IGEM:IMPERIAL/2009/Twitteria
Twitteria - A Novel Bacteria-Computer Interface
Description
We propose the idea of a new interface between biology and computing. This new interface would allow a new standard in synthetic biology for networking between a computer and bacterial culture.
Our idea aims to remove the need to have human monitoring of a culture, and to instead create a standard automated process.
We aim to move away from the manual monitoring and control of experiments, to a more automated system, with user defined commands.
This new standard would result in the creation of a new toolbox of Biobricks, designed to be compatable with a computer. Like the standard Biobrick catalogue, these will be designed to a set of criteria - reusability, modularity, standardisation and now compatability.
The possibilities of such a system are endless. It would allow complete automation of biological experiments, and fine control using the computational processes to monitor feedback loops etc.
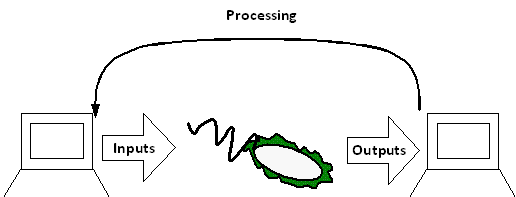
The input mechanism would be anything that is used to stimulate the culture to perform a certain function. For example, using microfluidics to add a certain amount of a transcription factor to stimulate expression of a certain gene.
The output mechanism would vary, allowing measurement of many different paramters of the culture. Possibilities include pH measurement, flourescence (GFP) expression related to gene expression, optical densitiy and colour (possibly used as a null driver for custom inputs). The aim would be to maintain the 'plug and play' functionality of traditional biobricks, whilst adding an increased level of control to the process.
These output mechanisms are designed to be compatible with inputs for the computer. In this way the system can be fully automated, with the computer responding instantly to signals from the cell culture. Programming of the software interface with usability in mind would give the possibility of adapting the settings for whatever inputs and outputs are desired.
The Big Picture: the Toolbox
The proper characterisation of various combinations of interfaces (or tools) would create a powerful suite enabling the biologists to computerize some aspects of data analysis.
Each tool would be composed of two halfs: the biobrick and the computational output as shown below, where BB and CI respectively stand for BioBrick and Computational Input
The biobrick standard is open-source, however in this toolbox, the CI (pH meter, turbidity meters...) are not. We aim to solve this problem via the creation of CIs from scratch.
These would be open-source as they would employ a readily available and immensely popular class of microcontrollers: Arduino (examples of these microcontrollers at the bottom of the page). Also, the kits are quite cheap and could potentially appeal quite strongly to the DIY community
This toolbox would not only comprise of "hardware" solutions but also contain "software" solutions that would link the different detectable variables to the major programming languages.
Applications for this particular last point are virtually limit-less. Ranging from Twittering bacteria (thanks Vincent) to automated email delivery when the bacteria are undergoing stress, potentially enabling the scientist to salvage weeks worth of experiments.
These particular applications have a great visibility factor and we think could be a great selling point for the more serious toolbox suite endgoal.
Arduino web-resources
Arduino official website
Hack A Day website features a number of arduino projects
Make features a number of arduino projects and associated tutorials
Instructables features a number of tutorials on arduino
Check out Vincent's profile page on OWW and scroll down to open source hardware, there are a number of very interesting projects there.
Application 1 - Automated Cell Feeder
This is the simplest example of how we can apply the BIB system. Like a one 1-1 function in maths, this demonstrates interfacing between one cell culture with one processor. Here, pH of the cell broth is monitored. As the cell culture grows, it uses up the nutrient media and changes its pH (or colour). A pH meter (or colourimeter) is used to detect the change. A program is written that responds to this change in pH (or colour) by extracting the expired media and adding in the new media. In this way, the researcher can go on his holiday and not worry about his cells dying. Furthermore, the computer can send a twitter feed or email the researcher that new media has been added.
Application 2 - Information transfer
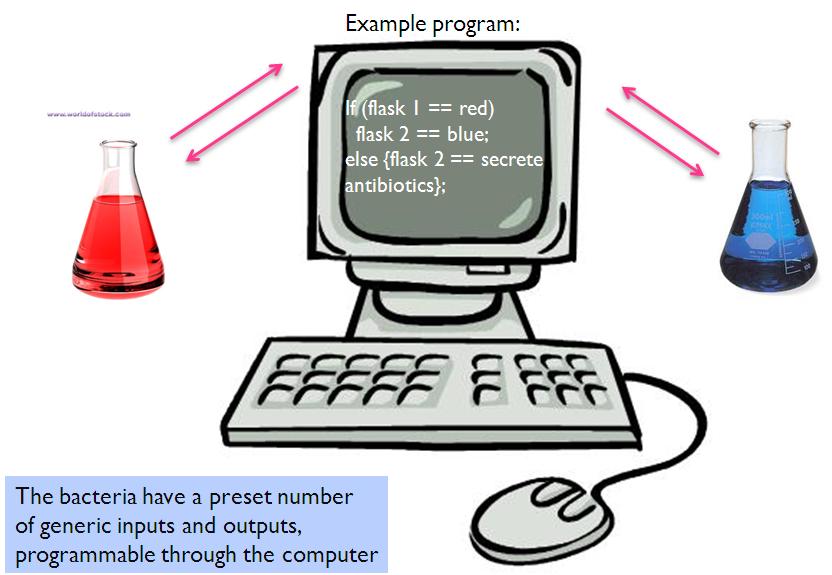
This is an example of a 1-many function in maths. Here, we have one computer interfacing with two (or more) cell cultures. The sample program shows that when the cells in flask 1 glows red under a stimulus, the computer causes say, chemical X to be secreted into flask 2 and the cells in flask 2 will turn blue. If however, the stimulus causes cells in flask 1 to turn green, cells in flask 2 will be made to produce antibiotics. Of course, the cells in both flasks are genetically engineered to respond to certain inputs and output different colours accordingly.
It could be conceivable to create a game based around this application of Information transfer. The popular Battleship game, when analysed more precisely is in reality an inherent exhibition of information transfer. Each player sends coordinates decoded by the adversary and that translate into a hit or miss. We could imagine two bacterial cultures communicating through a computer (or quorum sensing, signal subsequently analysed by the computer, to know what is going on). The first broth would make a first move by secreting a certain amount of molecule that would be detected by the computer. This would then "translate" or decode the information and equate into <....> Scrap that idea.
Example: Bacterial MasterMind Game
Mastermind rules/game
In short, you propose a number of combinations and each time you do, the opponent (whose combination is hidden) answers in code fashion: one white peg for each colour that is correct but not in the right position, one red peg for each colour in the right spot. By doing permutations and thus suggesting combinations (trial and error approach) you should (that is if you haven't been lobotomised prior to playing) work out the combination.
So now, the idea would be to implement that in cultures. I am no longer thinking about broths because it would be less handy for analysing output.
There are billions of ways to integrate this game.
- Easier, but also probably the least interesting is the 1 inducer -> 1 colour confirmation so you would streak a modified E. Coli strain horizontally on a "hidden" plate then cross streak (90°) in 4 different places with 4 inducers that would cause different colours (fluorescent proteins will probably be easiest) to be expressed by the bacteria. The "Guesser" the does the same but streaks the different inducers to "submit" his/hers educated guesses.
- This idea is however only basing itself on selection of FPs under different promoters: boooring!
- More complicated, thus iGEMMABLE: using broths this time. Using randomly injected (via computer) inducers, the combination is determined genetically but does not result in visible change (or it does but the plate is hidden (see bullet point # 3)) ie: protein synthesis, or reporters that are analysed continuously by the computer checking against a threshold to know whether they are expressed thus working out the combination. The guesser broth, would then temporally (not in the case of custom proteins) encode/produce signals (could be colour/fluorescence, could be proteins) that are detected by the computer and analysed against. The following logic operations can either be done by computer (easy) and translated into a biological signal (but not much interest maybe) or the logics performed by a -yet to be specified construct-
work in progresss
Application 3 - Universal Biosensor
This is an example of a many-many function. We can genetically engineer bacteria to respond to various inputs and give corresponding outputs. To detect diabetes, a drop of urine is added to the culture broth. Genetically engineered bacteria with glucose receptors will output GFP. To detect kidney failure, the same bacteria will also be genetically engineered to contain amino acid receptors, which will output RFP. In this way, we can easily detect diseases in humans. Hence, a universal biosensor is produced.
Application 4 - Production Line
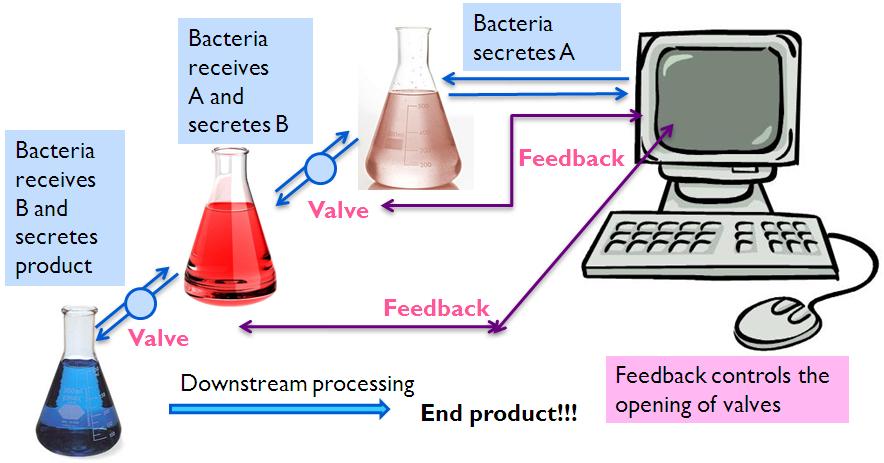
This shows a production line application of the BIB system. The computer is programmed to cause the first cell culture to start secreting A. When the concentration of A has reached the threshold, the probe detects it and the computer causes a quencher to be added into the cell culture to stop the reaction. The valve is open and A is secreted into the second cell culture. The same thing happens till the final cell culture, where our product of interest is secreted. Downstream processing is performed to isolate our product.
Application 5 - CAG
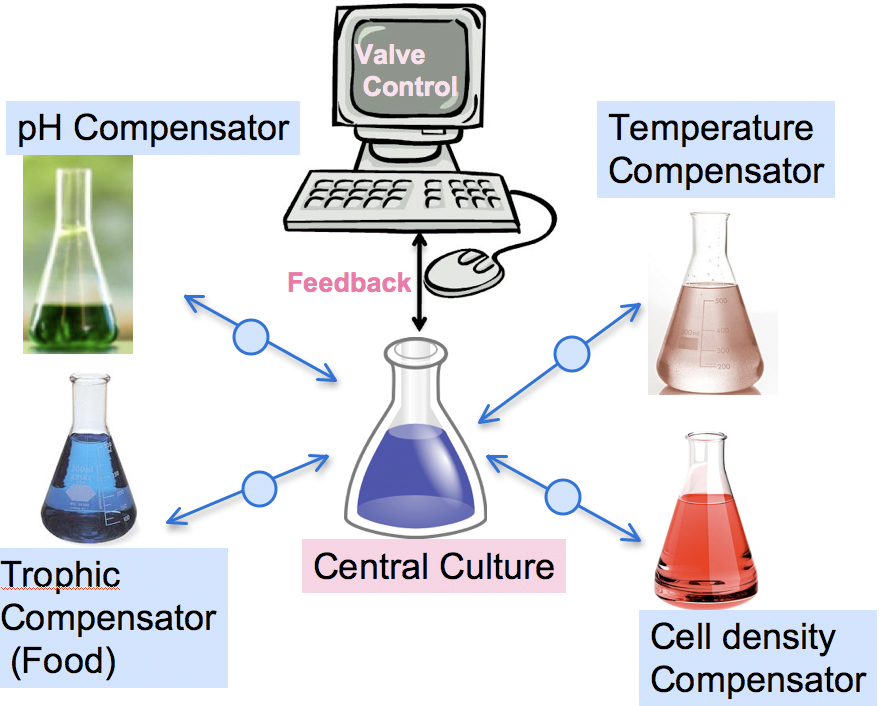

The aim of this project is to implement a supply-demand system to minimize stress incurred by a central culture.
In order to sense stress cues, we propose to use a number of existing biobricks (eg: BBa_K115035, TU Delft08) as well as creating a considerable number of them adapted to detect different signals (eg: Media:II09_EPA.pdf).
The bacterial output modules would allow to monitor environment variables at the molecular level. These sensors would result in the induction of a signal readable by computer assisted sensors.
This system would allow to predict (based on computer data analysis) stressful conditions and consequetly open the valve linking the stress-remediating and the central (futurely stressed) cultures.
Example: Central culture undergoing Temperature stress
The central culture's temperature is going up but the conditions are not stressful yet. A constitutively expressed temperature sensor linked to a luciferase output senses the shift in temperature and signals it through the computer interface.
As a result of temperature threshold crossing, the system starts sending a quorum sensing signal to the other flasks but the computer only operates the relevant valve. This accellerates signalling to induce the peripheral broth to produce the needed compound, in this case a Heat Shock Protein.