Haynes:PCR clip clone
by Karmella Haynes, 2015
Principle: The insert is PCR-amplified using an error-reducing, high-fidelity polymerase (Phusion) and primers that add restriction sites to each end of the insert. The PCR product is cleaned up using a DNA-binding column. A predetermined amount of clean DNA is cut at the ends (or "clipped") and the 5'phosphates are removed in a single 20 μL-reaction. Dephosphorylation prevents clipped ends from re-ligating. Enzymes are deactivated at 95°C. Slow cooling after this step allows any denatured DNA to re-anneal. A small amount of the reaction is used directly as insert in a ligation. The vector that is used MUST carry 5' phosphates because the insert has no phosphates.
Plan the DNA Ligations
In your notebook, make a list of all insert+vector and vector-only controls that you will set up ligations for. Include...
- a short name for the construct
- a short name for the insert, what it will be cut with (in parentheses), and its length in bp
- a plus sign followed by the vector, what it will be cut with (in parentheses), and its length in bp
EXAMPLE List: In line 1, the short name for the construct is AubR/MRV. The insert is Aub-PCR cut with EcoRI and XbaI, and it is 851 bp long. The vector is MRV, cut with EcoRI and XbaI, and it is 3201 bp long. The last line is the negative control. There is only one negative control because only one vector is being used for all of the ligations.
|
Phusion PCR
- Design insert amplification primers that have the appropriate restriction sites on the ends. The restriction sites must be ABSENT from the insert sequence itself.
- Set up a Phusion PCR reaction as shown in the table and the Thermal Cycler Program below. Use the table as a template to record your work in your notebook.
- Note: Use 50 - 100 ng of template DNA if the template is a plasmid. Use more DNA if the specific target is at a low copy number (e.g. GAPDH gene in genomic DNA, ACTB gene in cDNA library, etc.).
- Run 5 μL of each reaction on an agarose gel to check for successful amplification.
| Reagent | Volume | Expected: 1. Insert 1 = ### bp 2. Insert 2 = ### bp 3. etc. |
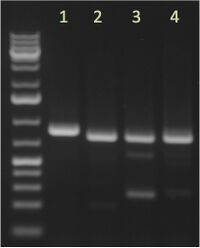 Example Image from 04/17/15; 5 μL/lane; 1% agarose; Ladder |
| DNA template* | 0.5 μL | ||
| 10 μM F primer | 1.0 | ||
| 10 μM R primer | 1.0 | ||
| 10 mM dNTPs | 1.0 | ||
| Phusion Pol. | 0.5 | ||
| 5X HF buffer | 10.0 | ||
| dH2O | 36.0 | ||
| 50.0 |
Thermal Cycler Program: (Note: set annealing temperature based on the Tm of your primers. The 72°C extension period depends upon the length of the amplified product. For products 1 kb or less, 30 seconds is sufficient. For products larger than 1 kb, use 30 more seconds per additional kb)
- 98°C, 3 min
- 35x[98°C, 10 sec; ##°C, 30 sec; 72°C, ## sec]
- 72°C, 10 min
- 4°C ∞
Clean up PCR Product(s)
- Use the Qiagen Qiaquick PCR Purification kit (cat. no. 28104 ) to purify the remainder of the amplified DNA (45 μL).
- In the final elution step, use 30 μL of elution solution.
- Measure the concentration(s) of the sample(s).
- Use the table below as a template to record your results in your notebook.
| Sample | OD260 | 260/280 | ng/μL |
| 1. Insert 1 PCR | ### | ### | ### |
| 2. Insert 2 PCR | ### | ### | ### |
| 3. etc. | ### | ### | ### |
Digest & Dephosphorylate the Insert(s)
- Set up restriction digest/ phosphatase reactions as shown in the table below
- FD = Fermentase FastDigest buffer/ enzyme
- SAP = shrimp alkaline phosphatase
| Reagent | Volume |
| DNA (500 ng) | up to 15.0 μL |
| 10X FD buffer | 2.0 |
| FD EcoRI | 1.0 |
| FD XbaI | 1.0 |
| Roche SAP | 1.0 |
| dH2O | x μL |
| 20.0 |
Thermal cycler program: Note: In the Haynes lab, use the LabNet OptiMax Thermocycler, Program "AnOlig RD"
- 37°C, 10 min
- 95°C, 5 min
- Ramp down to 25°C, 5°C/1 min. [90/1 min, 85/1 min, 80/1 min, ... 25°C/1 min]
- 25°C, ∞
Your insert is now ready to use. The final concentration is 25 ng/μL.
Perform Ligations & Transformations
- Set up the following ligation reaction(s) in sterile tubes.
Note: The vector that is used MUST carry 5' phosphates because the insert has no phosphates. - Transform competent E. coli using the appropriate/ preferred procedure. Both the 5-minute quick transformation (using DH5α-turbo cells) and longer transformation protocols work.
| Ligation | Neg. Ctrl. | |
| Insert DNA | ## | --- |
| 5'p Vector DNA (25 ng) | ## | ## |
| 2x lgn buf (Roche) | 5.0 | 5.0 |
| T4 ligase (NEB) | 1.0 | 1.0 |
| dH2O | --- | 3.0 |
| 10 μL | 10 μL |
--> Room temperature 5 min.