IGEM:Harvard/2006/Brainstorming Papers - Valerie
Papers:
1. http://www.cs.duke.edu/~thl/papers/NanoTrack.2005.pdf
This paper demonstrated how it is possible to attach a certain array of proteins onto a 2D DNA nanogrid. The self-assembling nanogrid served as a template for the self-assembly of streptavidin protein arrays. Intersection points at the grid where they wanted streptavidin to bind had a biotin group incorporated in order to mediate the streptavidin binding onto that site. From this method, they also talked about how to make 1D DNA nanotracks.
We talked a little about binding proteins or other molecules onto a DNA nanostructure and this paper shows that this is quite possible. We might not be able to do a large lattice, but we can perhaps apply their method of attaching proteins onto our own specific nanostructure. Also, we can try to attach different proteins onto different places? The beginning of the paper mentions several applications of doing so, which is quoted as follows: "Fully programmable DNA templated protein arrays, where individual proteins can be positioned with precision and specificity, could allow us to build function protein templates with nanometer dementions for single molecule detection. Such assemblies may also prove useful for future nanoscale "factories" by positioning enzymes at desired locations according to their functions to allow sequential reactions on substrates in a flow. Precisely controlled organization of protein molecules onto periodic 2D DNA lattices could also enable us to gain insights into protein structure using 2D cryoelectron microscopy diffraction."
2. http://www3.interscience.wiley.com.ezp1.harvard.edu/cgi-bin/fulltext/110526995/PDFSTART
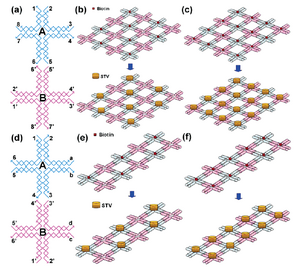
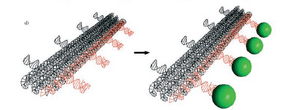
This paper is also talking about how protein arrays can be self-assembled using DNA nanostructures in the shape of a lattice. The biotin-streptavidin interaction provides only one type of protein-ligand interaction; while it is possible to fuse other proteins with streptavidin, this process is both complicated and may affect the functionality of the proteins themselves during the process. They then introduce aptamers, which are DNA or RNA molecules that have the ability to bind to other molecules such as proteins, nucleic acids, organic compounds, and organisms. There are a wide range of aptamers suited to a variety of proteins that guarantee specificity and high affinity. This method has three components: the DNA lattice nanostructure, a DNA-docking site with the aptamer sequence that will bind the protein of interest to the nanostructure, and the protein itself. In the paper, they use the thrombin-binding aptamer (TBA) which binds thrombin, a serine protease playing a major role in coagulation. When using linear DNA arrays, they noted that the arrays usually formed pairs with the attached thrombins sandwiched between, possibly due to protein dimerization.
The paper goes on to mention that another group was able to work out a system that controls the binding and release of the thrombin protein from the DNA nanostructure. Applications of this paper include the study of proximity effects of protein-protein interactions, perhaps allowing for direct visualization of such interactions due to the amplification of spatial resolution. Maybe we can steal this application idea and use it ourselves, since it sounds pretty good...The following is the paper mentioned above about the binding and release of thrombin: http://www3.interscience.wiley.com.ezp1.harvard.edu/cgi-bin/fulltext/108561908/PDFSTART.
Additional Literature:
1. http://www.cs.duke.edu/~thl/papers/SupraMolec.Chapt.12.2005.pdf
2. http://www.nsti.org/Nanotech2006/program.html
3. http://www.iop.org.ezp1.harvard.edu/EJ/toc/0957-4484/15/10
4. http://www.cs.duke.edu/~thl/pages/publications.html
New Ideas?
"As well as potential applications in the manufacture of nanostructures such as molecular electrical circuits, Professor Turberfield’s team are investigating using the DNA tetrahedra as containers for individual protein molecules: a technique which if successful could one day lead to new methods of drug delivery." if only they actually told us how we could do this...
2. "A biologist might use DNA origami to take proteins which normally occur separately in nature and organize them into a multi-enzyme factory that hands a chemical product from one enzyme machine to the next in the manner of an assembly line."
3. using DNA nanostructures to transport charges (for DNA repair?) - Barton lab at Caltech: http://www.its.caltech.edu/~jkbgrp/Publications.htm