Koch Lab:Publications/Drafts/Versatile Feedback/Paper
Link to Nature Precedings Version
Title
Versatile Control System for Automated Single-Molecule Optical Tweezers Investigations
Authors
Richard C. Yeh and Steven J. Koch
Abstract
We present a versatile control system to automate single-molecule biophysics experiments. This method combines low-level controls into various functional, user-configurable modules, which can be scripted in a rudimentary instruction language. The ease with which the high-level parameters can be changed accelerates the development of a durable experiment for the perishable single-molecule samples. Once the experimental parameters are tuned, the control system can be used to repeatedly manipulate other single molecules in the same way, which is necessary to accumulate the statistics needed to report results from single-molecule studies. This system has been implemented for an optical tweezers instrument for single-molecule manipulations, with real-time point-by-point feedback at a loop rate of 10-20 kHz.
Introduction
We wrote a software tool to facilitate and automate feedback control of an optical trap for dynamic single-molecule tethered-bead studies. Single-molecule experiments offer the potential to study the properties and behavior of the enzymes and other molecules that perform the chemistry of life, with a precision unavailable from bulk experiments. Measurements made with optical traps, using optical beads as force transducers (Svoboda and Block, 1994), have revealed clues about the mechanisms of motor proteins (Block; Yanagida) and the energies of biologically-functional substructures (Koch; Brower-Toland).
Our program is important because single-molecule experiments are notoriously hard to perform: the biological samples require hours of delicate preparation and have lifetimes on the order of seconds or minutes; the experimental apparatuses are sensitive to noise and require exquisite stability (Lang et al., 2002). The expense associated with building a laboratory to perform single-molecule studies motivates the creation of tools versatile enough to perform a variety of experiments as appropriate for shared instruments. To these requirements, our software enables the rapid design of a sequence of manipulation steps and the rapid tuning of relevant parameters governing the manipulations, to minimize the number of precious biological samples spent in the design phase of an experiment. Once the appropriate parameters are found, our software enables the precise repetition of any desired manipulation during the high-volume data collection phase of the experiment: scientific results of single-molecule studies are always statistical in nature.
This paper is organized in the following manner: Section 2 describes the design of the program. Section 3 present examples demonstrating the versatility of the program. Section 4 contains discussions and a conclusion.
Program design
- Overall structure and parameters to specify
- Modules
- General structure and parameters to specify; data acquired; performance
- Feedback modes
- Exit conditions
Our program behaves like an interpreter. The user may specify any number of steps to be performed. A flowchart of the data acquisition and feedback control side appears in Figure 1. The main program initializes and configures the data acquisition and optical trapping hardware per the user’s specifications, and sequentially executes each step. Each step consists of a module responsible for taking data, calculating a response (if necessary), controlling the apparatus subsystems, and deciding whether to loop (continue executing the step) or return to the main program. This last responsibility is the major contribution of this paper: rather than simply executing a sequence of steps, the system must programmatically determine when to go to the next step. This is akin to allowing a cook to boil pasta until it has a particular texture, instead of simply boiling pasta, or boiling pasta for a number of minutes. This process of specifying “stop conditions” is described in more detail below. After each step, the main program records the data acquired in the step and metadata about the program state, including the reason why each module exited, and proceeds to the next step if one exists.
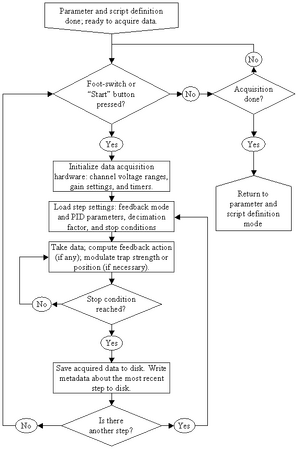
The metadata and acquired data are saved in the “header” and “data” files, respectively. Each header file is a simple free-form database saved in the National Instruments LabVIEW configuration file (similar to the Microsoft Windows for Workgroups initialization files < http://www.microsoft.com/technet/archive/wfw/2_ch6.mspx >) text format, and itself stores information about the data file format. Our data acquisition program saves data in a binary format. Our data processing and analysis programs produce daughter copies of the header and data (converted to text) files, automatically appending the applied calibration data and conversion methods to each newly-generated header file, so that every processed data file has its own detailed history of not only the manipulations used to obtain the raw data, but also the steps used to convert the raw data to its current form. A detailed listing of the information stored in every header file appears at the end of this article.
This program abstracts and combines the low-level manipulations of AOD Voltage (optical trap stiffness) and piezo stage position (sample position) into the most popular modes of feedback control: constant-velocity clamp (with stiffness modulation), often used in stretching studies; constant-force clamp (with position modulation), often used to monitor, hinder, or encourage the progress of motor proteins. Aside from those two modules, this program offers steps to locate the center of the tethered bead (for both long and short tethers); perform velocity and force clamping by steering the beam instead of moving the stage; perform force loading-rate clamping; hold the trap stiffness and position (no feedback) and take data; ramp stage position (no feedback) and take data; acquire a power-spectrum; await the footswitch; reset the acousto-optic deflector driver. These modular feedback programs are configured with a dialog box shown in Figure 2.
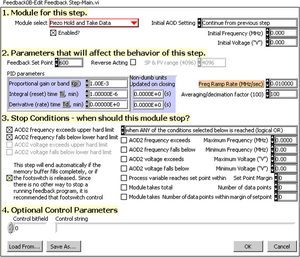
All settings are expressed in hardware units, because at the time that this program was developed, no precise calibration data were available. It would certainly be more convenient to say, “pull the tether with constant velocity until the force exceeds 60 pN,” than “Velocity Clamp with a particular feedback set point (corresponding to a calculated distance from the bead to the trap center) until the AOD voltage exceeds 4.0 V,” but the latter does not depend on (potentially erroneous) calibration data. In the initial design of the program, the optical trapping laser was steered with an acousto-optic deflector (AOD). The frequency of the signal driving the AOD set the position of the trap, and the amplitude determined the trap stiffness. Later, we used a piezo stage to position the sample relative to the trap, and then the AOD frequency settings were converted internally to intended positions and then to piezo voltages.
In section 1 of this dialog box, the user must select the module for this step from a menu of available modules. The “Enabled?” checkbox allows individual steps to be included or excluded from the script, akin to uncommenting or commenting lines in text programs. The “Initial AOD Setting” menu allows the optical trap position and intensity to be reset upon entry into this step. In section 2, standard PID feedback parameters are specified, if applicable to this module. Feedback is performed on the position of the bead relative to the optical trap, so the set point defines a desired displacement of the bead within the tweezer’s Hooke’s-law potential well. The “SP & PV range” field was disabled after we discovered how to obtain the bit-resolution of the data acquisition board programmatically. The “Freq Ramp Rate” field applies to the velocity clamp and other modules that move the trap position at a constant rate. The “Averaging/decimation factor” allows the user to specify the number of point-by-point acquisitions to be averaged (in a boxcar fashion) for each stored point. Section 3 of the dialog box allows the specification of the conditions that will cause this step to terminate. The interpretation of each condition is shown in Table 1. Section 4 of the dialog box provides a modicum of extensibility by allowing custom parameters to be passed to modules. The “Load From” and “Save As” buttons allow the step configuration to be set from or saved to a text file, in the same format that they are saved when the data are acquired.
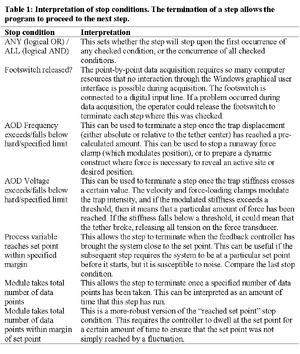
Within each module, point-by-point data acquisition and feedback is performed at rates of 10-20 kHz (our computer was a Dell Pentium 4 running Windows 98 Second Edition). The stop conditions are checked only against the averaged/decimated data. Additional modules may be developed and inserted as needed. For example, to find the center position of a tethered particle in a static fluid, a force clamp can be used to pull the bead to the left until the set point is reached, and then to the right with the same stop condition. A plot of the force exerted by the trap during this process appears in Figure 3 and the point of symmetry is usually defined as the closest approach to the tethering position. (Yeh, 2002)
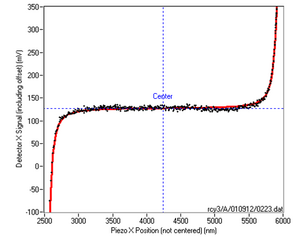
The metadata stored for each step includes: the stop condition or conditions causing the step to terminate; number of data points acquired; the instant position and stiffness of the trap (expressed in hardware units); the average point-by-point loop time (in microseconds); the measured detector offset voltage; the calculated tether center position; and the value of a timing register (used to calculate the precise delay between steps incurred for storing data to disk).
Our system’s primitive step-by-step instruction language does not allow for looping or branching except within the instruction modules themselves.
Examples
The reliability and flexibility of our system is demonstrated by the quantity and variety of experiments for which it has been used to take data (see, for example, Adelman et al., 2002, 2004; Brower-Toland et al., 2002, 2005; Johnson, unpublished calibration data; Koch et al., 2002, 2003; Shundrovsky, unpublished calibration data; Shundrovsky et al., 2004; and Yeh, 2002). In each of the following examples, a diagram depicts a cartoon of the dynamic experiment and the accompanying figure shows a plot of trapping force and trap position versus time, with arrows indicating transitions from one module to the next.
Velocity clamp for DNA stretching
[To be written.]
The script used to take these data has the following steps:
0. (Assume that the tethered bead is centered at the trap position.) 1. Initialize trap stiffness and position. 2. Find tether center. 3. Clamp the bead at a particular displacement from trap center while moving the trap at a constant velocity, increasing trap stiffness if necessary, until the footswitch is released.
Force clamp for RNAP / helicase experiments
Transcription experiments with RNA polymerase reveal how rates of transcription and pause/arrest probability depend on tension applied to the DNA sequence or RNA transcript molecules. The progress of transcription is shown in Figure __ as a change in the force-feedback controlled trap position as the RNA polymerase enzyme draws in or releases the sequence.
The script used to take these data has the following steps:
0. (Assume that the tethered bead is centered at the trap position.) 1. Initialize trap stiffness and position. 2. Find tether center. 3. Clamp the bead at a particular displacement from trap center while keeping the trap stiffness constant, moving the trap if necessary, until the footswitch is released.
Force clamp for nucleosome unwinding experiments
Chemical bonds under constant tension will eventually break. The failure times follow a distribution with the most-likely value dependent on the bond strength and the amount of tension. To acquire good experimental timing data on such events requires high temporal resolution (kHz) when the events occur frequently (tenths of a second). The data need not be acquired at the same high rate in the latter part of a stretching experiment, when events occur less often. To reduce the overall size of the data file while preserving the high-resolution data, we programmed a succession of force-clamp steps with identical parameters but increasing levels of averaging or decimation. Since our program understands not to reset the internal feedback registers between successive force-clamping steps, the transition from one step to the next occurs without disturbing the system, as shown in Figure __.
The script used to take these data has the following steps:
0. (Assume that the tethered bead is centered at the trap position.) 1. Initialize trap stiffness and position. 2. Find tether center. 3. Clamp the bead at a particular displacement from trap center while keeping the trap stiffness constant, moving the trap if necessary, for 10000 points (about 1 second) or until the footswitch is released. 4. Same as previous step, with decimation set to 10. 5. Same as previous step, with decimation set to 100.
Force-loading clamp
[To be written.]
The script used to take these data has the following steps:
0. (Assume that the tethered bead is centered at the trap position.) 1. Initialize trap stiffness and position. 2. Find tether center. 3. Clamp the bead at a particular displacement from trap center while moving the trap at a constant velocity, until a specific force (needed to open the DNA construct) is reached. By now, the construct is open. 4. Clamp the bead with a constantly increasing force (assuming a spring-force potential from the trap center) while modulating both the trap stiffness and the trap position, until the footswitch is released.
Discussions and Conclusion
From a system-design point of view, we can imagine several different use-cases or levels for controlling a small experimental setup, spanning: (1) direct physical or electronic manipulation of individual setup components; (2) computer-aided manipulation of individual setup components; (3) computer control of the entire system. In our instrument, there was a combination of levels always available: switches, safety lockouts, beam-steering telescopes, and microscope stage translators at level 1; and a control panel for adjusting trap intensity and position at level 2. Our software is intended for use-case 3 and overrides any instant setting of the level-2 controls, but cannot affect any level-1 controls.
The implementation by Lang et al. (2002) includes a joystick for use-case 2 to facilitate sample positioning before each experiment, and mostly runs at level 3. Their two-dimensional force clamp eliminates the tether center position error in our one-dimensional system (Yeh, 2002). The implementation by Jobin et al. (2005) includes a haptic device for use-cases 2 and 3, and can record and repeat the manipulations transmitted from the haptic device. This is particularly important for an atomic force microscope, but with the optical microscopes used with optical tweezers, level-1 manual positioning of the microscope stage can be done with a accuracy of 200 nm (Wang, 1995; Yeh, 2002), and video microscopy techniques offer much higher positioning accuracy. Further, every haptic manipulation device will be limited by the operator’s training. While simple modes of force and position feedback have obvious physical analogies, such as holding a constant weight against gravity, or moving an object with constant velocity, more-sophisticated manipulations, such as constant-jerk or force loading-rate feedback over many orders of magnitude (Koch and Wang, 2003), would be challenging for our clumsy fingers to manage.
Millett (1976) notes that software offers a degree of versatility for lab automation that cannot be matched by hardware implementations of feedback control, such as that described by Wang et al. (1995). Our system was motivated by the need for a control system that was comprehensive enough to change any parameter in our experimental setup and user-friendly enough to enable non-programmers to develop and run experiments. Before we finished the initial version of our system in December 2000, control systems in use in our lab were custom-designed for particular experiments. This limited the possible complexity of the experimental parameters and increased the opportunity for errors when adapting the systems for different experiments. Acquired data were not automatically traceable, especially when the control programs changed. Tweaking parameters or inserting additional control steps could not be done “on-the-fly” while samples remained viable.
Our program goes a step beyond that described by Cautero et al. (1994) by enabling the experimenter to specify not only any sequence of manipulations or feedback modes to be applied, but also the conditions to be met for the program to proceed to each subsequent step. This high-level instrument control provides a solution at use-case 3 enabling the reproduction of experimental conditions, the traceability of trapping data to the experimental parameters, and also a safe amount of tinkering and tuning of parameters and feedback sequences to suit a variety of single-molecule experiments.
It requires approximately five hours to train a single researcher to use our instrument, and probably one or two weeks for an experienced LabVIEW programmer and biophysicist to develop a new module. This software was originally developed with LabVIEW 6, but can be modified to run on LabVIEW 5.1.1 and 7.
What design principles can be extracted from this “singular solution for a particular application in a particular environment?” (Millett, 1976). The longevity of our system has been influenced by several factors:
- The program is versatile: it can be scripted to run all previously-known one-dimensional dynamic experiments.
- The program is comprehensive: it allows programmatic control of all currently-known experimental parameters.
- The program is extensible: new modules providing new functionality or feedback modes can be added. Additional configuration parameters can be specified.
- The program produces traceable data in a human- and computer-readable form: every data set is accompanied by a text file containing all script steps and all parameters used to obtain those data, as well as status indications and statistics about the program performance.
- The underlying hardware, with nanometer-scale position resolution and piconewton-scale force resolution, has not changed. If the hardware were to change, the main program would have to be rewritten.
- The preferred use case for which the program was written has not changed. The abstraction of the hardware controls into feedback modes is an appropriate level of description for scientists specialized in fields other than instrumentation.
Appendices
Acknowledgements
The authors thank Alla Shundrovsky, Arthur La Porta, and Benjamin E. Newton for developing or improving some key subroutines used in the program. We are grateful to Alla Shundrovsky, Brent D. Brower-Toland, Karen Adelman, and Daniel S. Johnson for use-testing the software and providing helpful comments to guide the functionality and user interface design. RCY and SJK were supported on NIH Molecular Biophysics Training Grant T32-GM08267 and other grants from the NIH and the Keck Foundation. SJK also supported by a grant from the US Dept. Ed. This work was performed in the laboratory of Michelle D. Wang at Cornell University.
References
References
Adelman K, La Porta A, Santangelo TJ, Lis JT, Roberts JW, Wang MD. (2002) “Single molecule analysis of RNA polymerase elongation reveals uniform kinetic behavior.” Proc Natl Acad Sci U S A. 99:13538-13543.
Adelman K, Yuzenkova J, La Porta A, Zenkin N, Lee J, Lis JT, Borukhov S, Wang MD, Severinov K. (2004) “Molecular mechanism of transcription inhibition by peptide antibiotic Microcin J25.” Mol Cell. 14:753-762.
Brower-Toland BD, Smith CL, Yeh RC, Lis JT, Peterson CL, Wang MD. (2002) “Mechanical disruption of individual nucleosomes reveals a reversible multistage release of DNA.” Proc Natl Acad Sci U S A. 99:1960-1965.
Brower-Toland B, Wacker DA, Fulbright RM, Lis JT, Kraus WL, Wang MD. (2005) “Specific contributions of histone tails and their acetylation to the mechanical stability of nucleosomes.” J Mol Biol. 346:135-146.
G Cautero, G Paolucci, B Brena, R G Agostino, R Tommasini, G Comelli and R Rosei. (1994) “A LabVIEW-based control system for a surface science experimental station.” Meas. Sci. Technol. 5:1002-1011.
Marc Jobin, Raphael Foschia, Sébastien Grange, Charles Baur, Gérard Gremaud, Kyumin Lee, Laszlo Forró, and Andrzej Kulik. (2005) “Versatile force–feedback manipulator for nanotechnology applications.” Rev. Sci. Instrum. 76:053701.
Koch SJ and Wang MD. (2003) “Dynamic Force Spectroscopy of Protein-DNA Interactions by Unzipping DNA.” Phys. Rev. Lett. 91:028103.
Koch SJ, Shundrovsky A, Jantzen BC, Wang MD. (2002) “Probing protein-DNA interactions by unzipping a single DNA double helix.” Biophys J. 83:1098-1105.
Matthew J. Lang, Charles L. Asbury, Joshua W. Shaevitz, and Steven M. Block. (2002) “An Automated Two-Dimensional Optical Force Clamp for Single Molecule Studies.” Biophys. J. 83:491-501.
Millett EJ. (1976) “Digital techniques in laboratory automation.” J. Phys. E: Sci. Instrum. 9:794-802.
Shundrovsky A, Santangelo TJ, Roberts JW, Wang MD. (2004) “A single-molecule technique to study sequence-dependent transcription pausing.” Biophys J. 87:3945-3953.
Svoboda K, Block SM. (1994) “Biological applications of optical forces.” Annu Rev Biophys Biomol Struct. 23:247-285
Wang MD, Yin H, Landick R, Gelles J, Block SM. (1997) “Stretching DNA with optical tweezers.” Biophys J. 72:1335-1346.
Yeh RC. (2002) “Design and calibration of optical tweezers for single-molecule studies.” Master’s thesis, Cornell University. Available at http://hdl.handle.net/1813/3611.