IGEM:Hong Kong HKUST/2009/Notebook/Investigation on the Functionality of RNA Thermometers/Entry Base
 Investigation on the Functionality of RNA Thermometers Investigation on the Functionality of RNA Thermometers
|
|||||||||||||||||||||
Authors
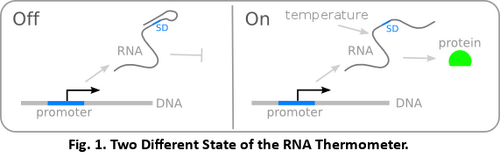
AbstractDesigning gene circuits with RNA thermometers can temporarily control the extent of expression by changing the temperature periodically. RNA thermometers, which are temperature-sensitive, are useful in engineering complex genetic circuits to produce specific and distinct results. It can regulate gene expression at post-transcriptional level by forming hairpin or stem-loop structure on the Shine-Dalgarno sequence (SD). At temperature higher than its unique activating temperature, the secondary structure of the RNA thermometer is altered and the occluded ribosome binding site is exposed, which enables translation. Without sufficient characterization data on the iGEM Registry and iGEM teams wiki pages which indicate the functionality of the RNA thermometers, our investigation was carried out to verified the performance of a list of RNA thermometer at 37℃ and 20℃, reported by the expression of fluorescent protein. IntroductionDue to its temperature-sensitive secondary structure, the RNA thermometer, which has its own activation temperature, is able to control the translation rate by exposing or occluding the ribosome binding site. On the left (off-state), when the environmental temperature has not reached its activation temperature, the hairpin or stem-loop structure of RNA thermometer will occlude the Shine-Dalgarno sequence (SD sequence) to inhibit the binding of small ribosomal subunit which in turn hampers translation. On the contrary, on the right (on-state), when the threshold temperature is reached, the secondary structure of the transcript will undergo conformational change (the secondary structure melts) such that the SD sequence will be exposed and the binding of ribosome is possible to initiate translation. In this investigation, we transformed genes with the six different RNA thermometers into the Escherichia coli DH10B strain, which were incubated at 20℃ and 37℃ and observed for the presence of fluorophore. Methods and MaterialPlasmid ConstructionTo investigate the functionallity of RNA thermometers, genes encode for monomeric red fluorescent protein (mRFP) were constructed with different RNA thermometers of interest and were transformed into E. coli DH10B strain. Cells were then streaked onto agar plates and were investigated. Six RNA thermometers, one promoter, one reporter with double terminators and a mRFP coding device were used, as listed below:
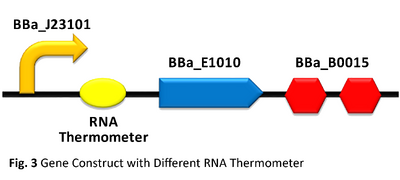
The three parts were assembled using the RFC10 standard as illustrated, with pSB1C3 as the backbone. BBa_K516132 was used as a positive control with BBa_B0032 as the reference RBS. Negative control was constructed with the parts: BBa_J23101+BBa_J04650. All constructed genes were transformed into E. coli DH10B strain.
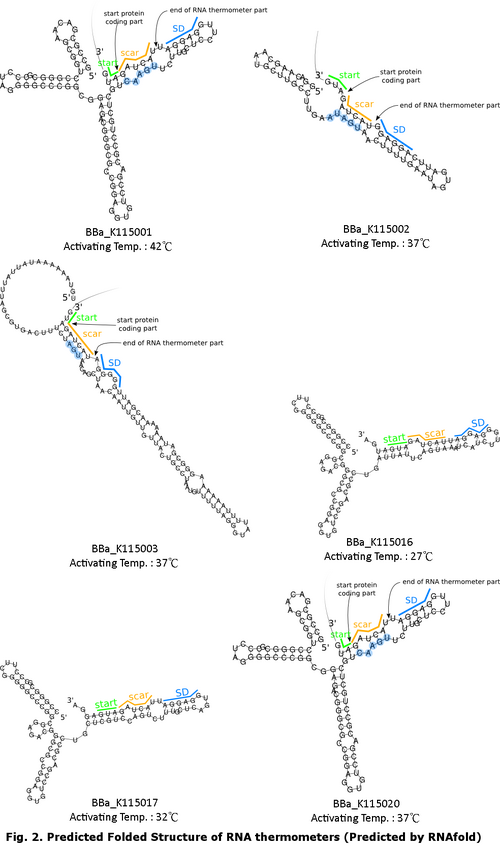
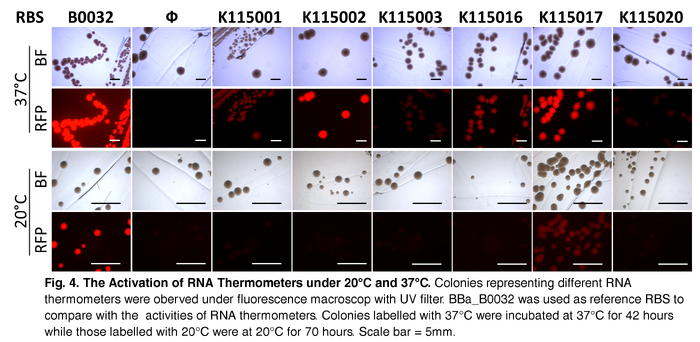
After incubation, both sets were investigated under UV light and white light with Olympus MVX10 Fluorescence MacroZoom to observe the fluorescence of the colonies. Agar Plate PreparationFor 800mL of agar, 8g of Bacto-tryptone, 4g of yeast extract, 8g of sodium chloride and 12g of agar were autoclaved and dissolved in double-distilled water. 800 μL of 25 µg/mL chlorapmhenicol is added before pouring. Result and InterpretationAt both temperatures, positive controls showed bright red colonies. Negative control at 37°C showed no trace of fluorescent while at 20°C dim red fluorescent. Contrasting with the negative control, all RNA thermometers incubated at 37°C showed red colonies, suggesting activation of all RNA thermometer at 37°C. For the set incubated at 20°C, when compared to negative control, only BBa_K115017 showed significant difference, indicating activation of this RNA thermometer at 20°C. Comparison of each RNA thermometer across two temperature settings suggests BBa_K115002, BBa_K115003, BBa_K115016 and BBa_K115020 performed as anticipated to function only at 37°C. While for BBa_K115001, appearance of red fluorescence at 37°C was not consistent with the expectation of activation at 42°C. Similarly, for BBa_K115017, appearance of red fluorescent at 20°C was not consistent with the expectation of activation at 32°C. DiscussionIt is expected that for the secondary structures of RNA thermometers BBa_K115001 and BBa_K110017 to melt and expose the SD for translation initiation, the temperature should exceed their corresponding activation temperatures of 42°C and 32°C respectively. Yet, in accordance with Fig. 4. The Activation of RNA Thermometers under 20°C and 37°C, characteristic red colonies carrying BBa_K115001 were observed in plate incubated at 37°C while that carrying BBa_K115017 were observed at 20°C. This suggests that even though the activation temperature is not reached, the secondary structure of the RNA thermometers becomes unstable, exposing the SD sequence, at a temperature lower than the corresponding threshold temperature. Another possible reason for this is the activation temperature of these two RNA thermometers may be lower than the proposed temperature. When building a complex logic circuit, in which the designers want to manipulate gene expression post-transcriptionally in an independent and specific fashion, RNA thermometers such as BBa_K115001 and BBa_K115017, which are vulnerable to a wide range of temperature, is not suitable for the implementation of an accurate temperature-dependent model. The fluorescence of BBa_K115003 when compared to that of the negative control plate, is observed to be very weak even though it reaches the activation temperature. While the consensus Shine-Dalgarno sequence of E. coli is 5’-AGGAGG-3’ (Gold & Stormo, 1987), it is possible that the pairing efficiency between the SD of listed RNA thermometers, i.e. 5’-GGGGGA-3’, and the E. coli small ribosomal subunit is reduced. The reduction in pairing efficiency subsequently decreases the rate of translation, resulting in a lower concentration of mRFP produced (John, 1991). This experiment is limited to qualitative analysis of each RNA thermometers. For a quantitative measure of the effect of change in temperature on gene expression level, flow cytometry can be applied to obtain relative amount of fluorescent protein in cells at different temperatures. This application is useful for, firstly, comparing the expression level of mRFP under regulation of two RNA thermometers having the same activation temperature. Also, by growing cell culture in a range of temperatures, comparison between the gene expression and its theoretical activation temperature is possible to determine the specificity of the RNA thermometer to a particular temperature. Knowing the relative fluorescence level allows us to occlude biased judgment based on the brightness of the colonies. ConclusionThe focus of this investigation has been on the functionality of the RNA thermometers. BBa_K115002, BBa_K115003, BBa_K115016 and BBa_K115020 have been identified to be potentially temperature-sensitive, however, further characterization have to be done to measure the activation temperature and strength of these RNA thermometers. ReferenceFig. 1. 2 Different States of RNA Thermometer. Retrieved February 8, 2015, from http://2008.igem.org/Team:TUDelft/Temperature_overview Fig. 2. Predicted Folded Structure of RNA Thermometers (Predicted by RNAfold). Retrieved February 8, 2015, from http://parts.igem.org Gold L., Stormo G. (1987) Translational initiation. in Escherichia coli and Salmonella typhimurium: cellular and molecular biology. eds Neidhardt F. C., Ingraham J. L., Low K. B., Magasanik B., Schaechter M., Umbarger H. E. (American Society for Microbiology, Washington, D.C), pp 1302–1307. Johnson G (1991). Interference with phage lambda development by the small subunit of the phage 21 terminase, gp1. Journal of Bacteriology 173(9): 2733–2738. PMID 1826903. | |||||||||||||||||||||