LuisM Turing Patterns with Stochastic π Calculus
A global Context from the iGEM Mexico team and Turing Patterns
One of the most important problems in developmental biology is the understanding of how structures emerge in living systems. Several mechanisms have been proposed, depending on the observed patterns. The so called Turing patterns are based on the interaction of two effects: diffusion of some chemicals, called morphogenes, and the chemical interaction between them. It has been highly controversial whether some patterns observed in several organisms are of this type. In particular, although some systems have been identified to be of activator-inhibitor type (the most popular Turing system proposed by Gierer and Meindhart), it is still questioned if pattern formation and more generally, the appearance of functional structures can be understood by means of Turing patterns or more broadly, reaction-diffusion mechanisms.
Talking about Patterns
After the last jamboree(Nov, 2007) some members of team started a quest with the objetive of generate turing patterns at the lab and since a theoretical way. A teammate Juan A. investigated the theoretical part to produce Turing, and in a nice afternoon of january he explained to Federico and me his results:
There are many ways to produce the patterns, but essentialy we need an Activator-Inhibitor system.
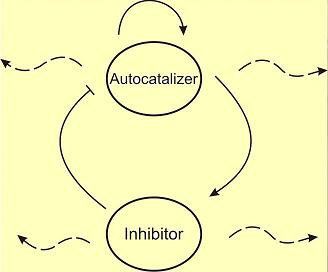
The Activator-Inhibitor system is composed of two parts: An Autoctalizer and an Inhibitor.
The Autocatalizer promotes its own synthesis (a positive feedback loop) and in turn induces the production of the inhibitors. When the Inhibitor is present, it generates a signal that repress the autocatalizer (Negative Control). The whole system might behave in an oscillatory way.
The autocatalizer and the Inhibitor produces diffusible signals that allow communication among other neighbors, in particular we need that the inhibitor signals spread faster than the autocatalizers' with the objective of getting syncronization in a global level and eventually generate oscillatory patterns. It is important to emphasize that no efective morphogen is needed: the internal dynamics and intracellular communication are sufficient to generate patterns.
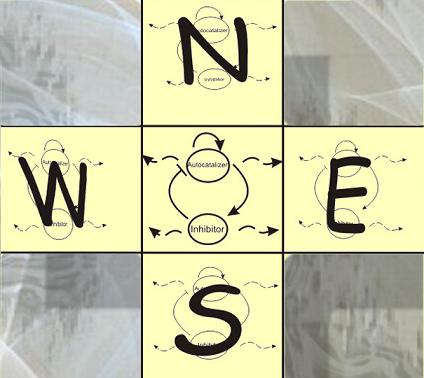
In a first approach to model this kind of system the most simple structure is an array of cells where each of them has internally the Activator-Inhibitor system. How these system takes in account the behavior of the neighbors the most simple neighborhood is the Von Neumann neighborhood from a cellular automaton, which consist of 4 ortogonal cells interacting with a central one:
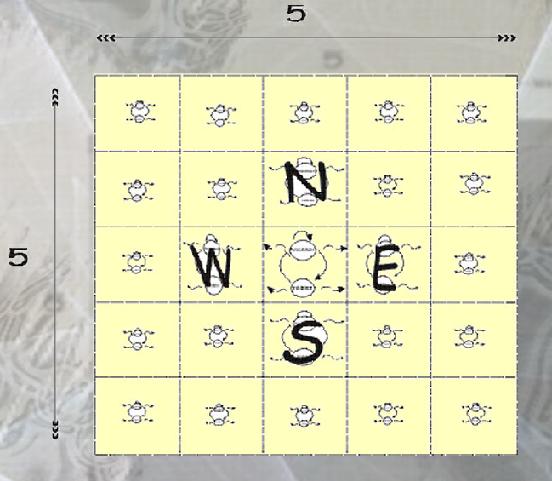
The fist model that I did was based on an array of 25 cells (5x5) and in the Stochastic Pi Calculus Model of the Complete Array (5x5) the diffustion signals form the Autocatalizers and the Inhibitors in the neighborhood are represented by the channels autocat'Neighbors' and inhibit'Neighbors' where the set 'Neighbors'={North,West,East,South} represent the system around a central one.
And by this way we have add a spatial distribution to the model.
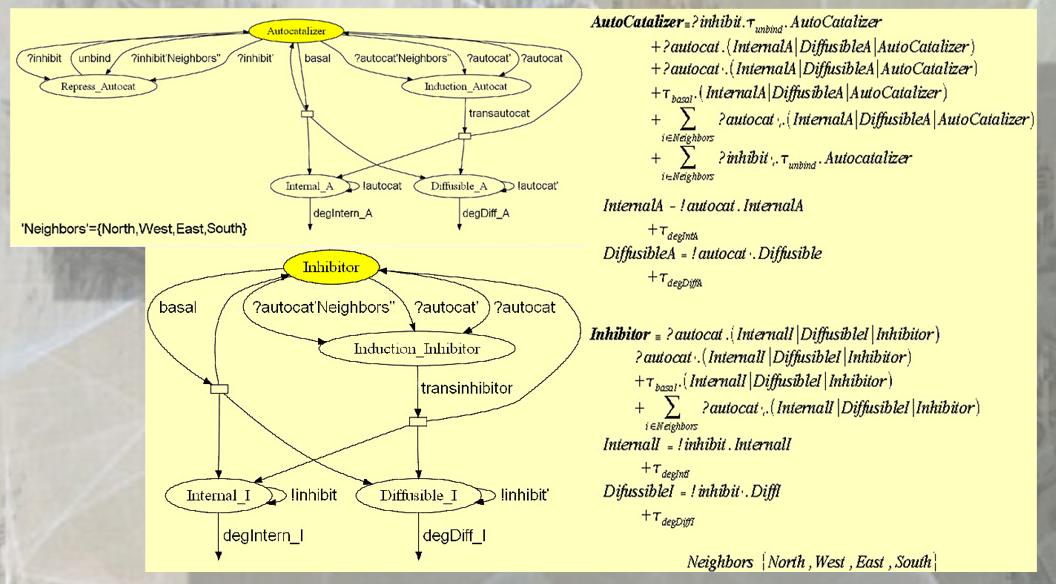
Once I had in mind how the global system behaves, i started to create the model in Stochastic Pi Calculus using the new notion of spaciallity that I defined with the neighborhood of Von Neumann and the model is the next:
The model with Stochastic Pi Calculus
Then the simulation in the Stochastic Pi Machine was done giving a file with the concentrations of the 25 different inhibitors, the results of the simulation says that there are communication in the system between the cells and when there are enough expression of cells around a central one, it can obtain a more expressed state represented with a high orange color. Some images of the results are the nexts: