IGEM:IMPERIAL/2009/Encapsulation/Modelling/Timer2
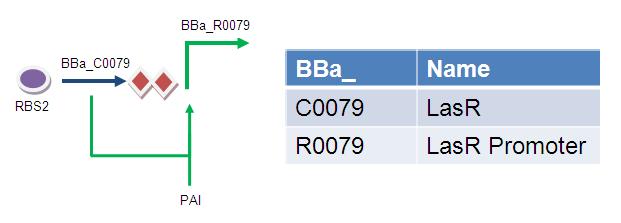
Diagram
David Roche 10:41, 24 August 2009 (EDT) : make it clear where this is part of the genetic circuit (which module?) and perhaps label the Biobricks with something other than the number so it is a bit more obvious! the key is ok, but perhaps include the whole genetic circuit so we can see how it all ties in. also LasR, is that the same as LuxR? not sure...
Operations
Truth table for the feed forward timer using LuxR
| INPUT | PAI | Output (encapsulation) |
| 0 | 0 | OFF |
| 0 | 1 | OFF |
| 1 | 0 | OFF |
| 1 | 1 | ON |
Hence, AND gate for encapsulation.
This is based on the assumption that both PAI and LuxR are above threshold.
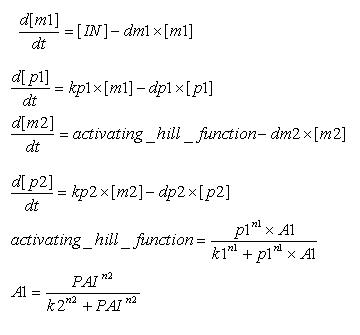
Model
[IN] - Concentration of LuxR
dm1 - Rate of degradation of mRNA 1
[m1] - Concentration of mRNA 1
[p1] - Concentration of LuxR
[kp1] - Rate of translation of mRNA 1 to LuxR
dp1 - Rate of degradation of LuxR
dm2 - Rate of degradation of mRNA 2, the mRNA coding for protein 2
[m2] - Concentration of mRNA 2
[p2] - Concentration of protein 2
kp2 - Rate of translation from mRNA 2 to protein 2
dp2 - Rate of degradation of protein 2
PAI - Concentration of PAI added
NB: protein 2 = encapsulation
David Roche 10:49, 24 August 2009 (EDT) : perhaps 'colanic acid producted' is better to use than 'amount of encapsulation,' (more quantifiable output). small thing i know, but for clarity
Simulations
Results
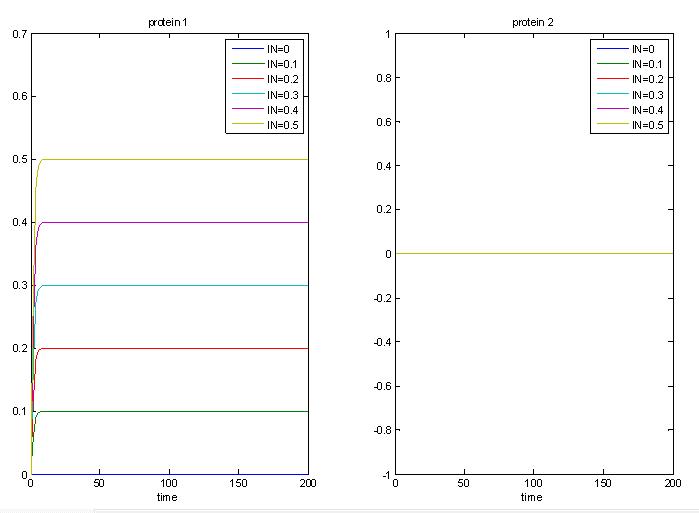
As a proof of concept that this system works as an AND gate, the graph below shows that when there is input, LuxR will be produced. However, since there is no PAI input, protein 2, will not be produced, and encapsulation will not occur.
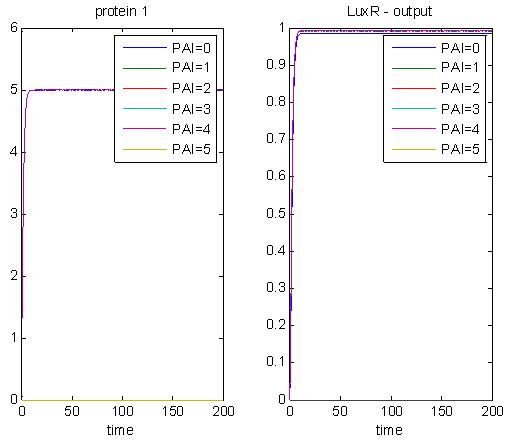
Here is another simulation, demonstrating the proof of concept.
The graph below shows that when there is LuxR and when enough PAI is added, there will be a stable output. The level of output depends on the amount of PAI added. A higher concentration of PAI means a higher encapsulation output. Here, the values of PAI are arbitrary, but will give an idea of how this system works.
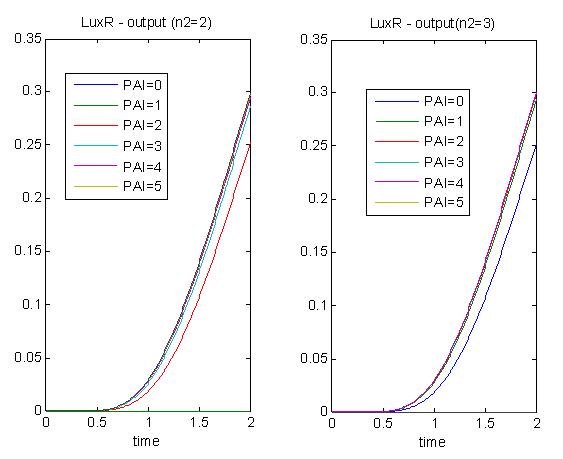
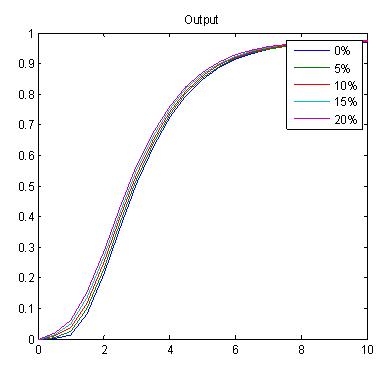
In the graphs below, we are varying the hill exponent of PAI (n2). In this sense, we are trying to explore the tunability of this timer. Varying n2 means we vary the steepness of the slope of the hill function. As n2 increases, we observe that the system acts more like a switch, and hence exhibits a decrease in tunability. As seen with the double inverter, the levels of PAI may affect the delay in encapsulation process, but tenability is within a very narrow range.
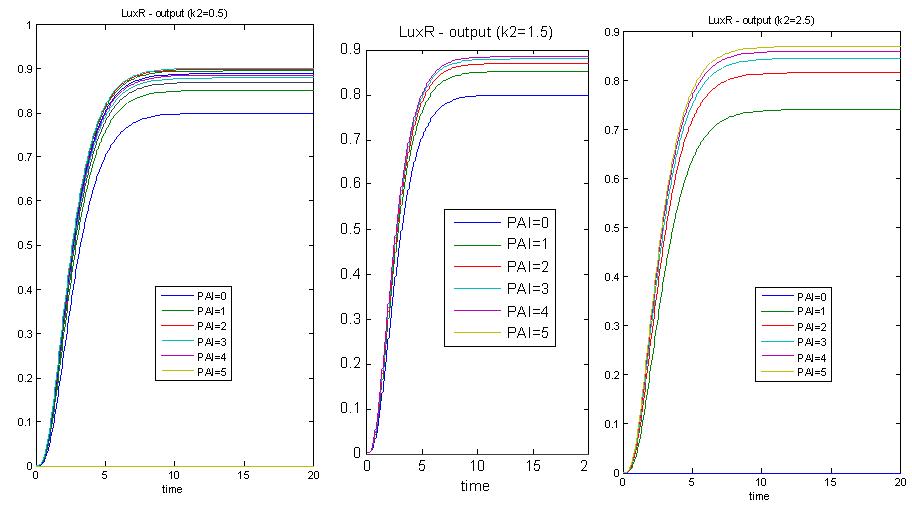
In these 3 graphs, k2(which controls the threshold of the p1 hill function) varies while other parameters are kept constant. K2 represents the threshold of the system, or the threshold of the PAI hill function. From the simulations, we realise that when k2 is large, we have less encapsulation. Whereas when k2 is small, we have more encapsulation, for the same amount of PAI. Hence, we have to trade off threshold with encapsulation. The significance of threshold is that we need to add more PAI to achieve the same effect.
When there is leakiness of the promoter, we see that when we include the effects of leakiness of the 2nd promoter, it will trigger encapsulation earlier. Therefore, in this timer, we want the output promoter to be tightly controllable.
Recommendations
- For lower values of PAI we get more apparent delays (assuming that sufficient protein 1 is present). The trade-off, however, is that we get a lower level of encapsulation, which is not desirable.
- For higher levels of PAI, the timer acts more like a switch and is not properly tuneable. (ie, miniscule levels of PAI will give us more tunability, but this is not what we want)
- We need to keep a relatively low hill exponent (between 1 and 3), if the hill exponent is too high, delays and tune ability will be lost, so the timer acts as a switch. The delay depends on the cut-off threshold defined in the hill function of LuxR and PAI.
- This design on its own is way too simple. If we want decent delays we will sacrifice on output yield again.
- The LuxR promoter controlling encapsulation cannot be leaky, as it would set off encapsulation earlier and mess the properties of the timer.