplified, the "forward primer" should match the first segment of base pairs and the "reverse strand" should match the reverse-complement of the last segment of base pairs. For example, to amplify 5'- TGTGTACGTACGGT-3' with ridiculously short primers, you would use froward primer: 5'-TGTGT-3' and reverse primer: 5'-ACCGT-3'.
For colony PCR, we typically use primers 20bp long. To insert a plasmid in a genome, we use primers that are 60bp long. 40 correspond to genomic DNA and 20 correspond to plasmid DNA. To follow the notation that we use, the genomic DNA should be capitalized and the plasmid DNA should be lowercase.
Materials
- A map of the segment of DNA that you want the primers to surround.
Example with NCBI Primer Blast
Suppose I want to amplify "pyADH1 PROMOTER," the white gene in this screenshot from Plasmid Viewer.
- Step 1- I will copy that segment of DNA, and I will include DNA a bit beyond both ends (blue highlighted region). Including extra DNA on the ends gives more options for which primers to choose. I included 12bp before the start of it and 11 bases after the ends. 5-30 bases is a reasonable range for most situations.
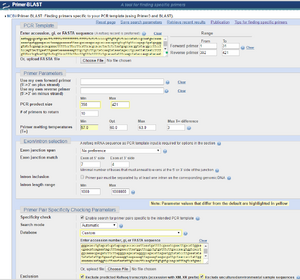
- Step 2- Paste the sequence in the PCR Template box here: NCBI Primer Blast
- Step 3- Fill in primer range. Since I want my PCR primer to be at least 19 bases long, I included 12 additional bases in the beginning, I included 11 additional bases at the end, and the entire segment is 421 bases long, I set the primer range to be 1-31 and 392-421.
- Step 4- Fill in Primer Parameters. I want my maximum product size to be 421. Since the "pyADH1 PROMOTER" is actually 398 bases long, that is my minimum product size. The other values can be left at their default values. However, it wouldn't be unreasonable to lower the minimum melting temperature to Tm=53 and reduce the maximum difference between the melting temperatures of the two primers to 2°C.
- Step 5- Fill in the "Primer Pair Specificity Checking Parameters." These parameters let it know which primers would bind elsewhere (off-target binding) so that those primers are not chosen. So it needs to know what other DNA will be in the mixture when the PCR is performed.
- Step 5 OPTION 1- If I were amplifying "pyADH1" from the yeast genome in a colony PCR, I would set the organism to "Saccharomyces cerevisiae (taxid:4932)". Then I would check the "Show results in new window" box, then I would click "GET PRIMERS."
- Step 5 OPTION 2- If I were only amplifying it from a plasmid, I would have even less off-target binding to be concerned about. I would then copy the entire plasmid sequence, convert it to a FASTA file on this website FASTA converter, I would then selected Database> Custom, then upload my FASTA file, then I would check the "Show results in new window" box, then I would click "GET PRIMERS."
- Step 6- Choose the best PCR templates from the new window. Verify that your sequence matches the recommendations on this page Designing_primers#General_guidelines
Notes
Please feel free to post comments, questions, or improvements to this protocol. Happy to have your input!
- List troubleshooting tips here.
- You can also link to FAQs/tips provided by other sources such as the manufacturer or other websites.
- Anecdotal observations that might be of use to others can also be posted here.
Please sign your name to your note by adding '''*~~~~''': to the beginning of your tip.
Contact
- Cameron J. Stewart 14:01, 20 June 2016 (EDT)