McClean:Yeast viability with methylene blue
From OpenWetWare
Jump to navigationJump to search
Overview
Assess viability of a culture of yeast by staining with methylene blue.
Methylene blue readily permeates yeast cells, but it is reduced to a colorless compound in living cells. Dead cells will appear dark blue when stained with methylene blue while live cells will be colorless.
Methylene blue also fluoresces, with peak excitation at 665 nm and peak emission at 690 nm.
Materials
- Mid-log culture of yeast
- 0.01% methylene blue
- Dissolve 0.01g methylene blue in 10 ml dH2O
- Add 2g sodium citrate dihydrate, stir until dissolved
- Bring to 100ml with dH2O
- Filter sterilize
- 95°C heat block
- PBS + 0.1% Tween (for flow cytometry)
Procedure
Staining cells
- Aliquot 2 x 1 ml mid-log culture into eppendorfs
- Heat kill one tube at 95°C, 15 min
- Combine culture and 0.01% methylene blue in a 50:50 mixture
- Asses viability using a hemacytometer (dead cells will be dark blue) or flow cytometry
If doing flow cytometry
- Dilute culture/methylene blue mixture 1:5 in PBS + 0.1% Tween
- Methylene blue excites at 665 nm and emits at 690 nm. On the UW flow core Attunes, this is the RL2 filter (633nm laser, 720/30 band pass filter)
- Bring a stained live and dead culture to draw gates
Notes
Please feel free to post comments, questions, or improvements to this protocol. Happy to have your input!
- List troubleshooting tips here.
- You can also link to FAQs/tips provided by other sources such as the manufacturer or other websites.
- Anecdotal observations that might be of use to others can also be posted here.
Please sign your name to your note by adding '''*~~~~''': to the beginning of your tip.
*Taylor D. Scott (talk) 11:47, 3 May 2019 (PDT): I tested this on the Attune NxT:
- Grow culture of yMM1032 (FY auxotroph) to mid-log in low-fluorescence media
- Heat kill 1 ml at 95°C, 15 min
- Create different ratios of live/dead cells (in triplicate) in a 96-well plate (pre-filled with 50 μl methylene blue) (see table below)
- Transfer 50 μl stained culture to 200 μl PBS+ 0.1% Tween
- Analyze on Attune NxT ("River"), RL2 filter
- Gate for single cells, then draw a gate for live cells in a RL2-H vs FSC-H view
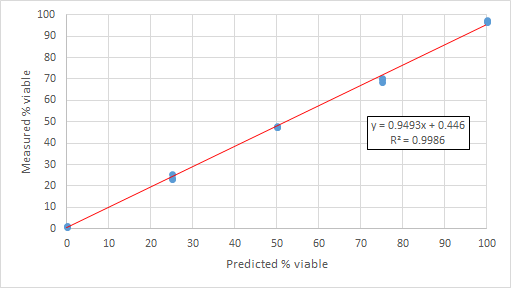
- Plot predicted % viable (from table above) vs actual % viable for each replicate
- Trend line is based on the mean of the 3 replicates
| Live volume | Dead volume | Predicted % viable | Measured % viable (mean) |
| 50μl | 0μl | 100 | 97.07 |
| 37.5 μl | 12.5 μl | 75 | 69.57 |
| 25 μl | 25 μl | 50 | 47.63 |
| 12.5 μl | 37.5 μl | 25 | 24.17 |
| 0 μl | 50 μl | 0 | 1.11 |
References
Relevant papers and books
- Painting, K and Kirsop, B (1990) - World Journal of Microbiology and Biotechnology 6(3) 346-7 PMID 24430080
- http://www.fluorophores.tugraz.at/substance/292
Contact
- Who has experience with this protocol?
or instead, discuss this protocol.