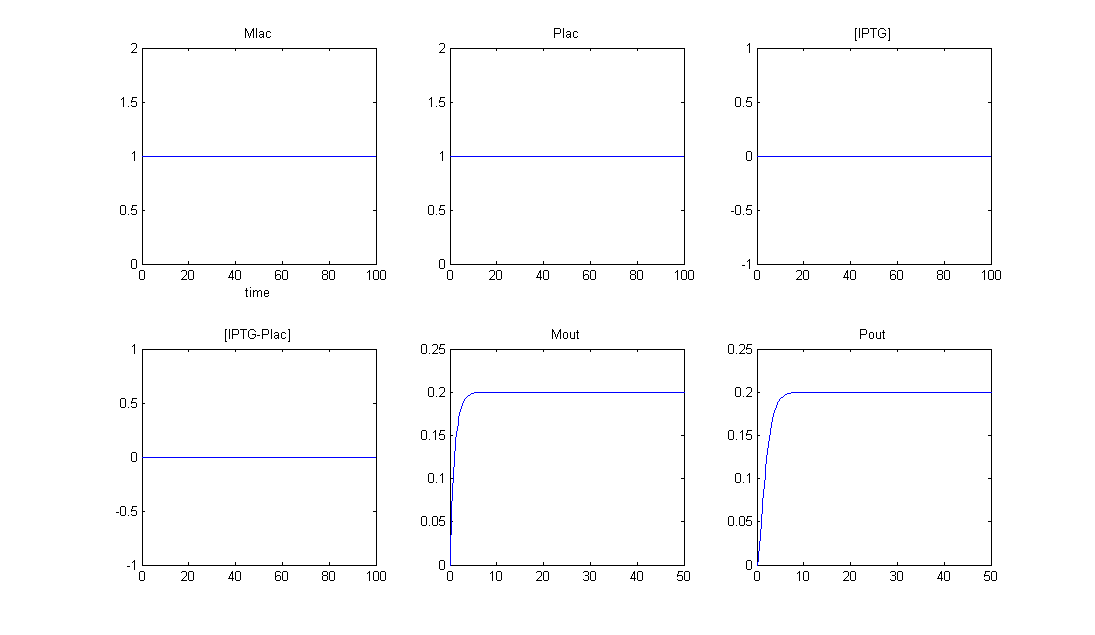IGEM:IMPERIAL/2009/M1/Modelling/LacI-IPTG
Preliminary Model 1: Genetic circuit: Action of the LacI/IPTG repressive pathway
Hypothesis
- LacI is constituitively expressed, and in the absence of IPTG, it represses the production of protein.
- MB's FB: THis is true, but this is not the only assumption made for LAcI...
- When we add in IPTG, we "de-repress" the pathway, as LacI is removed from the system and binds to IPTG to form a complex:
- LacI + IPTG ---> LacI-IPTG
- MB's FB: arrow goes both ways
- Assumption is : one to one binding
- This has the effect of reducing the amount of LacI, hence allowing for an overshoot in protein production.
- The system will then tend to its original state, once we stop adding IPTG
- Different amounts of IPTG will give us different sizes in the overshoot of protein production.
- MB's FB: these are not assumptions, they are predictions
- Assumptions: you are making big assumptions when you write the equations; please list them
Equations
MB's FB: Explain how you derive these equations and what they mean Please avoid M1,P1 etc.. the molecules have names!!!!
Legend
make the meaning of the terms more precise please
Table 1. Meaning of modelling terms
| Term | Variable/Parameter/function? | Meaning/description |
| [MLac] | Variable | mRNA for production of LacI protein |
| kmlac | Parameter | Constant for Mlac production |
| dplac | Parameter | Constant for Plac degradation |
| [Plac] | Variable | LacI protein produced (constitutively expressed) |
| kpl | Parameter | Production rate of lacI protein |
| dpl | Parameter | Degradation rate of lacI protein |
| [IPTG] | Variable | Concentration of IPTG (inducer) |
| kb1 | Parameter | Binding constant for the reaction [IPTG] + [Plac] ---> [IPTG-Plac] |
| kb2 | Parameter | Binding constant for the reaction [IPTG-Plac] ---> [IPTG] + [Plac] |
| [p1-IPTG] | Variable | Intermediate reaction complex that forms itself when IPTG is added to the system |
| [Mout] | Variable | MRNA that makes our protein of interest |
| dmo | Parameter | degradation rate of Mout |
| kmo | Parameter | production rate of Mout |
| [Pout] | Variable | Protein of interest |
| kpo | Parameter | production rate of Pout |
| dpo | Parameter | degradation rate of Pout |
| fh([p1]) | Function | Repressive hill function of LacI, acting on the output protein module |
| Th | Parameter | Switching concentration of Plac hill function |
| n | Parameter | Hill exponent |
Explanation of equations
- MB's FB:
- the meaning should be moved up the previous section
- More explanations needed
- Distinguish case without IPTG - explain its meaning and give simulations
- and with IPTG (start to real experiment) ; give simulations
- Equation(1): Equation describing the rate of transcription of Lac MRNA ([math]\displaystyle{ m_1 }[/math]). Note that at steady state: [math]\displaystyle{ \frac{d[m_1]}{dt}=0 }[/math] , so [math]\displaystyle{ m_1=\frac{k_{m_1}}{d_{m_1}} }[/math]
- Equation(2): Equation describing the rate of translation of Lac Protein ([math]\displaystyle{ p_1 }[/math]). Note that at steady state([math]\displaystyle{ \frac{d[p_1]}{dt}=0 }[/math]):
- In the absence of IPTG, [math]\displaystyle{ p_1=\frac{k_{m_1}*k_{p_1}}{d_{m_1}*d_{p_1}} }[/math], which would appear at a constant level.
- When we add in IPTG, the story is more complicated, so theoretically the equations become: [math]\displaystyle{ p_1=\frac{k_{m_1}*m_1 + k_{b_2}*(p_1 - [IPTG])}{d_{P_1} + k_{b_1}*[IPTG]} }[/math]. [IPTG] concentrations change over time, so this is like adding a "disturbance" in the system, that will effectively remove some LacI protein from the system. (Clearer in simulations)
- Equation(3): Equation describing rate of change of IPTG when added in. IPTG undergoes a secondary set of reactions with LacI when introduced, where the forward reaction destroys it and the backward reaction re-forms it:
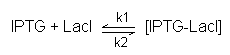
- Equation(4): Equation describing rate of change of LacI-IPTG temporary complex formedwhen IPTG is added in. Same reasoning as Equation (3).
- Equation(5): Equation describing the rate of transcription of [Mout]. The difference with Equation (1) is that before the promoter was constitutive, and now it is inducible. Now, at steady state:[math]\displaystyle{ Mout=\frac{k_{m_o}*\frac{Th^n}{{[p_1]}^n+ Th^n}}{d_{m_o}} }[/math]
- Note that the term [math]\displaystyle{ \frac{Th^n}{{[p_1]}^n+ Th^n} }[/math] is a repressive hill function of [math]\displaystyle{ p_1 }[/math] (LacI protein).Therefore, when [math]\displaystyle{ p_1 }[/math] is at a high concentration, we are repressing transcription of Mout.
- However, the action of removing LacI from the system, if it goes below threshold, we "de-repress" Mout transcription, so more protein is produced during that period of time.
- Equation(6): The system is originally at a steady state, given by: Pout = [math]\displaystyle{ \frac{k_po*Mout}{d_po} }[/math].
- So originally, the system will tend to a particular steady state.
- When we add in IPTG, this will change, so more Pout will be produced.
Simulations
Simulation 1: Outputs for no IPTG
How come Mlac and Plac are constant? Are you simulating the journey to SS? PS: there has been a change of notation
On image, make the plots thicker so they can be seen more easily waht were your parameters+units? (justify)
Explanations
- Mlac is at a steady state (constituitive expression)
- Plac is at steady state (constituitive expression)
- No IPTG in the system
- As a consequence, no intermediate IPTG-LacI complex
- Production of Mout and Pout is repressed, so in the absence of IPTG we only get a basal amount of protein of interest (at a very low level).
- NOTE: If LacI value is at a higher initial state, production is fully repressed.
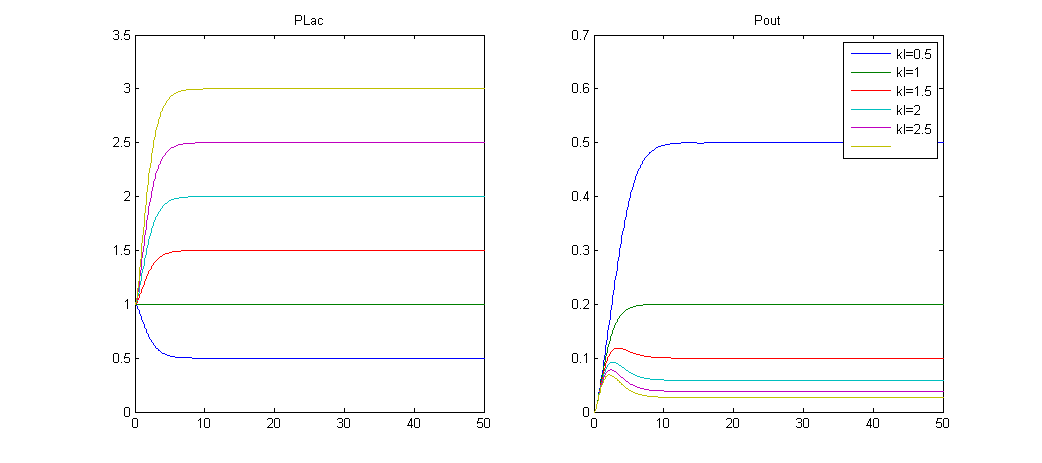
Simulation 2: No IPTG and different levels of LacI, effect on Pout
Explanations before images your simuloations are wrong: Plac does not return to its steady state!!! General commment: how did you choose your parameters? why did you vary the parameter you chose to vary?
Explanations
- The steady state of Plac, and consequently the repression of Pout is given by: [math]\displaystyle{ p_1=\frac{k_{m_1}*k_{p_1}}{d_{m_1}*d_{p_1}} }[/math].
- By varying kp1 (kl on the graph, which is PoPs of LacI protein) the system tends to a different steady state, the higher the value of kp1.
- The higher the value of kp1 at steady state, the more Pout is repressed prior IPTG induction.
- Therefore, we would ideally like levels of LacI expression to be high initially prior IPTG induction, in order to ensure that our production of protein of interest Pout is repressed before we trigger it.
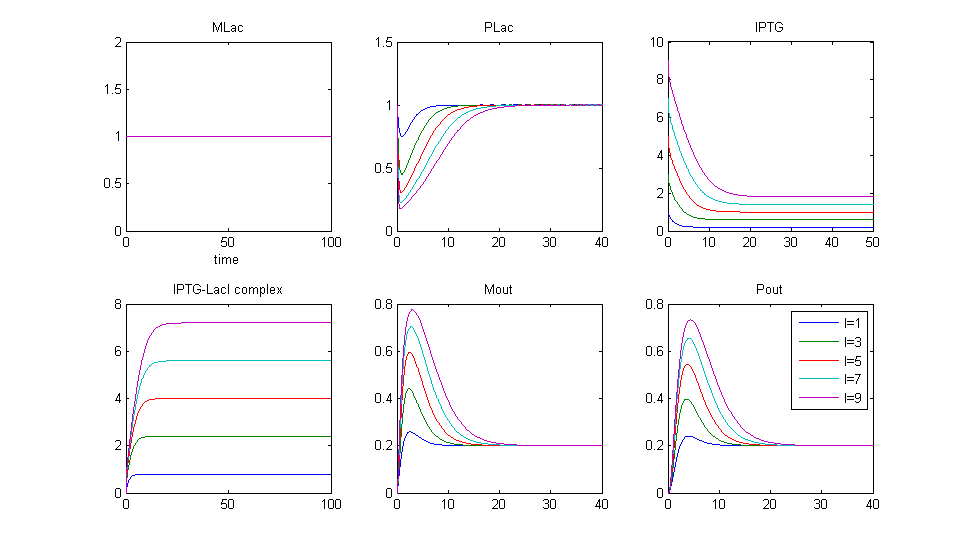
Simulation 3:Protein production for different concentrations of IPTG
Explanations
- At t=0, IPTG is introduced. This has the effect of removing some LacI from the system.
- LacI enters a secondary reaction:
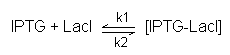
- The more IPTG we introduce, the more LacI is removed from the system so the deeper the "blip"
- IPTG is originally high and is used up over time, allowing LacI levels to return to their original state.
- When LacI is introduced, there is a bump in Pout production. The more IPTG added, the larger the bump.
- NOTE:We can see that there are flaws in this model, as IPTG toxicity has not been taken into account. In reality we will kill cells once a certain amount of IPTG has been added, hence decreasing protein production.
same remarks as above except simulations seem ok
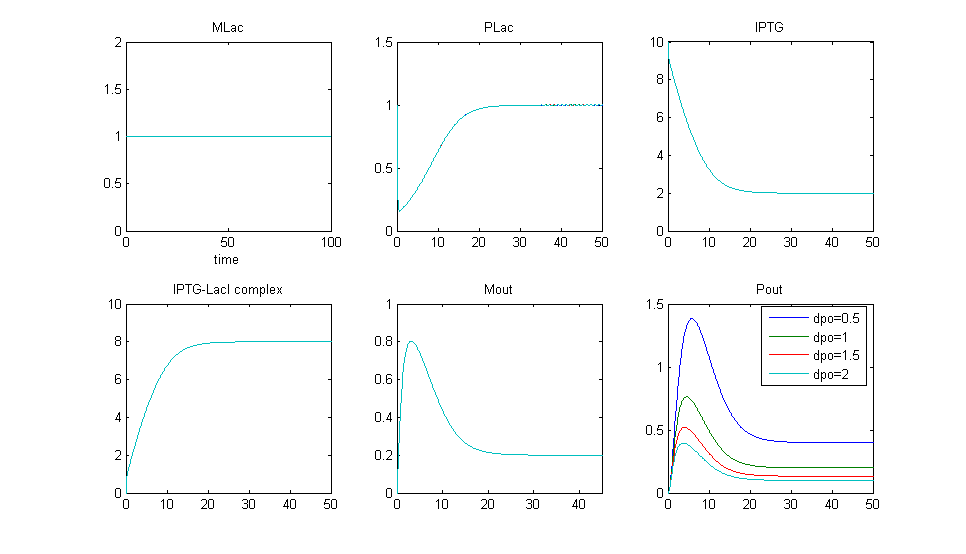
Simulation 4:Variation of Pout degradation term (dpo)
Explanations
- Once again, MlacI is constituitively expressed, and a fixed concentration of IPTG is added in, so PlacI is removed from the system.
- We have varied the degradation rate of Pout.
- We know from equation (6), that the steady state is Pout = [math]\displaystyle{ \frac{k_po*Mout}{d_po} }[/math], where d_po is the degradation term.
- Therefore, if our degradation rate is too large (fast) we will get less Pout produced and also for a shorter time (lower bump).
- Ideally, we would like the Pout enzyme to be stable, but this is not really under our control.
MBs FB: what is thwe purpose of these simulations???
Data Fitting
Remove until needed Create section with your preliminary conclusions
Matlab files
- ODE file
- Calling file
Media:II09_call_m1_circuit.ogg
References
- Zahn K. Overexpression of an mRNA dependent on rare codons inhibits protein synthesis and cell growth. J Bacteriol. 1996 May;178(10):2926-33. DOI:10.1128/jb.178.10.2926-2933.1996 |
-
Effect of Variable Isopropyl Beta-Thiogalactoside Concentrations on �-Galactosidase Activity,Avinash M. Baktula and Genise A. Nolan, Department of Biology, Western Kentucky University