M465:Western blots part 2 and Biolog assays
Western Blots Part 2
Last time we met you transferred proteins from your insect samples to a membrane. Your instructor incubated your blots in primary antibody overnight and this afternoon, they are ready for the next step of the protocol. Remember that the primary antibody targeted a Wolbachia-specific protein called wsp. We will detect the primary antibody using a labeled secondary antibody, which can be visualized using the Licor. This machine detects the fluorescent tag that has been affixed to our secondary antibody (see figures below).
Before we get to that point, we must wash our blot, add our secondary antibody and incubate it at room temperature for one hour. Follow these steps below to complete the protocol:
Remember: wear gloves throughout and never let your blot dry out!
1. Your blots can be found on the nutating platform in the back of the room. Locate your blot (it will be labeled with tape and group name).
2. Your blot is currently in primary antibody. Pour this antibody solution into a labeled 15 mL conical vial and keep it on ice. You will reuse it later in the semester so don't throw it away!
3. To your blot, immediately add TBST wash buffer, enough to cover the blot (about 10 mL) - this does not have to be precise! Just make sure it doesn't slosh out of the container while it is nutating.
4. Place your blot back on the nutating platform for 5 minutes
5. Repeat steps 3 and 4 two more times (three total washes), leaving the blot in the final wash buffer.
6 . During the final wash, obtain the secondary antibody from your instructor - it will be provided in starting block buffer.
7. Pour off the final wash and add the secondary antibody to your blot. This antibody is light sensitive so we will make sure to reduce the amount of light you expose the blot to by covering the container in an opaque lid.
8. Place your blot on the nutator and set a timer for one hour. Proceed to the next portion of the lab (Carbon Source Utilization Profiling) below during this interval.
Continuing the blot
9. Now that your secondary antibody incubation is complete, you will wash your blot three times using TBST (as in steps 3 and 4 above). After your final wash, you will walk over to Simon Hall with your instructor to image your blots.
Carbon Source Utilization Profiling
You have learned in other courses about the importance of carbon fixation by autotrophic photosynthetic plants. The inability to make carbon-carbon bonds and, therefore, to utilize carbon dioxide as a carbon source, is problematic for heterotrophic species including humans and all other animals. Fortunately there are bacteria that, like plants, are autotrophic and photosynthetic, although many others are heterotrophic, like us. Unlike us, however, bacteria are extremely diverse in the types of carbon sources they can use metabolically. Bacterial communities both compete and co-operate in utilization of available sources of essential, useable carbon. The health and longevity of the community is dependent on a continuous supply of useable carbon for all its members. Your investigation on carbon source profiling will identify carbon sources that can be utilized by the Drosophila microbiome.
Carbon source patterns using BIOLOG™ Ecoplates
Observing patterns of substrate utilization across isolates can provide evidence of functional capabilities of that microbe in its community context. Additionally, understanding how metabolic substrates can be used in the communities can help us understand the stability or flexibility of that ecosystem. Carbon sources are crucial anabolic raw materials for heterotrophic microbial growth. Microbes vary enormously in their ability to make use of carbon in different forms.
In the following assay, you will use Drosophila lysates, inoculated directly into the single carbon source wells of microtiter plates followed by spectrometric quantification of carbon utilization. If your microbes can use a particular carbon substrate, the metabolism of that carbon source is accompanied by a reduction of water-soluble colorless Tetrazolium salts (WTS) to become reduced purple formazans. WST-1 and in particular WST-8 (2-(2-methoxy-4-nitrophenyl)-3-(4-nitrophenyl)-5-(2,4-disulfophenyl)-2H-tetrazolium (MTT), are reduced outside of the cells. They combine with an electron mediator (phenazine methosulfate (PMS)), to yield a water-soluble purple product called formazan that can be measured spectrophotometrically at 590 nm. The color development is additive and directly proportional to the metabolism of each carbon source so the development of forazan can be followed over time. The intensity of purple color as a pattern in the wells is used to determine the metabolic footprint of your microbial community. We will the patterns to determine the metabolic diversity of your community (CMD). For these measurements to be meaningful, it is important to control for incubation time, and other microenvironmental factors as well as the requirement for saturating substrate and indicator concentrations. There are 31 carbon sources available on a BIOLOG plate. This set of substrates is far from exhaustive. However, these substrates were chosen for variety and are similar to many nutrients found in natural environments.
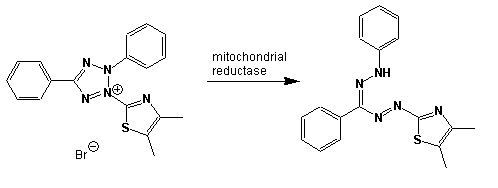
Scheme showing the reduction of MTT to formazan. Image created by Jenpen 21 September 2006
Source http://en.wikipedia.org/wiki/File:Mttscheme.png . Public domain use per Wikipedia Commons.
Colorless (2-(2-methoxy-4-nitrophenyl)-3-(4-nitrophenyl)-5-(2,4-disulfophenyl)-2H-tetrazolium) is reduced to purple formazan
The BIOLOG-ECO™ 96 well plates we will use contain 3 replicates of 31 carbon sources and three water control wells. The method for carbon source profiling that we are using is simple and rapid, but its interpretation must be carefully evaluated, recognizing that the methodology is imperfect. The references below discuss the pitfalls of the ecoplates in the context of community level analyses:
References and Resources: Biolog Carbon Source
• Garland, J.L., Mills, A.L. (1991) Classification and characterization of heterotrophic microbial communities on the basis of patterns of community-level-sole-carbon-source-utilization. Appl Environ Microbiol 57, 2351–2359.
• Garland, J.L. (1997) Analysis and interpretation of community-level physiological profiles in microbialecology. FEMS Microbiol Ecol 24, 289–300.
• Preston-Mafham, J., Boddy, l., Randerson, P.F. (2002) Analysis of microbial community functional diversity using sole carbon source utilization profiles-a critique. FEMS Microbiology Ecology. 42, 1-14.
Creating your Drosophila lysates and Inoculating your Plates
Do endosymbiotic community members (such as Wolbachia) alter the composition and or the function of the microbiota? Today you will ask this question by investigating what carbon sources can be used by communities of microbes found associated with four types of flies: 1) Wolbachia infected, 2) Spiroplasma infected, 3) co-infected and 4) uninfected. Each student will be provided with one of these types of flies. You will work collaboratively across the lab section to analyze the data resulting from this experiment. As a first step, you will make lysates.
- Salt Solution (137 mMol NaCl, 2.7 mMol KCl)
- P200 and P1000 micropipets with sterile tips
- Multichannel pipet (set to deliver 100 µL) and sterile tips
- BIOLOG EcoPlate™
- sterile plastic multichannel reservoir
1. Obtain a 1.5 mL tube with Drosophila flies from your instructor. Make sure to note what type of infection is found in this fly.
2. Grind the fly in the tube using 20 strokes with a sterile pestle (return to the beaker up front after using).
3. Using a micropipette, transfer the contents of the tube (500 uL total) to a 15 mL conical vial containing 12 mL of salt solution. Invert 2 times to mix the contents.
4. Move the Biolog ecoplate, your 15 mL conical vial, a sterile reservoir, and a multichannel to an available culture hood.
5. Pour the contents of the vial into the reservoir and, using the multichannel, pipette 100 ul of your diluted lysate into each well.
6. Be careful to preserve the BIOLOG Eco™ plate's and its cover's sterility (eg. don't place it face down on your bench). Remove the cover and transfer 100 µL of the dilution into each each well of the 96 well BIOLOG plate. Check visually the consistency of the amount of diluted extract in the micropipet tips after you have drawn up your aliquots to determine that you have no bubbles and that the quantity to be dispensed is the same. If the pipet tips appear unevenly filled or you have bubbles, do not dispense the inoculum into the wells! Start over. If you are unfamiliar with the use of multichannel pipets, ask your instructor to observe your technique.
7. Replace the cover of the plate and label one side of the cover. DO NOT LABEL ON the top or bottom to avoid interference in the light passage during spectrophotometric readings. Use a piece of your colored tape and include your initials, the date, and the isolate number.
8. Take a time 0 reading at A590 nm using the BioTek Synergy H1 384 plate reader (your instructors will show you how)
9. After each reading, place your covered back in the incubator (at the back of the room).
MEASURING MICROBIAL CARBON UTILIZATION AS A590 nm
The intensity of color change is monitored in each of the wells by taking spectrophotometer readings once a day at A590 nm. You will come in daily to collect these data until a peak absorbance is reached on more than 2 consecutive readings, this will likely require daily readings for a week. You must not miss more than 1 consecutive day. Make sure that you take a photo of your plate against a white background on the final day of measurement.
Carbon source utilization pattern:
How do we analyze our data? You could plot on a bar graph the average A590 nm absorbance of the three replicates (with error bars) on the final day of data collection on the y axis and the 31 different carbon sources on the x axis in one figure. (Remember that there is no such thing as a negative value for Absorbance so count anything that is less than zero as zero. Why might you seem to have a negative value?). Should you arrange those carbon sources in the order they are on the BIOLOG plate or is there a better way to organize them on your graph to show your main message(s)? What kind of information do you need in the figure legend?
Can you think of other ways to illustrate metabolic diversity of your community?


