IC Bioinfs/Project1/Overview
From OpenWetWare
Jump to navigationJump to search
OVERVIEW OF PROJECT
The project has 3 main sections-
Design of a relational database for data storage
- Designing a relational database to store the data- BioSQL relational model to comply with standard schema.
Implementation of DAS to visualize sequence data:
- Proserver(PERL)
- Dazzle (Java)
Development of a DAS compliant viewer to visualize the data:
- Flex
- GenoViz
Google App Engine
- GAE Home Page - links to download, "getting started" guides and other stuff.
PROJECT PLAN
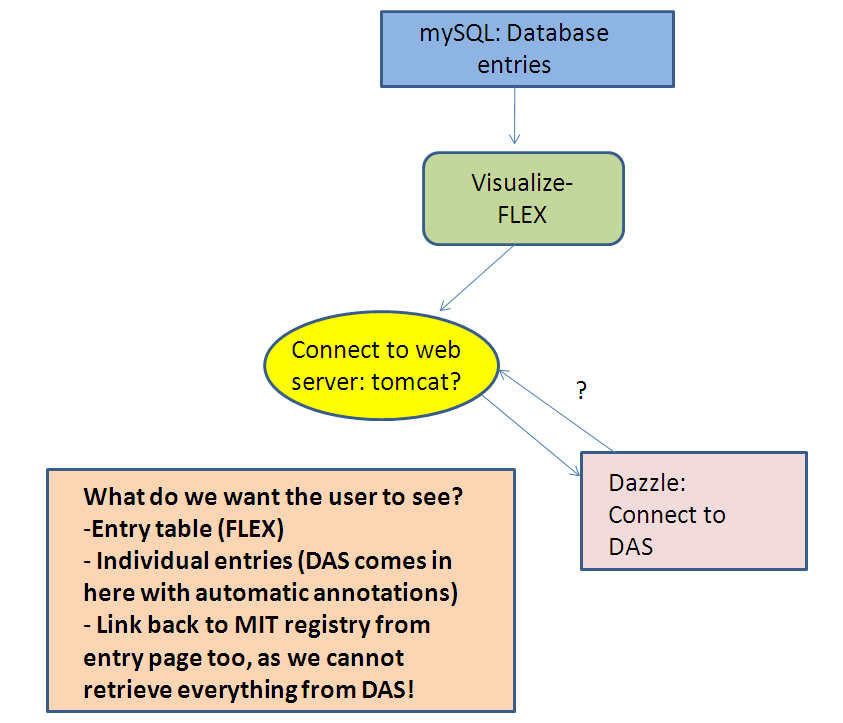
Derek's Model
As drawn on Feb 12th, during a group meeting.
The arrow from BioSQL to App Engine suggests the possibility of importing our tables to the Datastore.
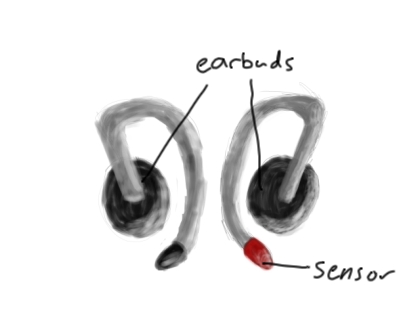
Older Model (?)
Temporary aims
- We would like to use DAS to improve on current annotation tools of the MIT registry, by being able to access external sources.
- The current MIT registry does not allow for more complex queries and filters, so we would like to improve on this by allowing for more refined searches
- Including an option to BLAST in and out of the registry will be highly useful too. The JBE allows the user to blast inside the registry.