Julius B. Lucks/Meetings and Notes/01212008 Arkin
From OpenWetWare
Jump to navigationJump to search
ColE1 System - Engineerable transcriptional control
- idea - want to genericise transcriptional control
- currently, theoretically, if want to build more complicated genetic networks, need more parts (promoters, terminators, etc.)
- problems - each have their own kinetics
- not necessarily orthogonal
- to build something up, have to do massive characterization, then put together, then tweak to get to work together
- parts proliferation problem
- every gate in the system has to be made up of a different non-interacting promoter, etc.
- still need to be tuned
- one approach is the big FAB approach (Knight, Endy) -
- build massive libraries of promoters and characterize them all in different cell contexts
- then in multiple RBS contexts
- then in multiple terminator contexts
- might have enough characterization then to be able to build something up
- HUGE cost, and not sure will work
- probably something like will first do on 10 most popular things, then stop there
- want to make the building of complicated system more amenable to predictable design
- another approach via RNA engineering - translational control
- Smolke - ribozymes - recognize metabolite - change conformation - allow translation
- Benenson - RNAi governing transcript stability
- these have been working in eukaryotes - mostly metazoans actually
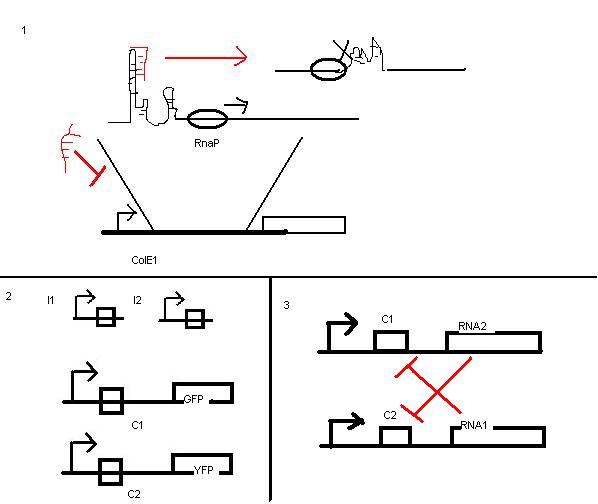
ColE1 - RNA control of transcription via antisense interaction
- what if we could use antisense interaction (like RNAi) to control transcritption
- see image to understand the system (1)
- ColE1 (72 bp region) - high copy number replication origin in E. coli (we actually use this in the lab)
- when transcribed, forms some complicated secondary structure that allows polymerase to carry on, so ON by default
- if certain antisense piece of RNA added, will bind to this, causing a DIFFERENT RNA secondary structure down the line, which does not allow polymerase to pass
- natural NOT gate
- true antisense mechanism - does not require other proteins to work
- looks designable
- could change the ColE1 sequence to recognize different key anti-sense sequences
- hopefully be able to do this with simple anti-sense matching
- hopefully be able to keep GC-content the same so that the thermodynamics would not change too much
- want many different (so orthogonal) ColE1 regions so can put them together in different ways
Initial Targets
- recreate original experiments
- try to find a sequence with a stronger repression factor
- prove orthogonal (2)
- put 2 orthogonal ColE1s behind two genes - 2 diff inputs, make sure only one on for each input
- reconstruct collins switch (3)
- get one system to produce anti-sense of the other
- to get this to work, would have to make sure the system is cooperative
- would need to measure the induction curve - if sigmoidal, see what can do with it
Applications
- since this is RNA mediated, doesn't matter where the RNA comes from
- could come from cancer cells (which are known to over-express certain RNAs)
- would have to design ColE1s to recognize these specific sequences
- the best papers make a new type of part - more powerful - more computation power - some application for these cells
- RNAi logic, Molecular turing machines - Kobi Benenson
- lots of power
- beyond Adelman
- more towards what I want to do
- figure out need computational power X to do thing Y - put in an application context
Problems
- currently only about a 3-fold repression - not that much
- may not be an anti-sense thing - could be something more complicated (Chris Anderson seemed to indicate)
- could be that the dynamic range is poor
- kinetics could be poor
- could be good for a NOT gate, but might need more logic on the promoter (like promoters that have several inputs - do an integration right there)
TOREAD
- talk to Anthony Carruthers (looking at changing mRNA degradation rates)
- also Jonathan Golder
- also David Tulga - works w/ Anderson - Xis and Int recombination systems
- alos Chris Anderson about RNA genes
- big ColE1 people in europe
- Sabin Brandtl
- Gerhard Wagner
- which papers like and why - which ones are the most exciting
Random Notes
- other systems kind of like this
- SacT terminator of B. subtilis - sacT binds to hairpin and opens
- anti-termination with no modifications of the polymerase
- SacT terminator of B. subtilis - sacT binds to hairpin and opens
- iGEM - FMN - translational control
- were trying to avoid the protein part - just another step to worry abou engineering
- Niles Pierce - multi-stranded RNA systems