Haynes:Luciferase
Cell Lysate Luciferase Assay
Principle: Cells expressing the luciferase reporter gene are lysed to release the luciferase enzyme into solution. D-luciferin substrate is added to the solution. Free luciferase enzyme then catalyzes a reaction that generates light (bio/chemiluminescence). In this application, the light signal serves as a proxy for the amount of luciferase gene expression. Note that data on individual cells is lost using this procedure, but it is useful for measuring relative average levels of reporter gene activity in different cell samples.
Kits we use in the lab include:
- Biotium - Steady-Luc Firefly HTS Assay Kit #30028-1 (kept at -80°C, thaw buffer at room temp before use!)
Below is a general protocol to help you plan/ prepare for a luciferase assay. This protocol includes a cell counting step so that luciferase signal can be normalized as luciferase per cell. This way, differences in luciferase signal will not be an artifact variations in cell input.
CELL CULTURING & HARVESTING
- Grow cells in a 6-well culture plate in standard growth medium.
- When cells are 90 - 100% confluent, there will be enough cells for the following processing steps.
- Harvest cells by standard trypsinization:
- Wash cells with 1.0 mL warm 1x PBS.
- Add 0.5 mL warm Trypsin-EDTA medium. Incubate at room temp for 2 - 4 min.
- Collect cells with 3.5 mL warm growth medium to deactivate the trypsin. Be sure to detach cells by pipetting up and down to detach the cells from the growth surface. Transfer resuspended cells into labeled 15 mL conical tubes.
- Pellet the cells at 1000 rpm for 3 min.
- Carefully aspirate off the medium while leaving ~50 uL medium coverign the pellet.
- Hold the tube firmly and flick the botom to break up the pellet.
- Resuspend cells in 2 mL cold FACS buffer (10% FBS in 1x PBS). Place tubes on ice.
- Add 1 mL of FACS buffer to an extra tube. This will be used to calculate background signal for the luc activity assay.
LUCIFERASE ACTIVITY ASSAY
- Prepare enough complete luciferase assay buffer for 3 replicates for all samples, 100 uL per replicate.
- Biotium kit Calculation 1: [ (# samples x 3 replicates) + 2 for piepetting error] x 100 μL assay buffer = total buffer to prepare.
- Biotium kit Calculation 2: (Total buffer to prepare / 100) x 2.5 μL 10 mg/mL D-luciferin stock = total D-luciferin stock to add to assay buffer
- Add total buffer to prepare into a 15 mL conical (if over 1.5 mL). Add total D-luciferin stock to add to assay buffer to the assay buffer. Cap tightly and mix. Shield the prepped buffer from light (simply place the tube under a cardboard lid, OR wrap in foil.
- Filter 600 μL cells from each 2 mL experimental sample through strainer caps.
- Transfer 3x 100 μL strained cells to 3 wells in 96-well plate (white opaque, flat bottom). Save remainder of strained cells on ice for flow cytometry/ counting.
- Load plain FACS buffer into three wells as a background control.
- Very quickly mix (by pipetting) 100 uL luciferase activity buffer into each 100 uL cell sample.
- Incubate the reaction at room temperature for the recommended time.
- Use the 96-well plate reader to measure the chemiluminescent signal. Three reads is recommended to capture output at maximum or steady state.
The current recommended program for reading luciferase activity on the Synergy H1 reader is:
- Protocol: KAH_luciferase
- Optimized specs:
- Double click 'Procedure' to edit
- Plate Type: 96 WELL PLATE
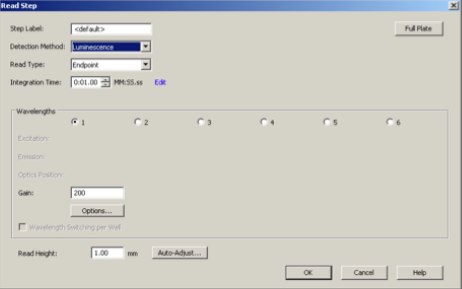
- Under 'Actions' click 'Read' to add a read step. Check the following:
- Detection method: Luminescence
- Read type: Endpoint
- Integration time 0:01:00 MM:SS:ss
- Wavelengths: 1
- Gain: 200
- Read height: 1.00 mm
- Use the Wang lab's Accuri flow cytometer. Be sure to check if anyone is using the cytometer while you plan to use it.
- Set machine to read 20 uL of cells.
- Set up the software to create a histogram of forward scatter (x-axis) vs. cell count (y-axis).
- "Clean" the cytometer with a non-cell water sample. When the cytometer reads close to zero events in water, it is ready to run the next cell sample.
- Flick the tube of strained cells to make sure that the cells are not settled at the bottom.
- Place the tube of strained cells onto the sample holder and run the sample.
- Gate the data to include only only the highest peak of cells. Use the exact same gate for all samples!
- Record the cell count per 20 uL in a note book.
- "Clean" the cytometer with a non-cell water sample. When the cytometer reads close to zero events in water, it is ready to run the next cell sample.
- Repeat the previous five steps for the remaining samples. Fill in a table like the one below.
| Sample | count/ 20 μL | cells / 100 μL |
| Sample 1 | value 1 | 5 x value 1 |
| Sample 2 | value 2 | 5 x value 2 |
| Sample 3 | value 3 | 5 x value 3 |
FINAL DATA PROCESSING
- Subtract the highest background value from all of the individual experimental values.
- Do the following calculation for each experimental luciferase value:
Sample 1 luciferase per cell = [Sample 1 Luciferase signal] - background signal / [cells / 100 μL]
Repeat this calculation for the remaining samples, using the appropriate cell counts for each unique sample. - Calculate the average of the three replicates for each sample, and calculate standard error.
- Plot this data as a histogram or dot plot, with positive and negative error bars.