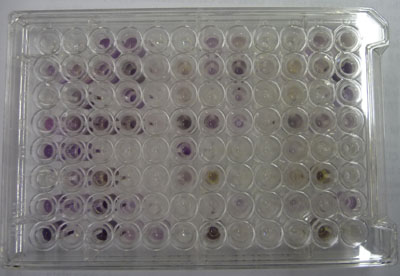
M465:Carbohydrateusage
Community Carbon Source Utilization Profiling:
Carbon Source Utilization
One type of metabolic diversity that we will assess in our investigation is physiological diversity in carbon source utilization.
You have learned in other courses about the importance of carbon fixation by autotrophic photosynthetic plants. The inability to make carbon-carbon bonds and, therefore, to utilize carbon dioxide as a carbon source is problematic for heterotrophic species including humans and all other animals. Fortunately there are bacteria that, like plants, are autotrophic and photosynthetic, although many others are heterotrophic, like us. Unlike us, however, bacteria are extremely diverse in the types of carbon sources they can use metabolically. Bacterial communities both compete and co-operate in utilization of available sources of essential, useable carbon. The health and longevity of the community is dependent on a continuous supply of useable carbon for all its members. Your investigation on community carbon source profiling will attempt to quantify some of that co-operation and competition.
Carbon source patterns using BIOLOG™ Community Level Physiological Profiling (CLPP)
Observing patterns of substrate utilization can provide evidence of functional diversity of a microbial community. Additionally, understanding how metabolic substrates can be used in microbial communities can help us understand the stability of an ecosystem. Carbon sources are crucial anabolic raw materials for heterotrophic microbial growth. Microbes vary enormously in their ability to make use of carbon in different forms. Using direct inoculation of community samples into a variety of carbon substrates will allow us to study and measure potential community carbon source utilization and provide us with evidence for our hypothesis that community diversity allows co-operative as well as competitive interactions.
In community level physiological profiling (CLPP) the metabolic properties of individual bacteria in the community contribute to the total metabolic capacity of the community. Mixed environmental samples are inoculated directly into the single carbon source wells of microtiter plates followed by spectrometric quantification of growth. If one or more microbes in the community can use a particular carbon substrate, the metabolism of that carbon source is accompanied by a capture of electrons from water-soluble colorless Tetrazolium salts (WTS) to become reduced purple formazans. WST-1 and in particular WST-8 (2-(2-methoxy-4-nitrophenyl)-3-(4-nitrophenyl)-5-(2,4-disulfophenyl)-2H-tetrazolium (MTT), are reduced outside of the cells. They combine with an electron mediator (phenazine methosulfate (PMS)), to yield a water-soluble purple product called formazan that can be measured spectrophotometrically at 590nm. The color development is additive and directly proportional to the metabolism of each carbon source so the development of forazan can be followed over time. The intensity of purple color as a pattern in the wells is used to determine a characteristic reaction patern classed a metabolic footrpint. We will use the patterns to determind average metabolic response (AMR) and community metabolic diversity (CMD). For these measurements to be meaningful, it is important to control for number of microbes, incubation time, and other microenvironmental factors as well as the requirement for saturating substrate and indicator concentrations.
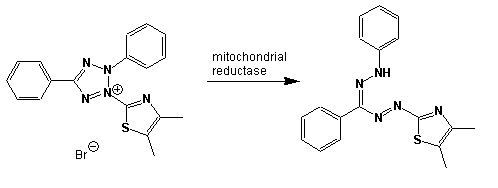
Scheme showing the reduction of MTT to formazan. Image created by Jenpen 21 September 2006
Source http://en.wikipedia.org/wiki/File:Mttscheme.png . Public domain use per Wikipedia Commons.
Colorless (2-(2-methoxy-4-nitrophenyl)-3-(4-nitrophenyl)-5-(2,4-disulfophenyl)-2H-tetrazolium) is reduced to purple formazan
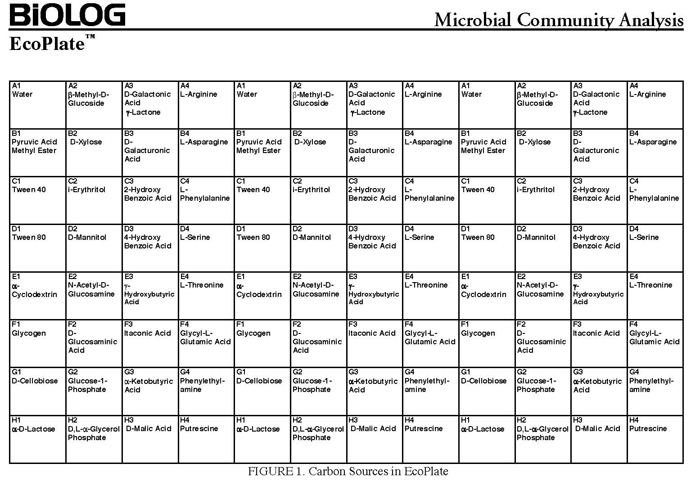
The BIOLOG-ECO™ 96 well plates we will use contain 3 replicates of 31 carbon sources and three water control wells. The method for community carbon source profiling that we are using is simple and rapid, but its interpretation must be carefully evaluated, recognizing that the methodology is imperfect. The following studies discuss issues and limitations of CLPP (community level physiological profiles) analysis.
These or similar references are provided in the lab Sakai site as .pdf files.
References and Resources: Biolog Carbon Source
• Garland, J.L., Mills, A.L. (1991) Classification and characterization of heterotrophic microbial communities on the basis of patterns of community-level-sole-carbon-source-utilization. Appl Environ Microbiol 57, 2351–2359.
• Garland, J.L. (1997) Analysis and interpretation of community-level physiological profiles in microbialecology. FEMS Microbiol Ecol 24, 289–300.
• Preston-Mafham, J., Boddy, l., Randerson, P.F. (2002) Analysis of microbial community functional diversity using sole carbon source utilization profiles-a critique. FEMS Microbiology Ecology. 42, 1-14.
BIOLOG Redox Dye Mix Brochure JUL07. http://www.biolog.com/mID_product.shtml.
PROTOCOL:
Carbon source patterns using BIOLOG™ Community Level Physiological Profiling (CLPP)
- 10mM phosphate buffer (sterile, pH 7)
- P200 and P1000 micropipets with sterile tips
- Multichannel pipet (set to deliver 100 µl) and sterile tips
- BIOLOG EcoPlate™
- sterile plastic multichannel reservoir
- Wear gloves throughout the entire protocol
- Do not cross contaminate your samples or the solutions
- Keep your work area clean: freshly disinfect your bench top before beginning
- Do not use a vortex at any point in this protocol unless it is specified that you should do so.
1. Use sterile serological pipets to make 12 mls of a 10-4 dilution (equivalent to 106cfu/ml). You will accomplish this by pipetting 10.8 mL of 10mM phosphate buffer into a large sterile tube and adding 1.2 mL from a 10-3 habitat extract serial dilution tube. (Note: You need 12 ml of diluted sample extract to inoculate all the wells of a BIOLOG™ECO plate (10 ml for the plate and 2 ml extra.) Our goal is to end up with 105 microbes in each well (in a 100μL volume).
2. Be careful to preserve the BIOLOG Eco™ plate's and its cover's sterility (eg. don't place it face down on your bench). Visually check the consistency of the amount of diluted extract in the micropipet tips after you have drawn up your aliquots to determine that you have no bubbles and that the quantity to be dispensed is the same. If the pipet tips appear unevenly filled or you have bubbles, do not dispense the inoculum into the wells! Start over. If you are unfamiliar with the use of multichannel pipets, ask your instructor to observe your technique.
3. Replace the cover of the plate and label along one long side of the cover. DO NOT LABEL ON the top or bottom to avoid interference in the light passage during spectrophotometric readings. Use a piece of your team color tape and include your initials, date, and any relevant sample code.
4. Take a time 0 reading at A590nm using the Spectramax340PC384 plate reader as described in the next section of these instructions.
5. After each reading, place your covered plate in a plastic container and incubate it at 37C in the location assigned for your lab section. Your instructors will ensure that they are kept humid to prevent dehydration
MEASURING MICROBIAL CARBOHYDRATE UTILIZATION AS A590nm
The intensity of color change is monitored in each of the wells by taking spectrophotometer readings once a day at A590nm. You and your partners should figure out a schedule to divide up the work of collecting these data until a peak absorbance is reached on more than 2 consecutive readings, this will likely require daily readings for 10-12 days. You must not miss more than 1 consecutive day. Make sure that you take a photo of your plate against a white background on the final day of measurement.
Time 0: Use the Epoch Synergy spectrophotometer (with a microplate reader) found in the Microbiology lab. You will measure absorbance as A590nm. Follow the directions for setting up the instrument and software and for exporting the data to Excel spreadsheets.
Using the Synergy H1 and the Gen5 Software
Epoch Synergy

Turn on the Synergy spectrophotometer using the toggle switch on the lower right corner front. Avoid pushing the small button to the left of the toggles switch for now. The machine will automatically start a required calibration. Allow up to 10 minutes for the Synergy H1 to warm up after the calibration and BEFORE you attempt to insert your plate.
The drawer may or may not open. You can manually close and open the drawer by using the small button on the lower front of the machine.
The laptop computer attached to the machine should be on. (If not, turn it on using the on/off ). The passwords for the account are on a post-it attached to the Synergy system.
When the warm up period is over, double click on the Gen5 shortcut on the desktop of the computer.
Select "New" and then select "Plate Type". You will be using a 96-well flat bottom plate.
Select Read. Under the read menu, select detection method (absorbance), read type (end point), and read speed (normal). Also set the absorbance to A590nm.
Select button, upper right of window, called "full plate"
Press OK, and the window returns to Read step
Press oK, and the window returns to the Procedure step
Press OK
The drawer should open. If it doesn't, you may manually open it by using the small button on the lower front of the machine next to the toggle switch.
Place your plate on the carrier and click OK.
After the read, be sure to SAVE the data onto a flash drive. You can also export your data to excel if desired and save.
Remove the plate and close the carrier.
Shut down the EPOCH and the computer.
Calculating Average Metabolic Response (AMR) and Community Metabolic diversity (CMD)
CALCULATING AMR: Average Metabolic Response over time and between sites and Average Carbon Source Utilization Ccore by the microbial community
Average Metabolic Response(AMR)= Σ(mean A590nm of triplicate carbon source tests - Mean A590nm of the three control wells) and less a threshold absorbance of 0.25 / 31 (the number of carbon sources tested)
The calculation of AMR provides a method to compare total metabolic activity by a community among aerobic or facultative microbes. The AMR is calculated as the average difference between A590nm values of the three replicates of each C source and the 3 water control wells. The following calculations can be built into an Excel template workbook: Take the average of each of the 3 replicate wells and subtract the average of the control wells from each of the carbon sources. A background absorbance threshold absorbance of 0.25 is substracted also. If you double click on the cells, you can check to see that the formulas and calculations are correct. The final calculation that is already built in is a division by 31 (the number of carbon source wells).
A high score indicates that there were many useable carbon sources for the community, a low score indicates that few of the carbon sources were able to be metabolized well. Calculate AMR for every date read. On the last page of the template workbook, please fill in the AMR table by day and overall average absorbance.
CALCULATING CMD: community metabolic diversity. CMD is a simple way to represent the total number of substrates able to be effectively metabolized by the microbial community. It's a measure of diversity in use of carbon sources. CMD is calculated by summing the number of positive responses (wells with a positive A595nm value after all the corrections) at each incubation time. Any negative values are considered 0 absorbance.
On your worksheet enter either a 0 or a 1. Zero indicates that there was a negative value or 0 for corrected A590nm. A 1 indicates a postive value of greater than zero. Once you have entered a 1 or a 0 as the "corrected" absorbance for all the cells, the template will calculate the CMD for that day. Complete the final CMD table on the last page of the template workbook.
GRAPHING THE DATA:
Once your team is finished collecting the data, make figures for AMR, CMD, and Carbon Utilization patterns.
To graph AMRs: You will plot the AMR value on the y axis against time on the x to provide a quick comparison of metabolic responses by day.
To graph CMDs: Plot the calculated sample's CMD values on the y axis versus time on the x to get a sense of community functional metabolic richness.
Carbon source utilization pattern:
Neither AMR nor CMD analysis provides information about the pattern of carbon substrates used in each community.
To examine the pattern of carbon sources used across communities, you could plot the average A590nmabsorbance on the final day of data collection on the y axis and the 31 different carbon sources on the x axis in one figure. (Remember that there is no such thing as a negative value for Absorbance so count anything that is less than zero as zero. Why might you seem to have a negative value?)
Can you think of another way to compare the patterns of substrates?