IGEM:IMPERIAL/2009/M3/Modelling/old/Codes/celldeath
Function code
function dy=ty_celldeath(t,y)
%%
global T km1 km2 k1 k2 k11 n1 n2 de dm growth death
%%
%equations
activating_hill_temperature =(k1*(T^n1))/(T^n1+km1^n1); %activating
dy(1)= activating_hill_temperature - dm*y(1);
dy(2)= k2*y(1) - de*y(2);
hill_function_death = (k11*(y(2)^n2))/(y(2)^n2+km2^n2); %activating
dy(3)= growth*y(3) - death * hill_function_death * y(3);
dy=[dy(1);dy(2);dy(3)];
Call
%function call
clear all;
clc;
%%
global T km1 km2 k1 k2 k11 n1 n2 de dm death growth E
%T=1
km1 =1;
km2 =1;
k1 =0.5;
k2 =1;
k11 =1;
n1 =1;
n2 =1;
de =1;
dm =1;
growth=1;
E =1; %enzyme
%death=3;
i=1; %%
death=1;
for T=[0:1:4]
[T,Y]=ode45(@ty_celldeath,[0:0.01:10], [0 0 1000]); %initially, has population of 1000
A1(:,i) = Y(:,1); %mRNA conc
A2(:,i) = Y(:,2); %killing enzyme conc
A3(:,i) = Y(:,3); %population
i=i+1;
end
figure(1);subplot(1,3,1);plot(T,A1); TITLE('mRNA');xlabel('time');legend('T=0','T=1','T=2','T=3','T=4');
figure(1);subplot(1,3,2);plot(T,A2); TITLE('Enzyme');xlabel('time');legend('T=0','T=1','T=2','T=3','T=4');
figure(1);subplot(1,3,3);plot(T,A3); TITLE('Cell population');xlabel('time'); legend('T=0','T=1','T=2','T=3','T=4');
%%
death=2;
for T=[0:1:4]
[T,Y]=ode45(@ty_celldeath,[0:0.01:10], [0 0 1000]); %initially, has population of 1000
A12(:,i) = Y(:,1); %mRNA conc
A22(:,i) = Y(:,2); %killing enzyme conc
A32(:,i) = Y(:,3); %population
i=i+1;
end
figure(2);subplot(1,3,1);plot(T,A12); TITLE('mRNA');xlabel('time');legend('T=0','T=1','T=2','T=3','T=4');
figure(2);subplot(1,3,2);plot(T,A22); TITLE('Enzyme');xlabel('time');legend('T=0','T=1','T=2','T=3','T=4');
figure(2);subplot(1,3,3);plot(T,A32); TITLE('Cell population');xlabel('time'); legend('T=0','T=1','T=2','T=3','T=4');
%%
death=3;
for T=[0:0.01:0.04]
[T,Y]=ode45(@ty_celldeath,[0:0.01:10], [0 0 1000]); %initially, has population of 1000
A13(:,i) = Y(:,1); %mRNA conc
A23(:,i) = Y(:,2); %killing enzyme conc
A33(:,i) = Y(:,3); %population
i=i+1;
end
figure(3);subplot(1,3,1);plot(T,A13); TITLE('mRNA');xlabel('time');legend('T=0','T=1','T=2','T=3','T=4');
figure(3);subplot(1,3,2);plot(T,A23); TITLE('Enzyme');xlabel('time');legend('T=0','T=1','T=2','T=3','T=4');
figure(3);subplot(1,3,3);plot(T,A33); TITLE('Cell population');xlabel('time'); legend('T=0','T=1','T=2','T=3','T=4');
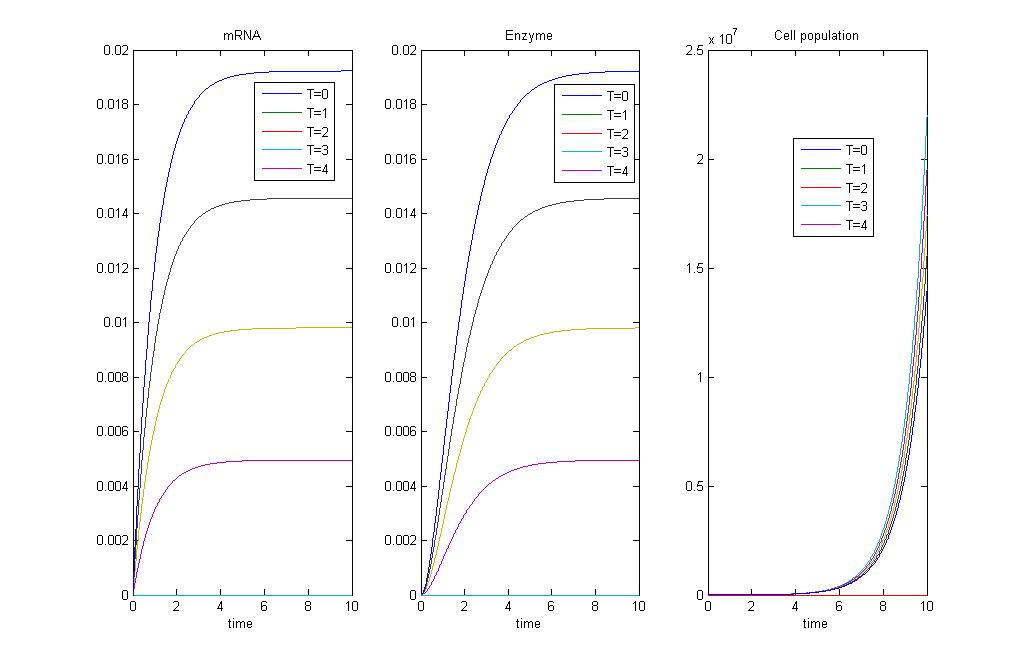
Results
Discussion
We can see from the simulation that as temperatures increase, there'll be an increase in production of enzyme and mRNA of TaqI and DpnII, the restriction enzymes that are involved in cell killing. As we make the death term greater than the growth term, we'll see increasing cell death, with increasing temperatures.