Haynes:BioBrick Method short
Assembly of BioBrick Parts: an Overview
gaattc gcggccgc a tctaga [BioBrick Part] actagt a gcggccgctgcag
EcoRI NotI XbaI SpeI NotI PstI
BioBrick Prefix BioBrick Suffix
Note: At the BioBrick Suffix, the NotI and PstI cut sites overlap at the underlined “c.”
The BioBrick itself must not contain any of the standard restriction sites (internally), or else it will be cut apart during the assembly process. Basic BioBrick assembly uses one of the two strategies:
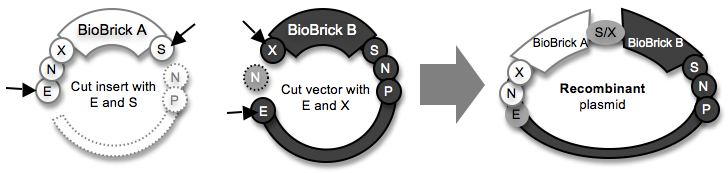
Ligating a “front insert” in a “front vector” to make A+B
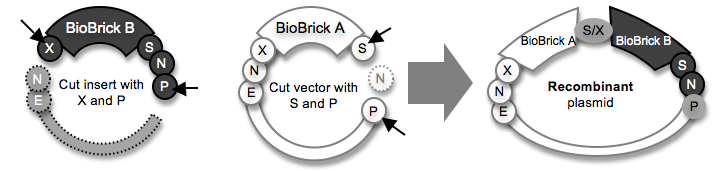
Ligating a “back insert” in a “back vector” to make the same A+B construct
Note: After DNA plasmids are cut, the desired DNA fragments are isolated and purified using agarose gel electrophoresis (ref). The fragments that are bordered by a dotted line are discarded.
A few important words about DNA assembly: In the diagram above, all of the DNA starts out as a plasmid (circular piece of DNA). Each plasmid has a region called a backbone (shown as a thin arc) that consists of the origin of replication (the DNA sequence that encodes information for making copies of the plasmid; the mechanism is very complex) and an antibiotic resistance gene (e.g., ampicillin resistance). After the plasmids are cut with restriction enzymes (ref), one DNA fragment is used as an insert (no backbone) and another is used as the vector (has the backbone). The final DNA ligation product (the recombinant plasmid) must have one backbone.
About the "S/X" scar sequence: When two BioBricks are ligated, the SpeI end ligates with the XbaI end. The resulting sequence is a mixed site (called a scar) that cannot be re-cut by any enzyme. Silver Standard assembly produces a 6 b.p. scar (two codons) so that proteins can be fused together without creating a frame-shift mutation.