IGEM:Harvard/2009/Lab Notebook/Week0
Tuesday: June 9, 2009
TAs arrived in the evening. Resuspended DNA in Biobricks library plates with 15ul of buffer EB from Qiagen kit. Samples turned pink, indicating resuspension. The samples were kept resuspended within the Biobricks 384 well plate and stored at –20C.
For Biobrick part locations: http://partsregistry.org/Help:IGEM_09_DNA_distribution
Parts resuspended:
| Registry # | Plate | Location | Resistance |
| BBa_120270 | 2 | 17B | Kan |
| BBa_120269 | 3 | 21D | Kan |
| E0040 | 1 | 14K | Amp |
| BBa_P1010 | 1 | 7M | Kan |
1ul of resuspended plasmid was used for each transformation. The transformation was only slightly modified from the standard protocol.
Wednesday: June 10, 2009
Although not very many, colonies grew for all plasmids except for part BBa_P1010, which was transformed into strain DB3.1. No colonies were detected.
Thursday: June 11, 2009
Miniprep of (1) high-GFP, (2) medium-GFP, and (3) GFP-vector. Done to extract DNA for diagnostic and eventual extraction digests, for production of low-GFP; also done to visualize differences in expression levels with different promoters. Followed Quiagen protocol for minipreps. Did three preps of 3 mL ea, (1) high-GFP, (2) medium-GFP, and (3) GFP-vector. Used Nanodrop to look at DNA concentrations, recorded on outside of vials. (1) was approximately 47 ng/uL, (2) was 90 ng/uL, and (3) was 200 ng/uL.
Diagnostic digest of GFP-vector. Done for practice, to see if digest works so we can do a larger scale digest and cut out the GFP and to visualize bands (GFP should be 700-800 bp in size). David made the mastermix, 5 uL GFP-vector DNA (~200 ng/uL) and 10 uL of master mix. Digest with EcoRI and SpeI for 1 hr at 37 degrees.
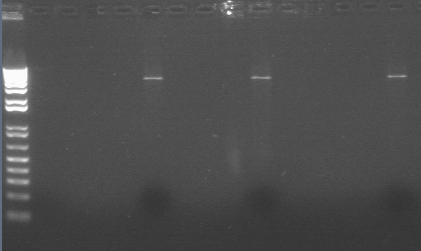
E-Gel for diagnostic digest of GFP-vector. Done to determine if the digest works. Need to load 20 uL. Only ladder has dye, samples do not need it because gels are buffer free. Prepoured gels. See picture of gel on website. Lane 1—Ladder, Lane 6—Undigested (5uL DNA, 10 uL water), Lane 7—Digested
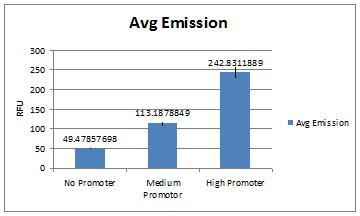
Use of spectrophotometer to compare expression of GFP under control of different promoters. Excitation was set to 488 nm and emission was set to 511 nm. Cells vortexed and added to cuvettes (make sure they are well mixed). Took OD for each sample and GFP fluorescence at approx 514. Found that all samples (all three, in quintuplicate) had similar OD values. Fluorescence under control of medium promoter was approx 2x basal levels of expression with no promoter. Fluorescence under control of high promoter was approx 2x levels with medium promoter (about 225 for high, 120 for medium, 70 for low; all ODs were about 0.75).
First, 1/10 dilutions of the overnight cultures we made, with LB broth. The Spec was blanked with LB. 1 ml of each sample was loaded into the cuvettes. Background was found to be 48.30 at 600 nm.
| Sample | OD600 nm | Emission 511 | ' | |
| Med | 0.232 | 51.58 | ||
| High | 0.261 | 57.75 | ||
| ' | Sample | OD600 | Emission 511 | Emission minus background | Corrected emission / OD | ' |
| No | 1-No | 0.684 | 82.13 | 33.83 | 49.45906433 | |
| Med | 2-Med | 0.691 | 132.69 | 84.39 | 122.1273517 | |
| High | 3-High | 0.674 | 224.06 | 175.76 | 260.7715134 | |
| No | 4-No | 0.735 | 85.29 | 36.99 | 50.32653061 | |
| Med | 5-Med | 0.701 | 130.51 | 82.21 | 117.275321 | |
| High | 6-High | 0.767 | 228.89 | 180.59 | 235.4498044 | |
| No | 7-No | 0.722 | 84.98 | 36.68 | 50.8033241 | |
| Med | 8-Med | 0.795 | 129.67 | 81.37 | 102.3522013 | |
| High | 9-High | 0.721 | 232.17 | 183.87 | 255.0208044 | |
| No | 10-No | 0.747 | 84.99 | 36.69 | 49.11646586 | |
| Med | 11-Med | 0.745 | 131.97 | 83.67 | 112.3087248 | |
| High | 12-High | 0.742 | 214.47 | 166.17 | 223.9487871 | |
| No | 13-No | 0.8 | 86.45 | 38.15 | 47.6875 | |
| Med | 14-Med | 0.757 | 132.99 | 84.69 | 111.8758256 | |
| High | 15-High | 0.715 | 219.16 | 170.86 | 238.965035 | |
| Superpos control from David | 16-Superpos | 1.08 | 3659.33 | 3611.03 | 3343.546296 | |
Friday: June 12, 2009
Outlined plan for today:
1. Digest of GFP and vector
2. Gel purification by using either
(a) The filter paper method (better yield)
(b) Qiagen Gel Purification Kit (contaminants go in column which might obscure spec results, but faster, with this method the gel gives nice results but the spec sometimes gives negative values)
Minipreps of psB3K3 to extract DNA. Anu and Amy did minipreps of my 4 mL cultures of psB3K3 transformed bacteria, and set up my digests. Digested with SpeI and EcoRI.
DNA spec results (values in concentration units of mg/microliters) XG low: 73.6 IB low: 38.2 AR low: 30.4 AG low: 91.3 NK low: 73.6 GD low: 92.0
Double digest protocol: Restriction enzymes: EcoRI + SpeI (New England Biolabs) Common digestion ingredients: (1) Enzymes - to cut the DNA and catalyze the "digestion" reaction (2) DNA - the digestee (3) Buffer (10x)- provides necessary nutrients to run the reaction (4) BSA - adds extra protein/enzyme cushioning which simulates the natural enzymatic environment, chemically innocuous (5) H20 - get up to final volume
GFP insert digest 30 microliters total volume (DNA in EB buffer) We have 17 microliters of DNA left since 2, 5, and 5 microliters of our stock was used up in the spec, digest sample, and undigest samples, respectively
This is our digest suspension (30 microliters total) 17 plasmid in EB 3 buffer (EcoR1 buffer: we are using this buffer becuase since we are doing a double digest, these are complimentary and the buffer works for both enzymes. Usually the reason why we are forced to do single separate digests is because of buffer incompatibility) 3 BSA 1 EcoR1 1 Spe1 5 H20
14x volume (we have 11 tubes)
So in total we have: 42 EcoR1 Buffer 14 Spe1 14 EcoR1 42 BSA 70 H2O
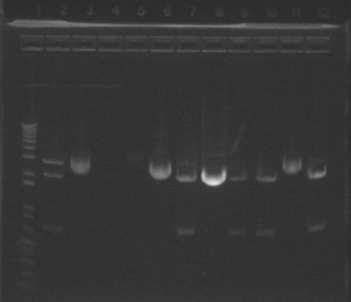
Making agarose gel for extraction of GFP from vector (GFP-vector). Ethidium bromide—0.5uL per mL Medium size gel tray holds 150 mL of agarose, when making 1% gel, 1.5 g agarose and 150 mL TBE. When making a gel use the same buffer to make the gel and as the buffer. Microwave for about 1-2 minutes, cap not tight, careful about sudden boiling, do not reboil after add ethidium bromide. Do not use short wave UV to visualize DNA, it is mutagenic. Poured two gels and ran out GFP-vector digest on one and pSB3K3 on the other one. When imaged using long wave UV found that GFP fragment was right size (about 700) and use filter paper extraction to get DNA. The psB3K3 band was the wrong size, expected about 2.35 kB, but got about 7 kB.
The filter paper extraction is much much more efficient, maybe 5/6, whereas with Quiagen best is going to be 1/10 extraction. After nanodropping, we found the concentration of the DNA to be around 8.6 ng/uL.
We also resuspended our primers.
On Monday we plan to assemble the low promotor and run a diagnostic gel.