Generating Distribution Charts and Cumulative Plots for GRNmap Weight Values in SPSS
This page outlines how to generate histograms and cumulative plots showing the distribution of regulatory weights present in Gene Regulatory Networks (GRNs) modeled with GRNmap. Specifically, the protocol written below was used to visualize the weight distributions for db1-db6 using SPSS. The results of this analysis can be found here.
Note: The instructions below were written using SPSS Statistics Version 21.
Step 1: Preparing the Data for Input into SPSS
Prior to beginning this analysis, use GRNmap to model the dynamics of the networks that you would like to analyze. Once this has been done, extract regulatory weight values from the "network_optimized_weights" tab found in the GRNmap output sheet. Although these values can be extracted using various different methods, you want to end up with a single column containing each of the weight values. One technique would be to transform the weighted adjacency matrix found in the "network_optimized_weights" tab into an edge list. Once this process has been completed, paste the single column containing all of the regulatory weights from the network into column A within a new excel file. Label the first sheet in this workbook "weight_value_conversions". Label cell A1 with the name of the network from which the weight values were derived. Then highlight column A and navigate to "Sort & Filter"-> "Sort Smallest to Largest". Assess the smallest and largest values present at the top and bottom of the column, respectively. Determine which value has the largest absolute value and note the designation of this cell (e.g. A29). Label cell B1 with the name of the network followed by "_scaled_normalized". In cell B2, write the formula: =(A2/ABS($A$__))*99.99. Replace the underscores with the number of the cell documented previously that contained the regulatory weight with the largest absolute value. Then press enter. In cell B3, repeat this process and also change A2 to A3 in the formula. Then press enter. Now highlight cells B2 and B3. Double click the black square at the bottom right hand corner of the highlighted region to automatically extend this formula for scaling and normalization to the remainder of the weight values. Doing so should leave you with column B having been expanded to match column A in length and containing values ranging from -99.99 to +99.99. This process normalized all of the regulatory weights to the largest weight value in the network and then multiplied by 99.99 so that the weights will be distributed from -100 to 100 in the final graphics.
Create a second sheet in the excel workbook and label it "SPSS_input". Copy the contents of column B. Then right-click on cell A1 in the new "SPSS_input" sheet and select "Paste Special..."-> "Values". This will copy the numbers from column B over without copying the formulas existing internally within the original cells. Now label cell B1 with the network name followed by "_coding". In this column, enter a 0 if the weight value in the adjacent cell of column A is negative or enter a 1 if this adjacent regulatory weight is positive. Alternatively, this coding can be expanded to include a third value if you would like to show low influence regulatory weights (labeled grey in GRNsight). Once the coding column has been completely filled out, the data is ready to be imported into SPSS.
Step 2: Generating the Weighted Histograms
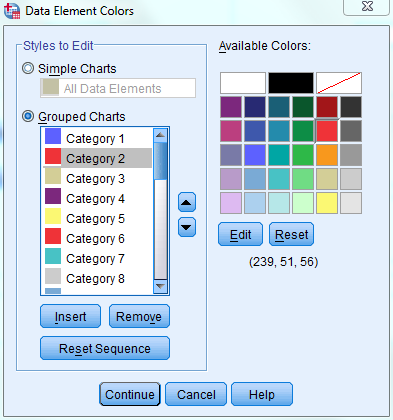
Launch SPSS, allowing the program several minutes to load. To open the excel file containing the data to be analyzed, navigate to File-> Open-> Data... A window will open prompting you to designate the path to your file. Navigate to the folder containing your file. By default, only .sav files will show up in the selection window. Change the file types displayed to "All Files" or "Excel" so that your excel file will be visible. Once it is, select the file and then click "Open". Another window will open asking you to designate the worksheet containing the data you would like to import. Select the worksheet labeled "SPSS_input" from the drop down window and click "OK". This will open your file in the SPSS data editor. Note that variable names and number formatting may be altered in the import process. To edit this formatting, navigate to the "Variable View" tab visible at the bottom left corner of the data editor. Although most of the formatting options are only for aesthetics, it is critical to check the "Measure" column to select the appropriate data type for each variable. Specifically, the "scaled_normalized" variable should be labeled as scale and the "coding" variable should be labeled as nominal. Further, you will need to specify the values for the coding variable. To do so, click on the cell in the same row as the coding variable and in the values column (should be labeled "None"). Then click on the button that appears in the cell labeled "..." to open the Value Labels window. In the window, enter your first value as 0 and enter the label as "Repression". Then click add. Now enter your second value as 1 and enter the label as "Activation. Click add again. Then select "OK" from the bottom of the window. Next, change the graph color coding options so that regulatory weights indicated as 0/Repression will be labeled blue and weights indicated as 1/Activation will be labeled red as in GRNsight. To do so, navigate to "Edit"-> "Options...". In the Options window, select the Charts tab from the top right and then click on the "Colors..." button. Select the "Grouped Charts" option within the "Styles to Edit" box. Then click on "Category 1" and select the blue color from the "Available Colors" menu to the right. Now click on "Category 2" and select the red color from the "Available Colors" menu. Once you have done so, your window should look like the sample provided below. If it does, click "Continue" and then "Apply" in the original Options menu. Now click "OK" to return to the data editor.
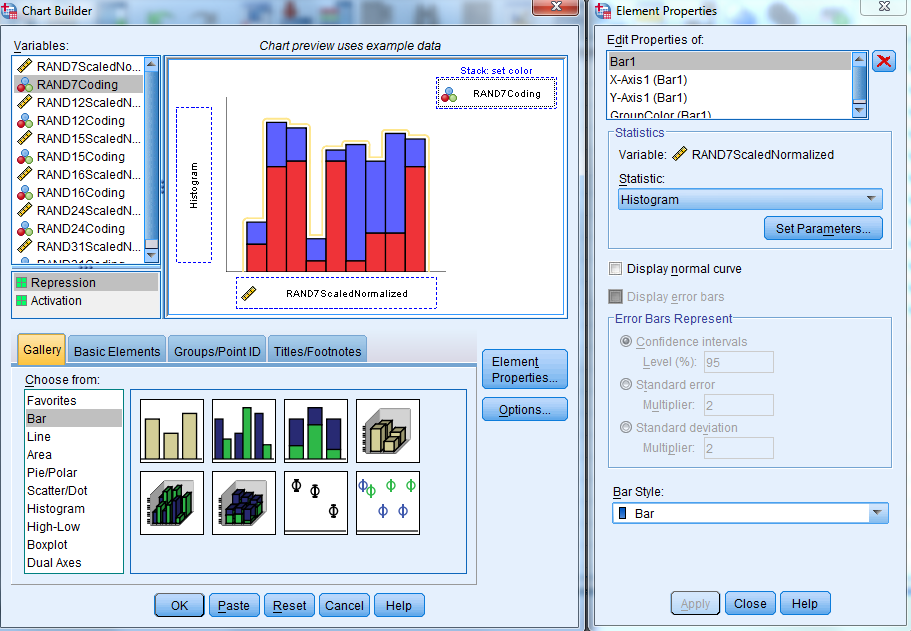
Now that the data and color coding has been properly formatted, it is time to generated the weighted histograms. To do so, click on "Graphs"-> "Chart Builder..." within the data editor. Click "OK" when presented with a popup to open the Chart Builder. In the gallery at the bottom of the window, click on "Bar" in the "Choose from..." box. Then double click on the "Stacked Bar" option to launch this format. Now click on the "Element Properties..." button to the right of the Gallery to open a second window. In the "Statistics" box, click on the drop down menu under the "Statistic:" option and select "Histogram". Next, click on the "Set Parameters..." option under the Histogram statistic designation. This will open a new window. In the "Bin Sizes" box, click on "Custom". Under "Custom", select "Interval width:" and enter the number "20" in the box to the right. This will fix the width of each bin to 20, which will fix the total number of bins to 10 given our -100 to 100 scale of weight values. Click "Continue" at the bottom of the "Element Properties: Set Parameters:" window. Finally, click "Apply" at the bottom of the "Element Properties" window. In the "Variables:" box in the upper left hand corner of the Chart Builder window, select the variable labeled "scaled_normalized". Then drag this variable into the box on the right labeled "X-Axis?". Now click on the variable labeled "coding" and drag it into the box on the right labeled "Stack: set color". Once this has been done, the Chart Builder window should look similar to the sample provided below.
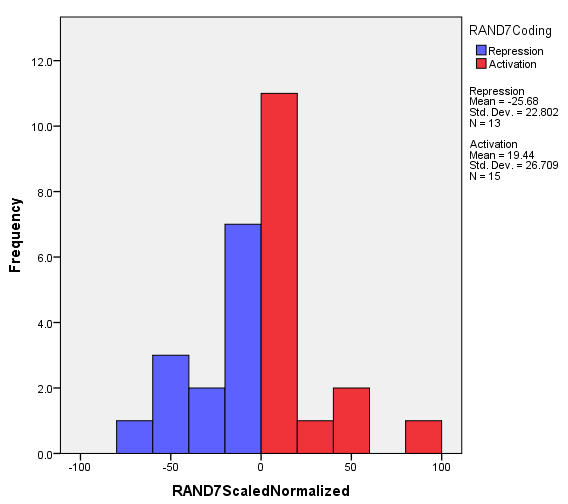
Now click "OK" at the bottom of the Chart Builder window to generated the weighted histogram. A sample weighted histogram produced for the network "RAND7", which is a random network derived from db5, is shown below.
Step 3: Generating the Cumulative Plots
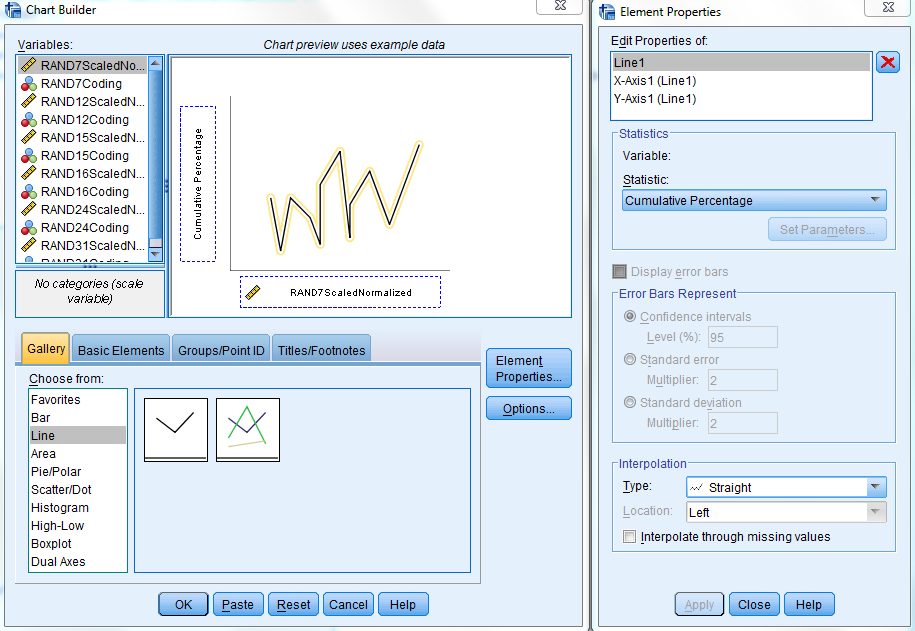
Next, to generate the cumulative plot, reopen the Chart Builder. In the Gallery tab, select "Line" in the "Choose from:" box and then double click on the "Simple Line" option. Click on the "Element Properties..." button to the right of the Gallery tab if necessary to reopen the Element Properties window. Within this window, click on the "Statistic:" drop down menu and select the "Cumulative Percentage" option. Then click "Apply" at the bottom of the window. Now select the "scaled_normalize" variable from the "Variables:" box in the Chart Builder and drag it to the box labeled "X-axis?" on the right. At this point, the Chart Builder window should look similar to the sample below.
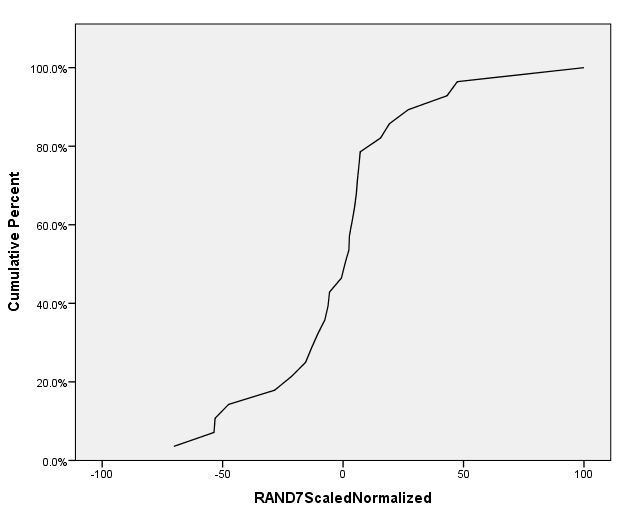
Now click "OK" at the bottom of the Chart Builder window to generate a cumulative percentage plot of the regulatory weight values within the network. A sample cumulative plot produced for the "RAND7" network is shown below.