James C. Clements: Week 8
Answer the following Discovery Questions from Chapter 4
5. (p. 110) Choose two genes from Figure 4.6 (PDF of figures on MyLMUConnect) and draw a graph to represent the change in transcription over time. *Note: Dr. Dahlquist said that this will be done on a seperate piece of paper to be submitted in class on Thursday.
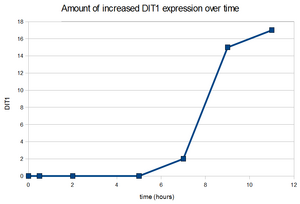
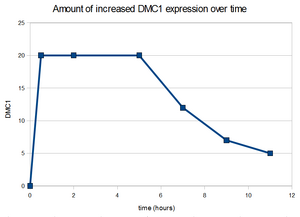
6b. (p. 110) Look at Figure 4.7, which depicts the loss of oxygen over time and the transcriptional response of three genes. These data are the ratios of transcription for genes X, Y, and Z during the depletion of oxygen. Using the color scale from Figure 4.6 (bright, medium, dim green, black, dim, medium, or bright red), determine the color for each ratio in Figure 4.7b.
| 1 | 3 | 5 | 9 | |
| gene X | black | med. red | black | med. green |
| gene Y | black | med. red | dim green | bright green |
| gene Z | black | dim red | med.-dim red | med.-dim red |
7. (p. 110) Were any of the genes in Figure 4.7b transcribed similarly?
Yes, genes X and Y were transcribed similarly.
9. (p. 118) Why would most spots be yellow at the first time point?
Equal amounts of cDNA from both the control and the experiment are present.
10. (p. 118) Go to http://www.yeastgenome.org and search for the gene TEF4; you will see it is involved in translation. Look at the time point labeled OD 3.7 in Figure 4.12, and find the TEF4 spot. Over the course of this experiment, was TEF4 induced or repressed? Hypothesize why TEF4’s gene regulation was part of the cell’s response to a reduction in available glucose (i.e., the only available food).
TEF4 was repressed. Proteins could not be made in translation perhaps because there was not enough ATP available for synthesis.
11. (p. 120) Why would TCA cycle genes be induced if the glucose supply is running out?
Inducing the TCA cycle genes could cause the flow of carbon towards glucose 6-phosphate which is lacking in glucose limitation.
12. (p. 120) What mechanism could the genome use to ensure genes for enzymes in a common pathway are induced or repressed simultaneously?
The genes in a common pathway could be located near each other and be on the same chromosome.
13. (p. 121) Given rule one on page 109, what color would you see on a DNA chip when cells had their repressor gene TUP1 deleted?
Red because of how many other genes would be induced because of the lack of a repressor gene (the spot for the repressor gene would be red, however).
14. (p. 121) What color spots would you expect to see on the chip when the transcription factor Yap1p is overexpressed?
Red because it induces transcription.
15. (p. 121) Could the loss of a repressor or the overexpression of a transcription factor result in the repression of a particular gene?
Possibly, if the expression of some genes are inversely proportional to each other.
16. (p. 121) What types of control spots would you like to see in this type of experiment? How could you verify that you had truly deleted or overexpressed a particular gene?
I would like to see control spots for cases when the cell clearly needs the gene to survive (in the case of of deleting the gene) and for cases in which the over expression of the gene is harmful for the cell. If the gene does not appear in those situations then the process will have been successful.