IGEM:IMPERIAL/2009/M3/Assays/BetaGal
Beta-Galactosidase staining (back up in case GFP does not work)
Aims
- We want to show that on coupling cI to the lambda promoter, cI represses the promoter activity.
- By linking F2620 to cI (TetR-LuxR-pLux) and the lambda promoter to the BBa_E0240 testing construct (RBS-GFP-TT), the expression level of cI, and consequently the transcriptional activity of lambda promoter can be determined.
- A quantitative measurement of the fluorescence over time is directly obtained. This will then be converted to the relative transcriptional activity of the promoter (PoPS).
Assay
The synthetic compound o-nitrophenyl-β-D-galactoside (ONPG) is also recognized as a substrate and cleaved to yield galactose and o-nitrophenol which has a yellow color. When ONPG is in excess over the enzyme in a reaction, the production of o-nitrophenol per unit time is proportional to the concentration of β-Galactosidase; thus, the production of yellow color can be used to determine enzyme concentration.
[1]
1 Miller Unit =
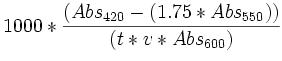 where:
where:
Abs420 is the absorbance of the yellow o-nitrophenol,
Abs550 is the scatter from cell debris, which, when multiplied by 1.75 approximates the scatter observed at 420nm,
t = reaction time in minutes, After sufficient color has developed, add 700 μL of Stop solution, mix well, and NOTE THE STOP TIME.
v = volume of culture assayed in milliliters,
Abs600† reflects cell density.
Equipment
- Spectrophotometer
Reagents
Standard Curve
- LacZ protein
Permeabilization Solution
- 0.8 mg/mL CTAB (hexadecyltrimethylammonium bromide)
- 0.4 mg/mL sodium deoxycholate
- 100 mM dibasic sodium phosphate (Na2HPO4)
- 20 mM KCl
- 2 mM MgSO4
- 5.4 μL/mL beta-mercaptoethanol
Substrate solution
- 1 mg/mL o-nitrophenyl-β-D-Galactoside (ONPG) - Sigma
- 60 mM Na2HPO4
- 40 mM NaH2PO4
- 2.7 μL/mL β-mercaptoethanol
Stop solution
- 1 M Sodium Carbonate (Na2CO3)
Protocol
1) pre-measure 80 μL aliquots of permeabilization solution into 1.5 mL microfuge tubes and close them
2) Grow cells until log phase
3) Measure Abs600 and RECORD IT! (preferably OD600 = 0.28 to 0.70) br>
4) Remove a 20 μL aliquot of the culture and add it to the 80 μL of permeabilization solution. The sample is now stable for several hours. This allows you to perform time-course experiments.
4) Add 600 μL of Substrate solution to each tube and NOTE THE TIME OF ADDITION
5) After sufficient color has developed, add 700 μL of Stop solution, mix well, and NOTE THE STOP TIME
6) Centrifuge briefly.
7) Carefully remove the tubes from the centrifuge and transfer supernatant from the TOP of the tubes to your cuvette(s). You are trying to avoid having particulate material in the cuvette so that scattering will not influence the reading
7) Use a spectrophotometer and record Abs420
(Recommended OD420nm reading should be 0.6-0.9)
Calculate Miller Units as:
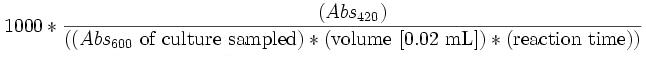
This formula yields approximately 1 Miller Unit for uninduced E. coli (low β-Gal production) and approximately 1000 units for a fully induced culture
Notes:
If the reaction goes too fast, your estimate of reaction time will have a large error. If you need to slow the reaction, you can use fewer cells and increase the amount of permeabilization buffer so the volume is still 100 μL
I start the samples 10 sec apart (with the time counting up) so it is possible to get accurate reaction times even if you only let your reactions go for a few minutes. I get very reproducible results with reaction times of 1.5-30 min. Doing each sample in triplicate will give you some confidence that your timing is reproducible