Jessica Karen Wong/Notebook/2007-6-22
From OpenWetWare
Jump to navigationJump to search
To Do
- Run gel of T9002 and E0240 PCR
- PCR Cleanup of T & E
- Digest T & E with Mfe1 and Nsi1
- PCR cleanup B, J, and R
- Ligate and transform B, J, R
- Log outline, schedule, and parts
Digest Cont'd
- heat shocked digests for J04650, B0032, R0040, and the plasmids
- put in 4 degree
Gel
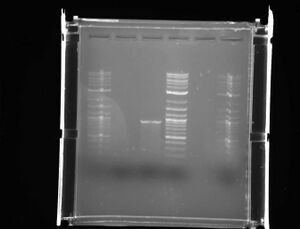
- Ran an analytic gel of the PCR product of T9002 and E0240
- L to R: Ladder, E, T, Ladder, space, Ladder
- E0240 gave no results and the T9002 fragments are 1kb when the desired product should have been 2kb
- T9002 has to B0015 sites in it and we conclude it must have cut only at the first site
- This also explains the absence of E0240 b/c they have the same end sequences
- Ordered new reverse, E0240F, and T9002F primers
Designing Primers
- New Reverse Primer (both T9002 and E0240) TCAGCCAT ATGCAT AAAACGAAAGGCCCAGTCTTTCGACTGAGCCTTTCGTTTTATTTGATGCCTGGCTCTAGTATTATTATTTG
- Melting temp without tail is 81.5
- Is so long because it must go all the way back to some of the GFP
New forward E0240 CTTAGTAG CAATTG TCACACAGGAAAGTACTAGATGCGTAAAGGAGAAGAACTTTTCACTGGAGTTGTCCCAATTCTTGTTGAATTAG
- Melting temp without tail is 80.9
- Length is to have similar melting temp as reverse
New forward T9002 CTTAGTAG CAATTG TCCCTATCAGTGATAGAGATTGACATCCCTATCAGTGATAGAGATACTGAGCACTACTAGAGAAAGAGGAGAAATACTAGATG
- Melting temp 80.0
Bold is the tail, italics is the restriction site.
PCR Purification
Did a PCR cleanup of the 3 backbones, T9002, E0240, B0032, R0040, and J04650
- Used 50ul of B, R, and J
- Used 90ul of T & E
- Protocol is in QIAquick spin handbook
3-way Ligation
Nishant Ligated:
- R0040, J04650, and 1AT3
- R, J, and 1AC3
- B0032, J04650, and 1AT3
- B, J, 1AC3
Overnights
- Checked competent cells by transforming with pUC18 and plating on both plain LB and Amp
- Set up 2 5ml overnight cultures of each plasmid (1AT3, 1AC3, 1AK3, and 3K3)
- 1 to miniprep and 1 to make glycerol stocks