IGEM:Harvard/2006/DNA nanostructures/Notebook/2006-8-14
From OpenWetWare
Jump to navigationJump to search
Goals for today
Microcon Purification Tweaking
- repeat Friday's mega PEG ppt on 5.0 (?)
- Micron experiments with 0.1% and 0.01% SDS in buffer
- ...and use 1x folding buffer and not water for washes
- also: perform control expt with 10 bp+ ladder, since according to Millipore documentation, the filter should retain ds DNAs longer than 100 bp
Streptavidin-Bead "Protection" Assay on Inside- and Outside-Biotinylated c5.0
- NB: no good purification of nanostructure from oligo has been achieved, but gel separation after elution should differentiate formerly bead-bound oligos from formerly bead-bound nanostructures
Redux of [Mg++], [oligos]
- Based on the inconclusive gels from last Monday, the titration will be redone with c5.0 at 8% PEG, 0.5M NaCl final.
Mg2+, Oligo-Concentration Titration w/ c5.0
Goals
- vary folding conditions ([MgCl2] and [oligo]) in order to determine best folding conditions for c5.0
- determine most efficient purification protocol (Microcon vs. PEG) based on recovery yields
Protocol
1. Working Stock Concentration
- concentrated 6 tubes of 96 μL c5.0D.L (no latches, outside-bound ligand) in Vacufuge so that [oligo]= 250nM * 6 = 1.5 μM
2. Folding Rxns
- used three different folding buffers varying [MgCl2]
- used two different [oligo concentrations]: 250 nM from the unconcentrated working stock, 1.5 μM from above
- folding conditions: 80[[:Category:{{{1}}}|{{{1}}}]] for 2 min., decrease 1[[:Category:{{{1}}}|{{{1}}}]] every 2 min. for 59 more times
Notes
- a,b - a and b of each are the same, just two different tubes.
- Oligos - 250 nM is 1x oligos, 1.5 {{um} is 6 oligos
| Trial | Oligos | p7308 (44 nM) | Folding Buffer (10x) | Water |
| 1a,b | 16 μL 250 nM | 9 μL | 4 μL 100 mM MgCl2 | 11 μL |
| 2a,b | 16 μL 1.5 μM | 9 μL | 4 μL 100 mM MgCl2 | 11 μL |
| 3a,b | 16 μL 250 nM | 9 μL | 4 μL 200 mM MgCl2 | 11 μL |
| 4a,b | 16 μL 1.5 μM | 9 μL | 4 μL 200 mM MgCl2 | 11 μL |
| 5a,b | 16 μL 250 nM | 9 μL | 4 μL 300 mM MgCl2 | 11 μL |
| 6a,b | 16 μL 1.5 μM | 9 μL | 4 μL 300 mM MgCl2 | 11 μL |
Microcon w/ detergent
- add 20 μL given nanostructure to center of YM-50 Micrcon tube
- add 480 μL given folding buffer, microcentrifuge for 6 min. at 14k rcf, and repeat dilution and spinning 4 more times
- yielded approx. 100 μL retentate, which was concentrated to 15 to 60 μL in a Vacufuge (about 30 min. at 45 [[:Category:{{{1}}}|{{{1}}}]]), depending on the sample
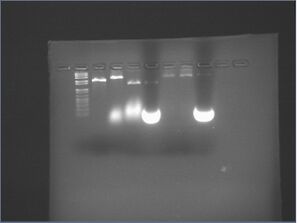
| lane | starting amt. of nanostructures | wash buffer | loaded onto gel |
| 1 | 7 μL 1 kb+ ladder | ||
| 2 | 2.25 μL p7308 | ||
| 3 | 10 μL unpurified 4.0.I | ||
| 4 | 20 μL 6hb | 1x folding buffer (10 mM MgCl2) | half of retentate |
| 5 | 20 μL 6hb | 1x folding buffer (10 mM MgCl2) w/ 0.1% SDS | half of retentate |
| 6 | 20 μL 4.0.I | 1x folding buffer (10 mM MgCl2) | half of retentate |
| 7 | 20 μL 4.0.I | 1x folding buffer (10 mM MgCl2) w/ 0.01% SDS | half of retentate |
| 8 | 20 μL 4.0.I | 1x folding buffer (10 mM MgCl2) w/ 0.1% SDS | half of retentate |
- ran 2% agarose gel at 80 V for 1 h. Gel appears to be of such low qualitiy that the results are inconclusive (ladder isn't clear)
- ran another 2% agarose gel at 60V for 1 h.
- results/discussion
- unclear why 6hb filtration failed (oligos were retained) (lane 4)
- 0.1% SDS gives unusual/unknown smears (lanes 5 and 8)
- under visible light, there are thin pink-red bands in the middle of these smears. is SDS breaking down the Microcon tube plastic?
- 0.01% SDS possibly gives higher yields than no SDS (lane 7 vs. lane 6), but it is not conclusively better yield, and it is still very poor overall yield (lane 7 vs. lane 3)
Streptavidin Bead "Protection"
- Goal:
- Determine whether c5.0 functions like a container - in other words, can it "protect" biotinylated sites on the inside from being bound by streptavidin? In order to do this, we can use 1 micrometer-in-diameter magnetic streptavidin beads (ie. bigger in diameter than the hole of the c5.0 barrel, so the streptavidin bound to the bead won't be able to access the internal biotin).
- If we remove the material that is not streptavidin-bound initially (hopefully, internally-biotinylated barrels) and then elute the material which was streptavidin-bound (externally-biotinylated barrels, free-floating biotinylated oligos), a gel run with the elutes will show DNA material for the externally-biotinylated structures.
- Structural Justifications:
- NEB's magnetic streptavidin beads are 1um in diameter, too large to fit into the 30nm-wide barrel of the c5.0.
- However, conceivably you could imagine the streptavidin binding region being able to snake into the barrel, as streptavidin's dimensions are 54 x 58 x 48 angstroms, or 5.4 x 5.8 x 4.8 nm, which would allow it to fit in the barrel.
- Biotinylated sites within the barrel are attached to oligos which are 3 Ts from the scaffold-oligo binding point, thus making this a possibility, but hopefully not a frequent one.
- Issues:
- No protocol found for the Fluka agarose streptavidin beads, and the NEB magnetic bead protocol's heating requirements seem unsuitable for keeping nanoboxes folded throughout the process
- Nanoboxes must remain folded post-elution because they must be gel-distinguishable from eluted biotinylated oligos, which we have not been able to pre-purify out by other means.
- No protocol found for the Fluka agarose streptavidin beads, and the NEB magnetic bead protocol's heating requirements seem unsuitable for keeping nanoboxes folded throughout the process
- Elution methods are of two types:
- 1. Overload with competitors - either streptavidin or biotinylated oligos
- But can't: biotin-streptavidin bond is so strong that dissociation by competition would theoretically take years
- 2. Denature the streptavidin using a) formamide, b) phenol, c) SDS + boiling
- But can't: all the conditions require heat that might damage the nanostructures (65[[:Category:{{{1}}}|{{{1}}}]] and up)
- Solution:
- Degrade streptavidin with trypsin
- Trypsin must be in solution w/o EDTA, which would otherwise chelate all the Mg2+ in the DNA nanobox solution that is necessary to keeping it folded.
- Degrade streptavidin with trypsin
- Protocol:
1. Incubate:
5uL beads (binding capacity: 2pmol/1uL, thus, 10pmol - far greater than the available binding sites in the DNA nanobox solutions)
35uL 1x folding buffer
10uL test solution
TEST SOLUTIONS:
--------------
a) H2O (ie. test = no biotin in solution)
b) biotinylated oligos (c5.0.8(b)) - 250uM per oligo in pre-working stock = 1mM biotinylation - for 1.6pmol = 1.6uL - thus, 1.6uL + 8.4uL H2O must be added for test solution
c) c5.0 E(b) (outside biotinylation) (was Microcon "purified" Tu 8.9) - ~2000fmol of binding sites/12.5uL, or 0.16pmol/ul - thus, 1.6pmol
d) c5.0 F(b) (inside biotinylation) (was Microcon "purified" Tu 8.9) - "
2. Mix
3. Pellet by drawing magnet down to bottom of tube (7 minutes using the old magnet)
4. Discard supernatant
5. Add 50uL 1x folding buffer
6. Repeat steps 2-5 three more times.
7. Trypsinize by adding to pellet:
3uL trypsin (1mg/mL)
27uL 1x folding buffer
8. Incubate 4hr-overnight @ 37 degrees C
9. Pellet by drawing magnet down to bottom of tube and remove supernatant to clean tube (1 minute, using the Magnetorack).
10. Run 20uL of each supernatant on 2% agarose gel (10mM MgCl2) for 1hr at 80V.
- Gel:
- please see 8.15.06 Notebook page