IGEM:Hong Kong HKUST/Investigations/ Plasmid Cloning of BFP Generator Using Standard Assembly
<html> <body>
Plasmid Cloning of BFP Generator Using Standard Assembly
</body> </html>
Authors
- Winnie Ho
- Claire Moran
- Mike Dorothea
- Sharon Lai
- Olivia Winata
- Luna Eresta Jaya
- Kashish Aggarwal
- Yiu, Stephanie P.T.
Abstract
This training was conducted with the aim to learn the basic skills in producing a desired protein by using DNA recombination techniques such as digestion, ligation and transformation. In this training, constructs were made by inserting a Blue Fluorescent Protein (BFP) generator (BBa_K592023) into a parental plasmid containing a promoter (J61002-BBa_J23110).
The original plasmid containing the promoter (J61002-BBa_J23110) was digested using SpeI-HF and PstI-HF restriction enzymes, while the original plasmid containing the BFP generator (pSB3K3-BBa_K592023) was digested using XbaI-HF and PstI-HF. As a result, five different possible constructs might be produced: the desired construct, a construct with the wrong insert orientation, a construct with two backbones, a construct with two inserts and plasmids that are self-ligated.
Hence, this investigation will explore on methods that can help investigators identify candidate colonies which contain the desired sequence/construct and separate them from the rest. Colony PCR was chosen for colony identification after ligation and transformation. To distinguish the orientation of plasmid cloning, the primers VR and VF2 (which have opposite orientations) were used. Once the PCR products (in the form of amplified, transformed plasmids) were run in gel electrophoresis, the candidates showing the expected band size were concluded to have the desired construct (J61002-BBa_J23110-K592023). These particular candidates were then inoculated.
Introduction
By using BioBricks Standard Assembly, we can assemble two BioBricks containing our desired parts into the same plasmid, this can then be transformed into a cell and cloned to obtain our desired results. The objective of our project was to create a recombinant cell containing a BFP generator from the BioBricks BBa_J23110 and BBa_K592023. BBa_J23110 is a constitutive promoter, and BBa_K592023 is an intermediate containing a BBa_K592100 Blue Fluorescent Protein and a BBa_B0032 RBS. After combining the two BioBricks by restriction digestion and ligation, then transforming the recombinant plasmid into a competent cell, blue fluorescent protein producing E. coli could be obtained. The details of the BioBricks used are shown in Table 1. BioBricks used in this investigation.
Methods and Materials
BioBricks
Table 1. BioBricks used in this investigation.
| BioBricks | Size (bp) | Plasmid | Description |
| BBa_J23110 | 35 | J61002 | RFP Producing Constitutive Promoter |
| BBa_K592023 | 724 | pSB3K3 | RBS (BBa_B0032) and BFP CDS (BBa_K592100) |
Restriction Enzyme Digestion
The plasmid containing the constitutive promoter (J61002-BBa_J23110) was cut by the restriction enzymes SpeI-HF and PstI-HF and served as the vector in the ligation, while the plasmid containing the BFP generator (pSB3K3-BBa_K592023) was cut by XbaI-HF and PstI-HF and was isolated as an insert. The buffer used was CutSmart and the restriction digestion process was carried out for 1 hour 30 minutes at 37°C. The volumes of the reagents used for this process are described in the following table.
Table 2. Restriction Enzyme Digestion Recipe
| J61002-BBa_J23110 | pSB3K3-BBa_K592023 | |
| DNA Amount (ng) | 300 | 500 |
| Sample DNA Concentration (ng/μl) | 67.14 | 192.3 |
| DNA Sample Volume (μl) | 4.46 | 2.60 |
| Xbal-HF (μl) | - | 0.20 |
| Spel-HF (μl) | 0.20 | - |
| PstI-HF (μl) | 0.20 | 0.20 |
| CutSmart Buffer (μl) | 1.80 | 1.80 |
| ddH20 (μl) | 11.34 | 13.20 |
Gel Electrophoresis, Purification, and Nanodrop Test
Gel Electrophoresis was performed for 30 minutes using 1% agarose gel and midori green as dye. The voltage and current used were 130 V and 400 mA respectively. The expected bands were 2726 bp for J61002-BBa_J23110 and 724 bp for BBa_K592023. After the gel photo was obtained, desired band was taken and purified according to the protocol. The purified band then tested for its concentration and protein and salt contamination by using Nanodrop machine.
Ligation
0.5 μL of T4 DNA Ligase was used to ligate J61002-BBa_J23110 with BBa_K592023. The proportion amount of backbone (J61002-BBa_J23110) to amount of insert (BBa_K592023) is 1 to 3.
Transformation
10 μL of ligated product was inserted into 50 μL of competent cell and put into ice for 20 minutes. After 20 minutes, heat shock and cold shock were performed. After undergo these processes, 1000 μL of LB was added into the tube, and the tube was centrifuged for 2 minutes on 6500 rpm.
Incubation
Two plates of agar containing the AMP antibiotics were taken out from the fridge, and the transformed bacteria containing the desired plasmid were spreaded on the plates. These plates were incubated overnight.
After colonies were cloned, some of the bacteria were taken as samples for PCR and streaked on new compartmentalized plates that served as back up for another PCR.
Colony PCR
First, mastermix containing 20 folds of 2 μL of 10X ThermoPol buffer, 0.5 μL of 10 mM dNTPs, 0.5 μL of VF2 as foward primer and 0.5 μL of VR as reverse primer, 0.15 μL of Taq DNA Polymerase, and 14.35 μL of ddH2O. This mixture was distributed to 16 eppendorfs which at the end contained 18.0 μL of the mastermix and added with 2.0 μL of template DNA before proceeding to PCR. The first step of PCR was initial denaturation to burst out the cell for 30 minutes at 95°C. Next, denaturation was done for 30 seconds at 95°C to separate the double stranded DNA. After that, annealing was done for 1 minute, followed by extension for 1 minute 30 seconds. The final extension process was conducted for 4 minutes 30 seconds, which was 3X longer than the main extension. The final step was holding for around 10 minutes at 10°C.
Gel Electrophoresis
The same step was performed as the previous gel electrophoresis.
Inoculation
5 ml of LB containing AMP antibiotics was added to named empty falcon tubes. The colony was inserted to the falcon tubes and incubate overnight.
Miniprep
Miniprep was done according to the protocol. Then nanodrop was performed.
Restriction Check
Restriction check was not applicable in this case because the restriction enzymes we planned to use were denatured, and since the backbone is J61002_J23110, if we use EcorI and PstI the negative results will show similar bands to the experimental setup.
Results
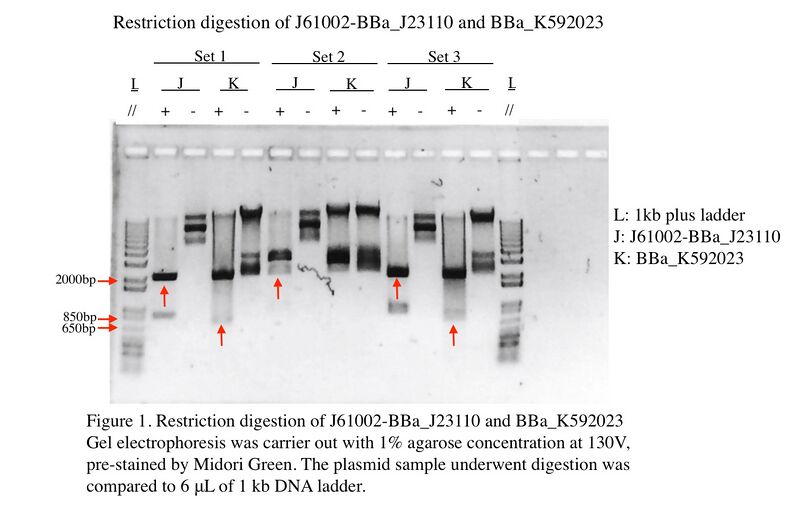
Figure 1 shows the Gel Electrophoresis of J61002-BBa_J23110 and BBa_K592023 candidates after digestion. There are three set of J61002-BBa_J23110 and BBa_K592023 we have used. Set one and set three show both expected band size which is 2276 base pair for J61002-BBa_J23110 and 724 base pair for BBa_K592023. After that, the expected bands were extracted for ligation.
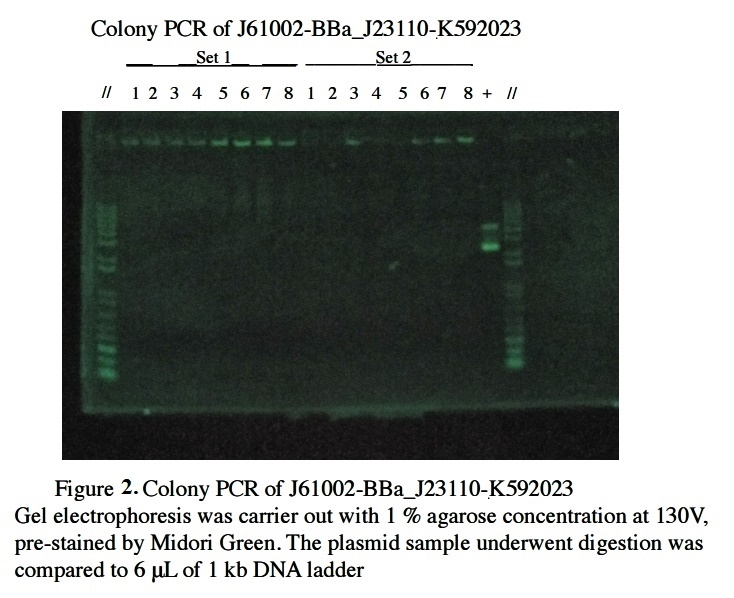
Figure 2 shows the Gel Electrophoresis of J61002-BBa_J23110-K592023 candidates after colony PCR. There are no any bands in the 8 candidates but band is shown in the positive control, that is using similar bp plasmid comparing to the decided plasmid. This results indicate that the PCR procedure had been functioning and errors occurred in the candidates.
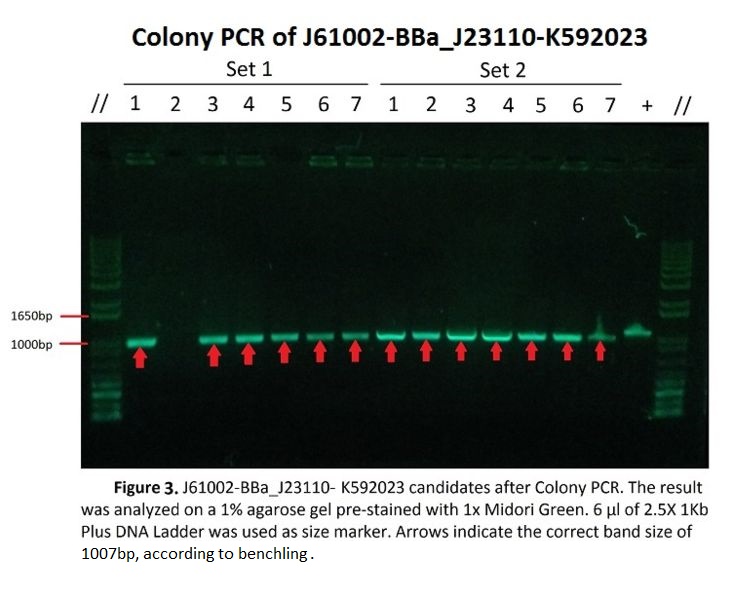
Figure 3 shows Gel Electrophoresis of J61002-BBa_J23110-K592023 candidates after colony PCR. As seen in the picture, the target bands of 1007 base pairs are shown, measured by comparison with the DNA ladder (1Kb Plus).
All of the candidate samples from both sets of the experiment showed the expected band, except for Sample 2 in Set 1 which showed no band at all. This, along with the uneven brightness of the bands might have been caused by some random and systematic errors during the experiment, which will be talked about under Discussions.
Discussion
On the third time of doing the PCR, bands with the expected size of 1007 bp appeared on the gel. This indicates that there is a high probability that the candidates from the PCR are in fact correct and contain the desired plasmid, J61002-BBa_J23110-K92023. Furthermore, the result is aligned with, and thus strongly supports the blue-colored appearance of transformed bacterial colonies under UV light upon incubation.
Nonetheless, only the last out of three trials of Colony PCR showed successful results. There are some probable reasons as to why the previous PCR was not successful. The first reason is due to the turbidity of the plasmid mixture upon loading into gel electrophoresis. There might have been too many cells in the samples, hence the gel could not separate and run the plasmids properly. Second, the amount of Taq polymerase that was added might be insufficient, thus the amplification was not enough to show distinguishable bands in the gel. Although quite superficial, human error when mixing the master mix could also highly contribute to the error.
Apart from this error, the result on gel electrophoresis after both restriction digestion and colony PCR both showed some smears and bands that are not sharp or with uneven brightness. This might be due to the incomplete digestion of plasmids for restriction digestion as well as human errors in loading the plasmid sample to the gel for both scenarios.
Improvements
In order to avoid smears in restriction digestion gel electrophoresis, perhaps more restriction enzymes or longer digestion time should be allocated in order to ensure complete digestion. Moreover, each of the loading of sample into the gel prior to electrophoresis should be done extra carefully, ensuring that the sample mixture was mixed beforehand and all the sample in the pipette has been completely loaded.
The process of making the master mix must be done with more care and precision to reduce systematic errors or failure. Also, adding more Taq polymerase into the mixture could also help to stronger amplify the DNA. In order to improve the accuracy of results, proper restriction check and sequencing should be done to further verify the candidates.
Conclusion
In conclusion, based on the PCR results, it is highly likely that the results obtained were the ligated product of BBa_J23110 and BBa_K592023 in J61002 backbone. However, restriction check and sequencing should be done to further verify the candidates.