Lidstrom:in vitro Pathway Assay: Sample Analysis via UPLC-MS/MS
Back to Lidstrom Protocols
Accela HPLC Protocol
Background
Column & Buffer
- We use a C18 column, and 5% methanol in our buffer.
- "Separations of complex mixtures normally require the elution of analytes from a buffered aqueous to an organic mobile phase composition (e.g., 5–100% organic). Thus, analytes are eluted in the order of their increasing polarity.The separation of very polar and/or ionic compoundsgenerally requires the use of a highly aqueous eluent composition (>95%) to provide some retention. Such a hydrophilic environment on the highly hydrophobic C18 stationary phase can lead to a loss of retention, increased peak tailing, and irreproducible results. This de-wetting effect (often incorrectly referred to as phase- collapse) (49) can be significantly reduced with the application ofLC columns with specially designed stationary phases (using polar embedded groups or polar end-capping, such as Aquasil/AQ columns)" - Mass spectrometry-based microbial metabolomics 2012 (Baidoo, Benke & Keasling)
- This is a "reverse phase" (hydrophobic) method
- "Separations of complex mixtures normally require the elution of analytes from a buffered aqueous to an organic mobile phase composition (e.g., 5–100% organic). Thus, analytes are eluted in the order of their increasing polarity.The separation of very polar and/or ionic compoundsgenerally requires the use of a highly aqueous eluent composition (>95%) to provide some retention. Such a hydrophilic environment on the highly hydrophobic C18 stationary phase can lead to a loss of retention, increased peak tailing, and irreproducible results. This de-wetting effect (often incorrectly referred to as phase- collapse) (49) can be significantly reduced with the application ofLC columns with specially designed stationary phases (using polar embedded groups or polar end-capping, such as Aquasil/AQ columns)" - Mass spectrometry-based microbial metabolomics 2012 (Baidoo, Benke & Keasling)
Ion Pairing
- Our method is ion pairing. (Tributylamine in buffer C)
- "The application of ion-pairing reagents in RP LC improves the to separations of these types of compounds. However, the addition of these ion- pairing (IP) reagents is often disadvantageous for MS detection (due to ion-suppression and the possibility of contamination after prolonged used)." - Mass spectrometry-based microbial metabolomics 2012
- ion pairing: for when analytes include basic or ionic compounds that are difficult to chromatograph by reverse-phase LC. In IPC, the mobile phase contains an ion-pair reagent that attaches to the stationary phase and creates a charged surface. The sample ion exchanges with the counter ion of the ion-pair reagent thus increasing interaction with the column resulting in greater retention of the sample.
References
- Ultrahigh performance liquid chromatography-tandem mass spectrometry method for fast and robust quantification of anionic and aromatic metabolites, 2010. This is the protocol Amanda & Justin used to get ours going.
- Their LC protocol:
- “Separation of compounds was achieved by an ion pairing-reverse phase method developed for ultrahigh performance systems based on previously published high pressure methods15,17 and eventually implemented on a Waters Acquity UPLC (Waters Corporation, Milford, MA, United States) using a Waters Acquity T3 end-capped reverse phase column with dimensions 150 mm × 2.1 mm × 1.8 µm (Waters Corporation, Milford, MA, United States) temperature-controlled at 40 °C. A gradient of mobile phases A (10 mM tributylamine, 15 mM acetic acid, 5% (v/v) methanol) and B (2-propanol) was used to separate metabolites (Table 1). The injection volume was 10 µL with full loop injection. The column was equilibrated for 4 min (4.4 column volumes) before each injection. “
- Their MS protocol:
- “Selective and sensitive detection of compounds was achieved by coupling the liquid chromatograph to a Thermo TSQ Quantum Ultra triple quadrupole instrument (Thermo Fisher Scientific, Waltham, MA, United States) using a heated electrospray ionization source (Thermo Fisher Scientific, Waltham, MA, United States). The mass spectrometer was operated in negative mode with multiple reaction monitoring. Electrospray ionization parameters were optimized for 25% 2-pro- panol at a flow rate of 0.4 mL/min and used for the entire gradient: spray voltage 2500 V, sheath gas pressure 80 arbitrary units, aux gas pressure 50 arbitrary units, ion sweep gas pressure 5 arbitrary units, capillary temperature 380 °C, spray temperature 400 °C. Tube lens voltage, collision energy, and fragment ions were optimized individually for all compounds (Table 2). Ion optics were set to 0.5 amu Q1 resolution, 0.5 amu Q3 resolution, 0.01 amu scan width, and 10 ms dwell time. SRM scans were organized in 11 time periods so that each compound could be detected for more than 1 min before and after expected retention time and a scan frequency of at least 2 Hz was maintained. Data Analysis. Both acquisition and peak integration were performed with Xcalibur software version 2.07 SP1 (Thermo Fisher Scientific, Waltham, MA, United States) and in-house integration software (Begemann and Zamboni, unpublished). The criteria for quantifiability of a standard compound were that a minimum of 4 neighboring dilution steps showed a linear cor- relation between the logarithm of the concentration and the logarithm of the peak area with a correlation coefficient greater than 0.99. Additionally, the peak area of the lower limit of the linear range had to be larger than the peak area found in water."
- Their LC protocol:
BEFORE SETUP NOTES
Columns:
- Waters Acquity C18 Column. Bigger blue box. (Water Acquity UPLC HSS T3 1.8 um 2.1 x 150 mm column)
Part no. 186003540)
- Also tried:
- Hypersil Gold AX, 100 x 2.1 mm, Thermo-scientific
- Hypersil Gold aQ, 100 x 2.1 mm, Thermo-scientific
- PFP column: Hypersil GOLD PFP, 100 x 2.1 mm, Thermo-scientific
- Hypersil GOLD 100 x 2.1 Thermo Scientific #25002-102130
Our Buffers:
- C = 5% (vol/vol) MeOH, 15mM acetic acid, 10 mM tributylamine
- for 10L: 2.38 mg/L tributylamine, 0.857 mg/L acetic acid, 5% methanol.
- for 10L: 9.468L dH2O + 23.8 mL tributylamine + 8.57 mL acetic acid + 0.5 L methanol
- tributylamine = ion pairing reagent
- do you have enough buffer C to do your run? (calculate it)
- (7/8) = estimate of fraction that is yellow (buffer C) (this is conservative)
- max use = 400 uL/min
- run = 36 min/sample
- # of samples = n
- If volume of buffer c is greater than (7/8)*(400 uL/min)*(36 min/sample)*n, you can start. If not, make more buffer C. Amanda makes 10 L batches. Justin says it is very sensitive to variation, so he waited for Amanda to make it on 6/20/2012.
- multiply (7 / 8) * 0.400 * 36 *n to get mL needed
- you can't run an experiment using two different buffer preparations. If there is not enough for your purposes, make more before you start and discard the leftovers. Look for the bottle labeled waste, or make one. Justin might dump it down the sink.
- wash is the same composition as buffer C but in a different bottle. The composition is written in orange tape but it says both 15 and 20 mM acetic acid. (15 is correct.)
Opening a New Column: (?)
- Record on spec sheet:
- Date opened
- Pressure and associated flow rate
- Solvent used
- Rinse all parts
- Insert guard column cartridge into guard column
- Don’t touch guard column cartridges while inserting it!
- Use socket wrenches to tighten screw into bottom of the guard column.
- Finger tighten guard column into the bottom of the real/separation column.
- Cap both ends with screw-in end-caps.
- Column is ready to be used.
Solvents
- A: Acidified H2O (H2O with 0.1% Formic Acid)
- B: Acetonitrile 0.1% Formic Acid
- C: Rotating Tap (Has our buffer on it now)
- Buffer C is an ion pairing buffer: 10 mM tributylamine, 15 mM acetic acid, 5% methanol. Stored in a 10 L bottle on the floor, under the desk. The recipe is written on the tape.
- D: 100% Acetonitrile or Methanol or Isopropanol
- Our experiment uses isopropanol
Normal Wash:
- Has its own bottle. We aren't currently using it
- 20% H2O
- 80% Acetonitrile
- 0.1% Formic Acid
- 20 mM NH4Acetate
Our Wash:
- Same as mobile phase C
PHYSICAL SETUP
Check these before running, you may not need to change any, or may need to adjust everything.
- Open an existing file (Janet started own folder on 6/20 from an old formate run of Amanda's)
- Study name: keep this general. Workbook: get more specific (date, etc.). You can have multiple workbooks per study.
- Check the change root study folder box if you want the workbook in a
Big Picture Steps/Things to Check Before You Start the UPLC-MS
- Right column?
- Right Wash Buffer?
- Washed needle?
- Right tray setup? (wells vs vials)
- Mobile phase/buffers are full enough?
- Right MS probe installed?
- Liquid N2 full enough?
Example workflow
- Check Buffer Levels and Correctness for run
- Change Column (Want to do this before changing buffers because you want to store it in whatever the previous user left at mobile phase D.
- Flush old column (~10 min)
- While column flushing, start spinning down samples and prepping filter plate (5 min)
- Insert new column (~5 min)
- Change Buffer, if not correct for run (~22 min)
- Set oven temp (helps you stay below pressure limit)
- Equilibrate Column (~15 min)
- Continue while column changing buffers and equilibrating column
- Sample Prep
- Change wash buffer
- Change MS probe, if necessary
- Change Autosampler Tray Settings, if necessary
- Clean plate seals
- Setup sequence file in LcQuan/Xcalibur
Changing the Column
Flush previous column before removing
- flushing previous column: mobile phase D
- mobile phase D may be 100% acetonitrile, methanol or isopropanol. Use 0.5 mL/min, 10+ min
- flush the old column with its buffer in preparation for storage
- Open Xcalibur.
- Go to Instrument Setup --> Acela Pump (icon on left bar) --> Direct Control (words on top bar).
- Check the box next to “Take pump under direct control.”
- Change flow rate to 500 uL/min
- If the pumping stops do to being over pressure, decrease the flow rate and run longer.
- Type 100% next to solvent D (100% Acetonitrile)
- Press Play.
- Run for at least 10 minutes (a timer counts)
Stop pump and remove previous column
- Note: All necessary wrenches are inside the bottom of the autosampler box (AS). The pumps can continue while the column is being changed or they can be paused.
- Manually unscrew the output (top) of column: the flexible red hose and one hex-shaped piece of metal should be all that remains.
- (while holding column) Using two appropriate wrenches unscrew the very bottom connector from the metal HPLC input.
- all that should remain is a metal wire with red tape, a hex-shaped piece of metal and a metal nipple.
- most columns have a short extra column on the input with a small, 1cm “guard column” which is easily replaced if damaged. It is housed in a short metal pre-column. This should be stored with each column.
- Place caps on the removed column and place back in the appropriate box, usually on top of the MS unit. (Box and end-caps for column in use should be stored on the top of the MS)
Insert your column
- Uncap the new column. (6/20/2012: acquity (blue box) pore size = 1.8 um, dimensions: 2.1x150 mm)
- Read direction of flow. Flow goes from the bottom up, so make the arrows on the column point up
- Attach bottom of new column, being careful to align threads to avoid leakage, using wrenches.
- Crank the bottom thread tight: this is ~ 400 atm of pressure.
- Manually attach top plastic connector. Push red tubing in hole first, then screw in connector
- Dry connections so any liquid will indicate a leak
Equilibrate your column
- Equilibrate column to starting conditions (& make sure column is on tight)
- In Acela Pump Direct Control, set starting conditions to run (e.g. 100% Solvent C, 0.4 mL/min).
- In Xcalibur, click instrument setup -- > “Accela pump” menu -> Direct Control. A new window opens. Check “take pump under control,” change parameters, then hit “Play”.)
- Can set temp = 40oC to keep pressure below limit. The method sets the oven temp later, so this is not critical.
- Pressure should equilibrate over ~1 min to ~570 bar. Check for leaks at the column and guard column, adjust as necessary.
- If pressure is only 400 bar, there is probably a leak. Recall 1 bar is about 1 atm, so this is HIGH pressure!
- Adjust as necessary = lower flow rate if pressure > 700 ish. 6/20/2012: Amanda set the oven temp to 40oC because the pressure was too high.
- If it hits the high pressure limit you set, it will stop pumping and the pressure will drop. You must lower the flow rate and turn it on again to try again.
- Run the column for 10 minutes to equilibrate and catch leaks. (10 min starts once you have 0.4 mL/min and pressure < 600 bar)
- If you are running < 400 uL/min, running more than 10 min can compensate for this lower flow rate. Amanda bumps it to 400 for a couple of minutes before calling the job done.
Changing the MS Ionization Unit
MS ionization unit: APCI (chrome)/ESI (matte brown) – (we use brown) Careful because parts may be hot!
- APCI is kept in a white box on the shelf above the computer, labeled APCI. The ESI is usually stored on top of the MS unit facing aack.
- Press button on front of MS to switch flow to waste
- Loosen the knob holding the ionizer head in the chamber and gently pull out the head, twisting to remove the alignment pin. Set aside head.
- Remove all cables from the ionization chamber, open up the two fasteners and pull directly back to remove (don’t forget the waste hose!)
- Put in desired ionization unit
- To put in ESI, remove the pin inside the chamber with needle nose pliers, rinse with methanol, place pin inside Ziploc bag in apci box.
- To put in APCI, take the pin from the Ziploc bag in apci box, rinse with methanol, dry with kimwipe, and place into pin-holder in the chamber. #Touch the pin with neither bare nor gloved hands, lest you besmirch it. Pliers should be nearby.
- Clean cone that points out at you if dirty (there is some rust/crust). See cleaning notes.
- Place chamber back on MS unit, closing fasteners
- Insert new head, lower to appropriate depth (C in my protocols), tighten
- Attach all cables. Note that for APCI the high voltage connects to the chamber directly, not to the back of the ionizer. Also note that for ESI the sample input line from the valve goes first through a metal connector (to ground) and a separate input line connects that metal intermediary on the chamber to the back of the ionizer.
Change mobile phases (C & D) if necessary
- C = tributylamine + acetic acid + methanol
- D = isopropanol
- Remove filter line from old bottle
- Rinse filter line with the new mobile phase
- Put filter line in bottle of new buffer.
- Attach a syringe to the line coming out of the purge valve in the pump unit under the autosampler
- Open the purge valve
- Start purging the line
- Open Xcalibur.
- Go to Instrument Setup --> Acela Pump (icon on left bar) --> Direct Control (words on top bar).
- Check the box next to “Take pump under direct control.”
- Change flow rate to 1000 uL/min
- Type 100% next to the mobile phase you’re changing
- Press Play.
- Run for 20 min.
- Close purge valve.
- Remove syringe and dispose of content
Changing the Wash/Flush Buffer
- Wash buffer is the same composition as buffer C, but it has a separate bottle. If you need to change it, you need to wash the needle afterward.
- Remove filter line from old bottle
- Rinse filter line with the new flush buffer (Amanda rinsed with water that was stored up there; can also rinse with water or new flush buffer in a beaker. here "new flush" = water because our solvent is mostly water)
- Put filter line in bottle of new buffer
- Put paper towel in Autosampler where wrenches live to catch extra fluid.
- Use "Wash Needle" command to flush lines
- Open Xcalibur or LCquan.
- Go to Instrument Setup --> Acela AS (icon on left bar) --> Direct Control (words on top bar).
- Choose “Wash Needle” from the drop down box.
- Change volume to 6000 uL (6 mL)
- Make sure the source (reservoir) is set to “Flush Bottle”
- Press apply. (Autosampler door must be closed)
- Repeat 2 more times. (it is fast(1-2 minutes); you can hear a hum pulsing when it is going.)
Choose tray type
- Quit Xcalibur/LCQuan and open the Instrument Configuration program on the desktop
- Choose Acela Autosamper (AS) under Configured devices
- Choose the and the correct type (e.g. for plates, use 96 well microwell plate + carrier + riser and for vials, use 1.8 mL vials)
- Quit configuration program
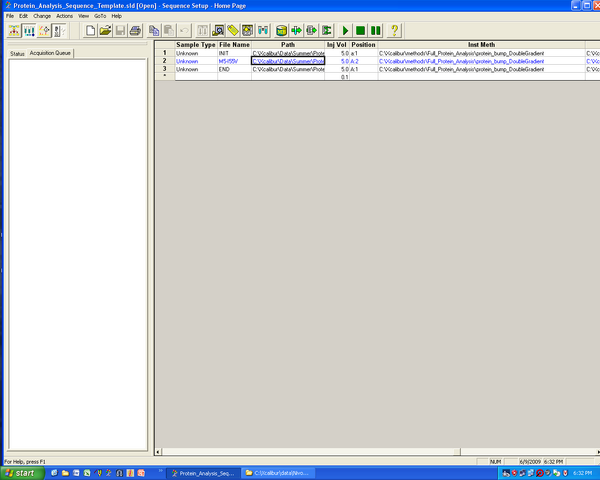
- In Xcalibur under sequence setup (lower image) choose “Change tray names” and choose new tray type.
- for 96-well plate: scroll down to "96 Well Microwell + Short Microwell Carrier + Riser Plate"
- It looks like the direction of sampling is wrong (bottom one selected) but Amanda says it is fine.
- Put/remove in riser and carrier as needed.
- pop out vial tray, put in riser (heavy metal block), then carrier tray (holds 3 plates)
Notes
- For plates positions are enumerated A:A1, etc. (first A indicates first of three plates), while for tubes numbering begins A1, etc.
- The plate positions are from the front, closest to the door, in the order, A,B,C.
Preparing & Loading Samples
- centrifuge 1.5 mL tubes (you quenched rxn in) for 10 min at 14,000 rpm & 4°C
- Move supernatant to 96 well plate (type = MSRLNO4: don't use another type! others are for different hydrophillic/hydrophobic)
- get a plate from cold centrifuge room: above plate reader that they are also used for
- MARK THE WELLS you use. Otherwise someone may try to use them again
- Tape the filter to the collection plate (don't tape the lid on)
- put on plate shaker for 5 min at 1,000 RPM without a lid (noisy)
- NOTE: load carefully to avoid splashing! Keep cover on for loading, then remove.
- centrifuge in plate centrifuge in cold room for 5 min at 1,500xg (not 5,000 RPM or 1500 RPM)
- after centrifugation, remove filter & lid (put in used pile if unused wells exist)
- Load a few wells with 75:25 acetonitrile:water
- Set up 3 or 4. We only inject 5 μL each time, but the extra wells are to make sure that nothing building up in the blank by rotating between a few.
- Amanda makes it fresh each time. Reagents are in ~ 2 inch tall bottles in cabinets by the liquid N2 tanks.
- use 150 uL/well: this is the max you can have in a well with a plate lid. This volume is large such that if there is needle contamination, it is diluted
- Note: yes, you started with 200 uL sample but you lose some to the filter, so the volume ends up being around 150 uL/sample.
- Add standard(s)
- Amanda has standard(s) in little vials in the -20°C freezer in the hallway. They are set up to be diluted in 3 volumes of acetonitrile (4x stock). This time, she had me add 15 uL to 45 uL of acetonitrile.
- Put a clean rubbery cover/plate seal on top.
Preparing & Loading Samples
Cleaning plate seals (easier to do a bunch at a time)
- Rinse with Methanol
- Rinse with Mass spec water
- Rinse with Methanol
- Set out to dry on paper towels or blow dry with air hose if you need it immediately
Load Plate into machine
- See if correct tray is in, then load into machine
- which of the 3 spots do you put the tray in?
- orientation matters: if handle is toward you, put 1A in the top left.
- The closest tray to the handle is A and the farthest is C.
Changing the Liquid Nitrogen Tank
- 1/4 tank might last a whole day, but it is good to check on it. Make sure you have much more if you are going to leave it over the weekend.
- close the use valve on the empty tank
- get wrenches from the drawer under the MiSeq. They are big & adjustable.
- take off the regulator - can be physically difficult. Counter clockwise, as you'd expect.
- put regulator on the use valve of the new tank
- Open use valve on new tank & set pressure to 100 PSI by using big blue turn valve.
- Note: a problem with full tanks: since it is mostly liquid, there isn't a lot of pressure. There is an orange doughnut-shaped tag that shows what the pressure should be. Open the pressure building valve until the pressure reaches the stated pressure.
- Note: on 8/31/2012 we had a tough time getting it open and only got a little bit of flow.
Enter sample names in LCquan
- enter samples into setup page of interface. This order indicates the order of sampling.
- enter wells as you entered them on the plate up high, then paste below the rows as you plan to use them. Put some sort of obvious spacer row inbetween
- sample blanks for the 1st 2 injections. The first one or two are always junk.
- then do a standard, and a blank.
- do some blanks (75:25 acetonitrile:water) every 7-10 samples at natural brake points
- do blank, standard, blank at end.
- you can select a subset of these to run (don't worry if you aren't planning to use every well)
- Tell it "yes" when it asks you if you want to wait for the instruments to be ready
Running with pre-setup software
- check that N2 is on (100 PSI) & the tank is full enough (look at bobs on top)
- if the pressure drops too low, the run stops with a generic mass spec error.
- set oven temp to 40°C, can do this under the Acela AS -> Direct Control -> Set oven temp
- check that ____
- _____
Obtaining Data
- the program forces you to save before exiting
- to download data, click on quantitate screen, then File - Export -
- settings: one file for all components. arrangement = Quan Results Grid. You can change it to comma separation.
Finishing with Equipment
- flush column? Leave it on?
- next person should flush it and remove it. If it is going to be a while before anyone uses it again, you can flush your column and store it. ?? Can you leave it without a column?? better ask first.
- Leave liquid N2 on
Misc
- Do we need to order more liquid N2 tanks or does someone take care of it?
- Tell Adam
(?) SOFTWARE SETUP
- Set up a method for a new run (usually already completed previously)
- Open Xcalibur and choose instrument setup
- Choose “Acela pump” to setup gradient, e.g. starting at 95% A to 5% A. Click in the last row and start typing to add a new section. After changing the gradient program, note how long it is and update the run length for the TSQ Quantum and Acela PDA.
- Choose TSQ Quantum to setup MS. Click the “…” button to select your tune (tsq) file and be sure to update the parameters from the relevant tune, e.g. switch to SIM if necessary, etc.
- Can add masses to be detected for each scan event by adding the parameters to the table at the bottom of the window.
- If you didn’t do so under Column directions, equilibrate column to starting conditions, ~10 minutes.
- Manually run at x% A. E.g. if the run starts at 40% A, run at 40% A, following instructions above under “(1) Column”
- Set up sequence.
- In windows, create a new folder for your run, usually in C:\Xcalibur\Data
- click “sequence setup”
- If you’re using a version of an old sequence, simply open, save as, and edit.
- Adjust “File Name” (e.g. compound1), Path (click to select), injection volume. Position goes A:A1… for plates, A1… for tubes. Click to browse for the desire method, already created previously.
- Use the “fill down” button (two downward pointing arrows) to copy values down or automatically increment plate values (A1, A2…) after selecting desired column.
- Usually a sequence should start and end with a blank, e.g. PBS
- Run stds at the beginning, beginning with lower concentrations first.
- Run the sequence by pressing the run sequence button (right pointing arrow with a sheet on top).
- To start just one or a set of the sequences select the relevant row and click run sample (right pointing arrow with a test tube)
- Before starting, make sure that all instruments needed are “on” in the Status tab. E.g. if mass spec is used in the method the machine should be “ready to download”
- In the window that opens, check/uncheck post-processing as needed, make sure that the Acella AS is the starting instrument, and make sure box is checked such that pumps turn off after runs.
- Click the “acquisition queue” tab to see the loaded set of sequences. If you need to stop any sequence, check the box and press the “delete” key.
- If not using the divert valve, make sure that the MS is set to load/detect.
Observing Results on Baker Computer
- Internal standard contains:
- 12C: 3PG, PEP, pyruvate,
- doesn't contain: formate,
13C:
PROTEIN SETUP (EXAMPLE) – I don’t use this {From the Baker Lab's protocol, we don't use this}
Overall guidelines: - Generally use 1mg/ml protein, absolute max is 2 mg/ml; at higher concentrations precipitation will clog the switches - 10-15 ul sample per vial, with insert - 50 ul sample per well in plates
Column: BioBasic-4, labeled “Current” Ionization: ESI head Buffers: Water/Acetonitrile each with 0.1% formic, equilibrate to 95% water/5% AceN Flow Rate: 500 uL/min (in rare circumstances use slower flow for higher sensitivity)
Setup template (open in xcalibur; saved in read-only format on Desktop): Protein_Analysis_Sequence_Template.sld Save the template under a different name/directory Adjust sample names; include a blank (e.g. PBS) in first and last run At run time, be sure that Processing boxes are checked for Qual and Programs.
ProMass will run automatically Mass error is +- 0.02% To view results: Promass results folder, index.html See data sheet on top of computer for common protein modification masses
To reprocess data, if anything goes wrong, hit the “summation” icon in xcalibur.
Tuning
MS Tuning notes for Accela HPLC/MS:
Note: When not running, inject/waste mode, otherwise load/detect mode.
- Sample injection:
- Use direct injection of salt and buffer free sample solution. For example using 1mM of the molecule of interest in 50% AceN/50% 0.1% formic acid water. Do not use any buffer, e.g. HEPES, or include any salts, as they may build up in the MS.
- Start with direct injection (we didn’t do this):
- To directly inject use a single tube with the needle adapter (on top of the MS; usually the needle adapter would be attached to the T-junction).
- Rinse the syringe (250 uL or 500 uL as needed) with solvent (5-6 times)
- Load sample as prepped above. Syringe should be placed into the syringe injector after it is attached to the input tube. Follow instructions in step (2) to turn on syringe pump.
- After you see compound in direct injection, go to inline injection (we started with inline injection):
- Use the T-junction. Attach the input line from the switch to one leg of T, attach input from syringe to other leg, and attach third leg to input of ionizer. Using direct control of the pump (icon in Quantum tune with a syringe flowing in to three collector flasks) run the normal HPLC buffers – note that if the syringe is not loaded in the injector the pressure will push the syringe plunger out!
- Vanilla Tuning
- Open Quantum Tune and run the MS (click on “pause” button in upper-left corner so it becomes a green triangle)
- Before changing anything chose “save as…” to select a new name for your tune, so as not to save over someone else’s hard-won tune file!
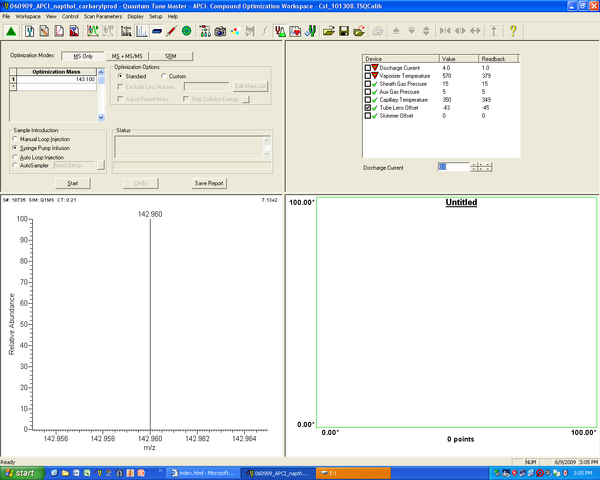
- Choose MS only optimization
- Choose – or + at the big blue “-“ button, here the example shows negative optimization – thus the ions will be negative, with -1 mass. Here the molecule has a mass of 144.2 , so the optimization mass will then be 143.2.
Set the optimization mass based on the -/+ setting
- Start the pump – Go to Setup menu and choose “syringe pump settings”. In the window that opens turn pump to “on” and set a flow rate, e.g. 5 or 10 uL/min. In that window also set the syringe volume, e.g. 500 uL, as appropriate.
- Set up MS device parameters -- Press the “tuning fork/green flask” icon towards the middle (Optimize compound dependent devices) to adjust scan parameters and pick which ones will be optimized. The define scan section will open in the upper right corner (as shown in the figure). Click in the check boxes to the left of each parameter to allow the automatic optimization to adjust that value – e.g. above the tube lens offset will be optimized.
- Define the scan – press the “multicolored bubbles” (Define Scan) button. For the simplest case choose Full Scan in centroid mode (the vertical bars button), and do a scan from 50 to 500 Daltons.
- Start the optimization – Press start in the upper left section. The program takes over, automatically optimizing the given parameters to maximize signal – this is shown in the lower-right hand corner. When it is done it will offer to save optimization-scanning info as a pdf (save it or cancel). If all has gone well, the spectrum in the lower-left corner now shows a big peak for your compound. Save the tune file (*.tsq) and you’re done!
- After doing some tuning with direct injection be sure to switch to inline injection and optimize parameters there – take special note of the gas flow rates, which need to be higher for the inline injection (because the liquid flow rate is higher).
- Stop the MS running by clicking the green triangle in upper left hand corner. Quit the Quantum Tune Program.
- Non-Vanilla Tuning
- Hand optimization – if the tuning is done and is showing you a peak, but you want it to be stronger, turn on the optimize compound dependent devices window. Start the trajectory view (icon with a watch band and a big green peak) to show total current (TIC) as a function of time to allow you to keep track of whether signal is improving. Now adjust each parameter, e.g. capillary temperature by clicking on it and editing in the space below; some parameters may adjust slowly, such as temperature, so keep track of the “Readback” value. The photocopies over the monitor give guidelines for these parameters – e.g. 400 to 600 for Vaporizer temperature in APCI, etc.
- Change the scan type – press the Define Scan (bubbles) button. Choose SIM to focus on one region of mass rather than recording a full spectrum. Set the center and the width of that mass region, e.g. 142.9 and 0.01 to define a very narrow band around a mass. This will increase sensitivity by integrating for a much longer time at each mass.
- Change the MS mode – choose MS/MS in the Optimization Mode region. Check “adjust parent mass” and “step collision energy” boxes. Then click start to start optimization. First a normal ms optimization will be done (and the parent mass adjusted to the best peak). Then the collision energy will be adjusted until a maximum is found – the data are shown in the lower right hand corner. Expect a gradual peak as a function of collision energy with a well-defined maximum. If the optimization fails, however, there is no peak as a function of collision energy and the program reports a failure to find fragments. The compound is rock-solid and won’t be amenable to MS/MS.
- Hand optimization – if the tuning is done and is showing you a peak, but you want it to be stronger, turn on the optimize compound dependent devices window. Start the trajectory view (icon with a watch band and a big green peak) to show total current (TIC) as a function of time to allow you to keep track of whether signal is improving. Now adjust each parameter, e.g. capillary temperature by clicking on it and editing in the space below; some parameters may adjust slowly, such as temperature, so keep track of the “Readback” value. The photocopies over the monitor give guidelines for these parameters – e.g. 400 to 600 for Vaporizer temperature in APCI, etc.