IGEM:IMPERIAL/2007/new pages/DNA concentrations
Testing DNA concentration of pTet-LuxR-pLux-GFPmut3b In vitro
Aims
To determine if the optimum concentration of pTet-LuxR-pLux-GFPmut3b in vitro.
Materials and Methods
Refer to protocols page.
Tested on (Insert link)
Results
 |
Controls:
- Negative Control- Nuclease Free Water was added instead of DNA
Constants:
- Temperature - 25°C
- Total Volume - 60µl
Raw Data
Discussion
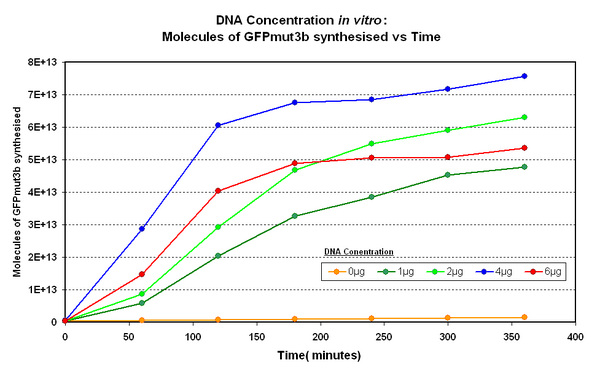
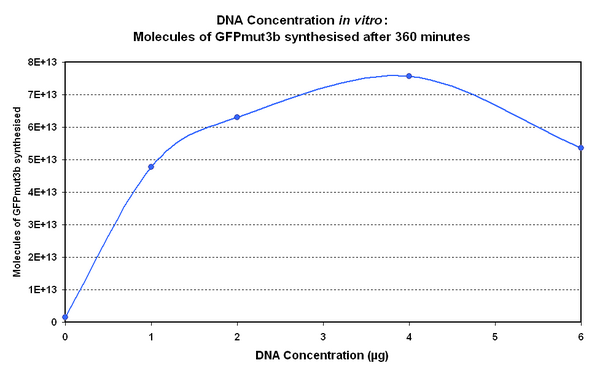
Figure 1.1 and Figure 1.2. shows us that the synthesis of GFPmut3b increases with DNA added up until 4µg of DNA. Above this the number of molecules of GFPmut3b produced decreases.
Another interesting observation is that the 6µg of DNA begins producing GFPmut3b greater than 2µg but then levels of before 2µg. This is thought to be because the data displayed on the figures are averages of 2 samples, one of the samples levels off sooner than the other and so brings the average down. However, the results do tell us that 6µg is lower than the 4µg, therefore the 4µg is the optimum DNA for our in vitro chassis.
This agrees with promegas guide on using the commcerial cell extract in vitro chassis. The guide states that 4µg is the maximum and above this there is problems with premature termination of translational products
Conclusion
To conclude the following approximations can be made:
- Optimum DNA concentrations - 4µg of DNA