Haynes:GOEnrichment
Intro: Gene Ontology
So you discovered that a set of genes all become activated when you treat cells with a drug. What do the genes "do?" How will the phenotypes of the cells change as a consequence of activating these genes?
To help answer such questions, a group of scientists built a large list of standard terms to describe the functions of genes. It's very important to have a standard vocabulary, especially when many scientists are sharing information. For instance, one scientist might write about "secretion of extracellular matrix proteins" while another, who is studying the same gene reports the function as "cell surface matrix component delivery." It is important to establish which phrase is acceptable, especially when most scientists now days are working with hundreds and thousands of genes that all need to be described.
Another interesting problem...when more than one gene cooperates to control a single function, if the function has many different names, then it is hard to correctly classify the genes into a single functional group.
"The Gene Ontology (GO) project is a collaborative effort to address the need for consistent descriptions of gene products across databases." Read more at the Gene Ontology Consortium home page at http://geneontology.org/
The three major categories of the Gene Ontology are:
- "Biological Process" - describes the process in which the gene product is involved
- "Molecular Function" - describes the biochemical function of the gene product
- "Cellular Component" - key cellular structure(s) that contains the gene product
Tool: GOrilla
Intro: These instructions will help you to use the Gene Ontology enRIchment anaLysis and visuaLizAtion tool (GOrilla) to search for enriched GO terms in a target list of genes compared to a background list of genes. The software searches for GO terms that are enriched in the target set compared to the background set using the standard Hyper Geometric statistics. Significant enrichment of a certain GO term suggests that your specific group of genes is associated with some biological process, and that this association is not just by chance.
Procedure:
- Go to http://cbl-gorilla.cs.technion.ac.il/
- Set "Choose organism" to the relevant organism (e.g., Homo sapiens = human, Mus musculus = mouse)
- Set "Choose running mode" to "Two unranked lists of genes (target and background lists)"
- In the "Target Set" field, paste or upload a list of genes that you want to analyze. txt format, one gene symbol per line, is recommended for the upload option
- For the "Background Set," copy-paste or upload a complete list of all gene symbols for your organism. Use your own or one of the following:
- Human genes - GOBg_Human_092014.txt
- Mouse genes - GOBg_Mouse_092014.txt
- Set "Choose an Ontology" to one of the following three options. It is recommended that you run an analysis for each separately (do not select "All") for publishable results...
- "Process" - is "Biological Process"
- "Function" - is "Molecular Function"
- "Component" - is "Cellular Component"
- Click the "Search Enriched GO Terms" button to run the analysis.
- After processing the results, use the back button on your browser and repeat the analysis with a different "Choose an Ontology" setting.
Results:
- The analysis outputs three important types of data:
- A GO term hierarchy tree, where GO terms are shown in boxes connected with lines. Most GO terms are specific sub-classes of parent terms.
- The color scale indicates P-values. The P-value represents the likelihood that the enrichment value for that GO term could be the same for a random list of genes. Therefore, the smaller the P value, the more significant the enrichment.
- A ranked table, where the GO terms with the smallest P-values are at the top. Click the "Show Genes" link to see the gene symbols that are associated with the GO term in that row.
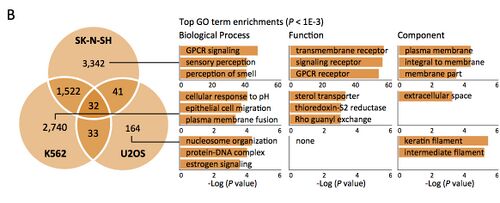
How to Make Bar Charts - small P-values are converted into positive numbers for easy-to-interpret comparisons
- Run an analysis for "Process" and get results.
- Open an Excel spreadsheet.
- Make a table like the hypothetical example below.
| Target list | GO Category | Term ID | Term | P-value | No. genes | Neg Log 10 |
| U2OS | Process | GO:0007186 | G-protein coupled receptor signaling pathway | 1.38E-47 | 340 | 46.86012091 |
| GO:0050907 | detection of chemical stimulus involved in sensory perception | 4.48E-41 | 176 | 40.34872199 | ||
| GO:0050911 | detection of chemical stimulus involved in sensory perception of smell | 7.91E-41 | 165 | 40.10182352 | ||
| Target list = description of your input list GO Category = what you selected for "Choose an Ontology" Term ID = "GO Term" from the result table Term = "Description" from the result table P-value = "P-value" from the result table No. genes = the value of b Neg Log 10 - Use this formula for the value of these cells: =-(LOG(#P-value#,10)) ...where #P-value# is the cell that contains the P-value | ||||||
...cont.
- Add more data as necessary. Three to Five top GO terms are sufficient.
- In a new column next to Neg Log 10, replicate the Neg Log 10 values (use the "=" function, or copy-paste the values). You will convert these into horizontal bars (next steps).
- With these new cells selected, open Format > Conditional Formatting...
- In the dialogue box, click "+" to add a new rule
- Set style to "data bar", minimum type = "number", minimum value = "0", maximum type = "number", maximum value = the highest rounded value of Neg Log 10, and select "show data bar only"
- Click "okay"
- You should see horizontal bars that correspond to the Neg Log 10 values
- Repeat this entire procedure for "Function" and "Component", adding the new data to your spreadsheet, with the appropriate labeling to keep the results organized.
- You can select the cells with bars and "paste-as-pdf" into Powerpoint to incorporate them into a final image. In the example below, Dr. Haynes used Powerpoint drawing objects and text boxes to add axes and numbering.