IGEM:Harvard/2006/Adaptamers/Notebook/2006-7-27
7/27
We now have three assays to try:
1) millipore nitrocellulose membranes
The protocol is described [here]. We may learn how to use this from the good people of Liu lab.
2) Biacore 3000 at the Bauer Center for Genomics
A complex piece of machinery is able to detect how well substrates are binding to a surface (to which the protein in question is bound). We spoke to Clair Reardon of the Bauer Center today; she will soon send us information about using it.
3) Streptavidin beads
Happily, the DNA aptamers to streptavidin were developed to bind streptavidin conjugated to agarose beads. The hope is that we add aptamer, wash, elute, and hopefully we will see aptamer. Hopefully.
We tried adding thrombin to a protein membrane and seeing if we can visualize DNA with EtBr. The answer is.. no. Well sort of.
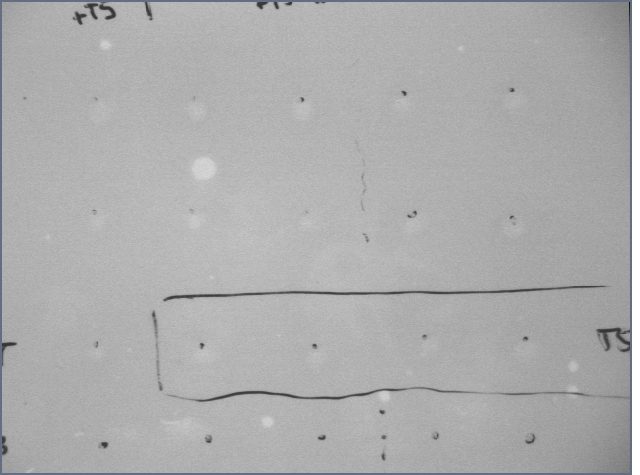
Dots are just above where I added solution.
First 3 rows of dots have thrombin added to them; last row is blanks
DNA was added to all dots. For rows 1,2,4: T5 added to column 1, 2, 3; S5 added to column 4, 5. For row 3, T20+S20 added except for column 1, where T5 was added. (this was a pretty informalexperiment)
Although there are slightly lighter spots under the transUV in the first few lanes, there is no distinction for the type of aptamer which is added. Furthermore, the UV light might just be highlighting certain areas that had more solution added to them instead of EtBr. Regardless, you can't see much.
In light of the secondary structure complications of the thrombin aptamer, I mixed together T20 and S20 to see if the complex, which no longer has the unfortunate binding site concealment, could bind to thrombin.
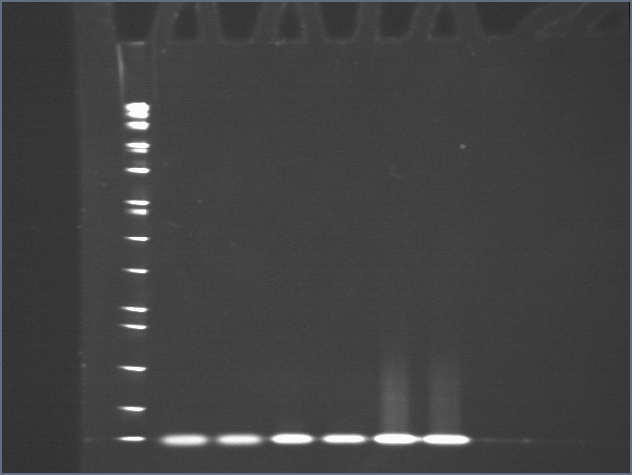
Lane 1: ladder
Lane 2: T20
Lane 3: T20 + thrombin
Lane 4: S20
Lane 5: S20 + thrombin
Lane 6: T20+S20
Lane 7: T20+S20+ thrombin
Oddly enough, we saw no gel shift for T20+S20, which we should have seen as demonstrated previously. While it is unclear that our adaptamer had actually formed, what is clear is that thrombin remained unbound.