IGEM:Harvard/2006/DNA nanostructures/Notebook/2006-7-19
From OpenWetWare
Jump to navigationJump to search
Protection assay
Incubation and dilution protocol, take 2
- Goal: to see the lower threshold of imaging streptavidin bound to biotin
- and to see if we can image any streptavidin-biotin construct on a gel at all
- Protocol: mix 6 μL 2 μM streptavidin with 6 μL 1 μM c.3.2.7.2b oligo, to give a final oligo concentration of 500 nM, and incubate at room temperature for 5 min.
- dilution and electrophoresis:
- each lane contained 2 μL of different dilutions (as per table below), 6 μL of water, and 2 μL of loading dye
- lanes 7-12 were loaded with the same mixtures as lanes 1-6, respectively
| lane | amt of 1x strep-biotin mix (μL) | dilution | H2O added (μL) | final oligo concentration (nM) | amt of oligo in 2 μL diluted mix (fmols) |
| 3 | 2 | 1x | 0 | 500 | 1000 |
| 4 | 1 | 5x | 4 | 100 | 250 |
| 5 | 1 | 10x | 9 | 50 | 100 |
| 6 | 1 | 40x | 39 | 12.5 | 25 |
| 1 | control: 1 μL 2 μM streptavidin (control) to give 2000 fmols streptavidin | ||||
| 2 | control: 2 μL 1 μM oligo (control) to give 2000 fmols oligo | ||||

- ran on native polyacrylamide gel for 30 min. at 120V
- no DNA imaged with EtBr staining
- GelCode Blue staining shows a band in lane 3 with the same motility as control streptavidin in lane 1
- this is probably free streptavidin
- unclear where streptavidin-biotin complex is
- expt appears to be consistent with manufacturer's notes that the lower detection limit is 8 ng (151 fmols streptavidin, MW=52.8 kDa) [1]
Imaging biotinylated oligos
- goal: show that we can image biotinylated oligos on a PA gel
| lane | non-biotinylated DNA (Lewis' S5) (2 μM) (μL) |
biotyinylated DNA (3.2.7.2b) (1 μM) |
H2O (μL) | Tris-glycine loading dye (10x) (μL) | total amt of DNA (fmols) |
| 1 | 3 | 0 | 5 | 2 | 6000 |
| 2 | 1 | 0 | 7 | 2 | 2000 |
| 3 | 0 | 5 | 3 | 2 | 5000 |
| 4 | 0 | 3 | 5 | 2 | 3000 |
| 5 | 0 | 1 | 7 | 2 | 1000 |
| 6 | 0 | 0.5 | 7.5 | 2 | 5000 |

- run on a 12% native PA gel at 120V for 15 min.
- clear bands appear in lanes 1-4
- faint bands in lanes 5-6 are likely artifacts, as lanes 7-12 (empty) contain them as well
Brainstorming for future assay
Run two experiments in parallel, the first a control.
- assmebly
- assemble a nanostructure with outward- (1) and inward- (2) facing biotinylated oligos
- incubate assembled nanostructures with fluorescently-labeled streptavidin
- close the container lids
- baseline protein assay
- precipitate nanostructures
- pour off supernatant (containing excess streptavidin)
- resuspend nanostructures in water
- measure streptavidin concentrations in each container (pre-digest)
- these are baseline values, and we expect that they should be similar if incubation efficencies for each nanostructure are similar
- protease digest
- digest each nanostructure with protease
- particular protease and protocol must be finalized
- digest each nanostructure with protease
- measure streptavidin concentrations in each container (post-digest)
- these values should differ if biotinylated streptavidin can still be digested (container 1) and if a closed nanostructure protects its cargo (container 2)
- confirmation of presence of streptavidin
- treat both containers with DNAse
- measure streptavidin concentrations
- streptavidin is expected to be bound to biotin, but biotin not bound to oligos
- treat both containers with protease
- measure streptavidin concentrations
- values should all be close to zero
Container 4.0
Reagents (each expt in respective 0.2 mL PCR tube)
| Experiment | Scaffold | Oligos | Folding buffer | dH2O | total volume |
| -latches -aptamers |
9 μL p7308 | 16 μL xxx (250 nm) | 4 μL 10x | 11 μL | 40 μL |
| +latch1 -aptamers |
9 μL p7308 | 16 μL xxx (250 nm) | 4 μL 10x | 11 μL | 40 μL |
| +latch2 -aptamers |
9 μL p7308 | 16 μL xxx (250 nm) | 4 μL 10x | 11 μL | 40 μL |
| 6 hb | 9 μL p7308 | 4 μL 6hb.v5 (0.99 μM) | 4 μL 10x | 23 μL | 40 μL |
| -oligos | 9 μL p7308 | - | 4 μL 10x | 27 μL | 40 μL |
| -scaffold | - | 16 μL xxx (250 nm) | 4 μL 10x | 20 μL | 40 μL |
Annealing protocol
- start at 80[[:Category:{{{1}}}|{{{1}}}]]
- 60 cycles: wait 2 minutes, decrease 1[[:Category:{{{1}}}|{{{1}}}]]
- hold at 4[[:Category:{{{1}}}|{{{1}}}]]
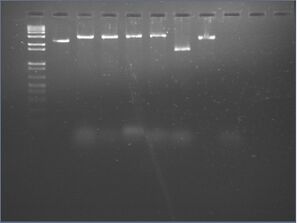
Gel analysis
- 2% agarose gel supplemented to 10 mM MgCl2 and with 3 μL 10 mg/mL EtBr (100 mL gel)
- run in 1x TBE supplemented to 10 mM MgCl2
- 45 min at 130 V
| Lane | Contents | Loading Buffer (10x TBE/glycerol) |
| 1 | 1kb DNA ladder (10 μL) | 1.1 μL |
| 2 | (-) | 1.1 μL |
| 3 | Ib (10 μL) | 1.1 μL |
| 4 | 4.0 -latches -aptamers (10 μL) | 1.1 μL |
| 5 | 4.0 +latch1 -aptamers (10 μL) | 1.1 μL |
| 6 | 4.0 +latch2 -aptamers (10 μL) | 1.1 μL |
| 7 | 6hb (10 μL) | 1.1 μL |
| 8 | 4.0 -oligos (10 μL) | 1.1 μL |
| 9 | 4.0 -scaffold | 1.1 μL |
- results