IGEM:IMPERIAL/2009/Encapsulation/Timers
Timers
Timer Prototype 1.1:
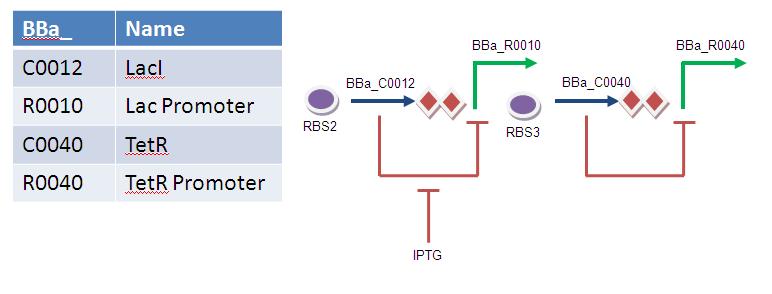
Overview: This timer works on a double repression mechanism. Varying IPTG concentration provides a variable threshold which must be overcome by the accumulation of LacI for the switch to flip. The production of LacI will ultimately be linked to the accumulation of protein.
1) Initial state: LacI=OFF, CI repressor = ON, Output = OFF
2) LacI accumulates until it saturates the IPTG.
3) Once the IPTG is saturated, LacI represses the LacI responsive promoter. This decreases the expression of TetR repressor.
4) A fall in TetR repressor levels results in de-repression of the TetR responsive promoter.
5) Final state: LacI = ON, TetR repressor = OFF, Output = ON
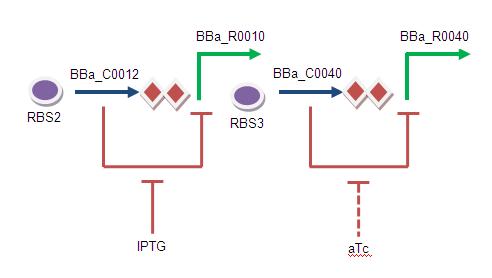
Second Input:
Just as LacI is blocked by IPTG, TetR is blocked by aTc (anhydrotetracycline). aTC inhibitory levels of TetR are about 20 fold lower than the levels to inhibit growth as an antibiotic.
Device Assembly:
1st Attempt:
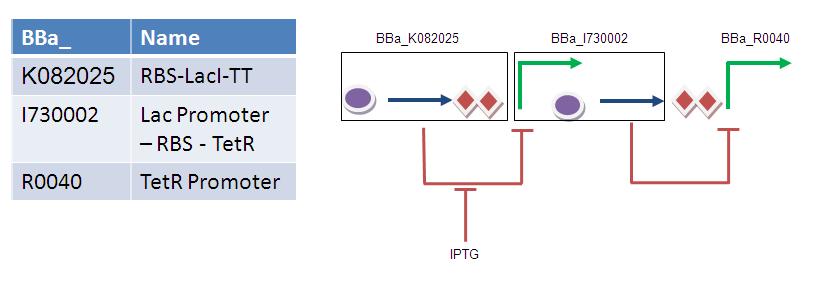
BBa_K082025 consists of an RBS fused to LacI followed by a double terminator. This part works although it does not appear to be present in the 2009 starter kit. Similarly, BBa_I730002 works although it does not appear to be in the start kit. If these two composite parts were used, then assembly of the timer module would consist of three ligation reactions:
1) BBa_K082025 + BBa_I730002
2) BBa_K082025 + BBa_I730002 + TT
3) BBa_K082025 + BBa_I730002 + TT + BBa_R0040
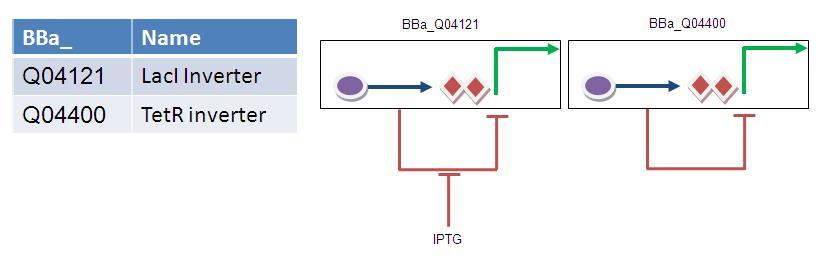
2nd Attempt:
Rather stupidly, I failed to look under the Registry's inverter section. Of course the timer is simply a double inverter... Therefore it can be assembled in one simple ligation step....
Both of these parts are available in the 2009 starter kit, however it should be noted that the LacI inverter has an illegal BglII site at position 1165.
Testing Timer Prototype 1.1:
Testing construct 1:
Overview:
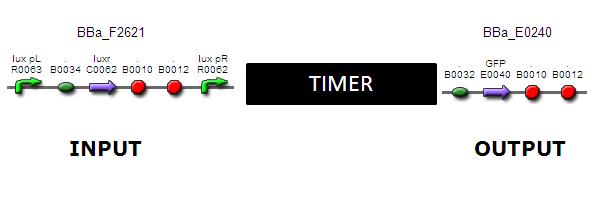
Treat the timer module as a black box. Use a POPs input by placing a characterised promoter upstream of RBS2. Obtain a system output by placing a GFP reporter down stream of the TetR promoter.
Input:
To characterise the timer, it must be possible to measure output as function of time. To achieve this, input must be inducible. The device F2621 is well studied and would be suitable for timer characterisation.
Output:
BBa_E0240 takes a POPs input and gives a measurable output of GFP.
Tuning:
Once the transfer function has been elucidated, it will be possible to investigate timer tuning via the addition of IPTG.
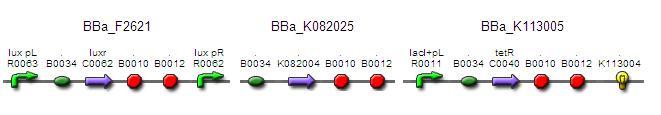
Testing construct 2:
This testing construct only requires 2 ligation steps:
Although, while this might have similar properties to our timer, it might not strictly count as characterisation of our own timer. It is also not known whether the two parts that are required are functional.
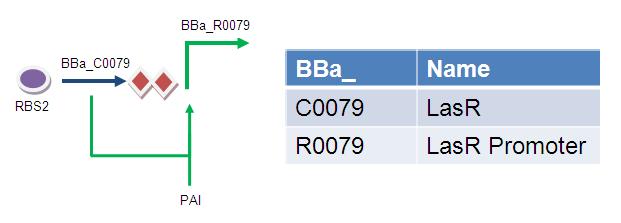
Simple Feed-forward Timer:
The Registry entry suggests that this promoter requires both PAI and LasR (e.g. it acts as an AND gate). However, it might be that LasR is sufficient for induction and PAI might just enhance induction (e.g. very leaky AND gate).
Note, I have not been able to find these parts fused together as Biobricks - therefore assembly would require 3 ligation steps (unlike the double inverter timer which only requires 1 ligation step).
Other Timer Info
PcstA (glucose-repressible promoter)
BBa_K118011
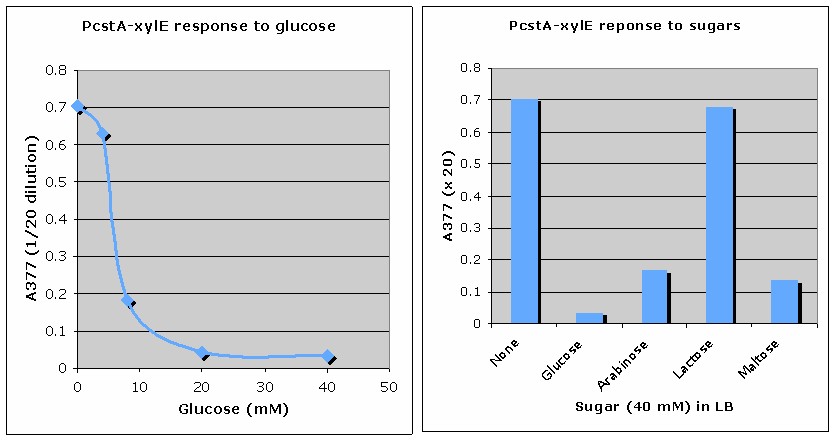 http://2008.igem.org/Team:Edinbrugh/Results/PcstA-xylE
http://2008.igem.org/Team:Edinbrugh/Results/PcstA-xylE
New Glucose Responsive Timer
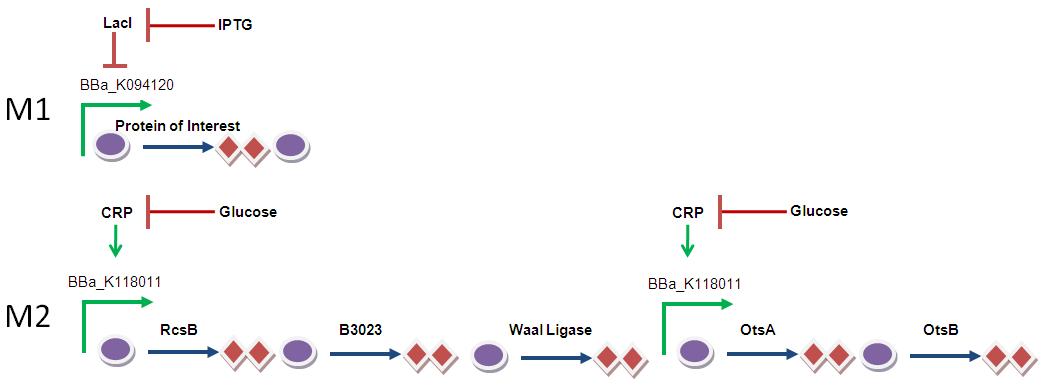
BBa_R0011 - this Biobrick is repressed by LacI and induced by IPTG. Perhaps this would be useful for the induction of the protein of interest. It can be used in combination with Kun's glucose-repressible promoter (BBa_K118011).
In the timer outlined below, M2 is triggered when the glucose in the medium runs out. M1 is selectivly triggered via the addition of IPTG, before which endogenously expressed LacI blocks expression of M1.
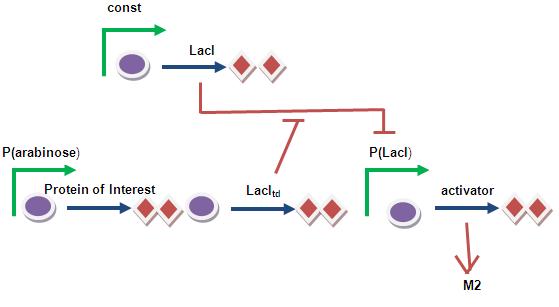
Lac(trans-dominant) Timer System
Uses the idea of transdominant mutants of LacI.
Altered repressor subunits that have lost the capacity to bind operator but can still aggregate with wild-type repressor subunits forming inactive tetramers. Therefore, the operon is expressed even in the presence of normal repressor. [1]
Problem is that the leaky promoter LacI is at the end of the circuit= might induce encapsulation inappropriately
Interesting and helpful paper on a simple timer [2]
Timers have many biological and engineering appplications. Design of mechanisms and characterisation of parts will provide a reusable timer module.
We could have a timer module as part of Modules 1, 2, 3 or 5 of our project.
Previous iGEM teams have explored different genetic circuits required to produce a timer.
Considerations for our timer:
- Separate timer for each phase vs. one timer that is ‘continuous’ between phases
- Thresholds
- Periodicity
- Start/stop vs. oscillation vs. combination of both
- Reset function
- Pre-programmed function(s)
- James Chappell 05:34, 21 July 2009 (EDT):What would the specs for our timer be? You should try to identify these first then look at the existing timers and see if they meet the specs, if not then look for new designs. Also Might be good to add diagrams of how the circuits work.
Specifications for a Timer in Each Phase of our Project
Module 1 - Compound:
- Induce production of compound at a specific time? - eg in response to something.
Module 2 - Encapsulation:
- Induce sporulation/encapsulation once desired threshold of compound X has been produced.
Module 3 - Killing Strategy:
- Induce death after encapsulation.
- Have a preset timer that induces death once the bacterium is encapsulated? (Chiba'08)
- Induce death immediately - would a timer be required? Why not just use a promoter? We could set a threshold rather than using a timer mechanism.
Module 5 - Delivery
General Specs
- Needs to be generic as much as possible (standard, independent output), there is no point in making a timer that only serves our application
- Needs to be inducible and resettable
- Needs to be tunable
- (optional, ideally) Monitor protein production and feedback on it
RiboTimer
Riboswitches are small pieces of RNA that regulate the translation of a gene. Many different times exist, however, because of the tunability property we seek in our timer, catalytic riboswitches are probably our best bet as they are concentration dependent.
The big problem is trying to engineer a riboswitch is too ambitious for this particular project as the main focus is encapsulation. Therefore if we want to pursue this route we should look for existing riboswitches rather than considering to engineer one.
Existing riboswitches: [3]; [4]
Riboswitch structure/engineering: [5]
Previous iGEM Timers
Chiba2008
Linear accumulation of non-degradeable signal molecules over time, which are then detected by receivers; in turn activating the switch to an 'on' state.
KULeuven2008
KULeuven used the 'InverTimer' in their 'Dr. Coli' project to induce destruction of their bacteria once the time lapsing between disease signals had reached a sufficient period.
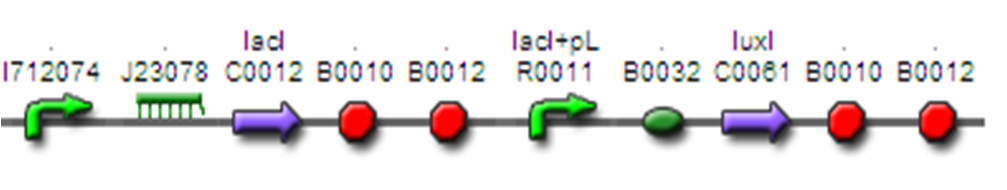
Their system is a LacI based inverter, controlling LuxI production. LuxI produces AHL (quorum-sensing molecule) which in turn gradually activats LuxR and its controlled promoter in the next module/device.
NYMU-Taipei2008
Taipei made two timers based on a three-component system:
Starter, Counter, Stopper
The counter component of the system is an oscillator. They required two timers with different periods so used different oscillators to fit the purpose of each timer.
Cyanoxilator
- A Cyanobacterial oscillator using the Kai proteins in E.coli.
- Time period of between 14 - 60 hours (...wide range there??)
Reloxilator
'A tuneable intracell-synchronized relaxation oscillator based on the combination of:
- Using the lysogenic genetic swtich from phage lambda and giving it tuneable oscillatory properties.
- Using the intracellular synchronising properties of Vibrio fischeri.
- Time period of ~46 minutes.
The timers are stopped once protein production reaches a pre-set threshold. Once this threshold is reached, the activity of the output promoter changes.
References
- Betz JL. Cloning and characterization of several dominant-negative and tight-binding mutants of lac repressor. Gene. 1986;42(3):283-92. DOI:10.1016/0378-1119(86)90232-5 |
- Ellis T, Wang X, and Collins JJ. Diversity-based, model-guided construction of synthetic gene networks with predicted functions. Nat Biotechnol. 2009 May;27(5):465-71. DOI:10.1038/nbt.1536 |
- Topp S and Gallivan JP. Guiding bacteria with small molecules and RNA. J Am Chem Soc. 2007 May 30;129(21):6807-11. DOI:10.1021/ja0692480 |
- Sudarsan N, Wickiser JK, Nakamura S, Ebert MS, and Breaker RR. An mRNA structure in bacteria that controls gene expression by binding lysine. Genes Dev. 2003 Nov 1;17(21):2688-97. DOI:10.1101/gad.1140003 |
- Topp S and Gallivan JP. Riboswitches in unexpected places--a synthetic riboswitch in a protein coding region. RNA. 2008 Dec;14(12):2498-503. DOI:10.1261/rna.1269008 |