Fong:Quantification
From OpenWetWare
Jump to navigationJump to search
<owwmenu font="arial, helvetica, sans-serif" bold="1"
color="White" bgcolor="gray" hovercolor=#FF9933
bghovercolor="black" topfontsize="7" fontSize="7" pagewidth="750"
image="Cth.JPG" lab="Fong">
Home=
Lab Members=People
Research=Research
Internal=Internal, Protocols=Protocols, Chemical Inventory=Inventory
Links=Links
</owwmenu>
Overview
This procedure is used to quantify carbohydrates in solution. Primarily, this is used to observe cellobiose or glucose uptake rates of various organisms including C. thermocellum and T. fusca.
Materials
- Samples (
- 96-well PCR plate & cover
- 96-well Spectrophotometry plate
- Multi-channel pipette*
- Buffer
- DNS reagent (use gloves & lab coat)
- Prepared standard curve solutions
- PCR Thermocycler
- Multi-well spectrophotometer
*Not required, but certainly handy.
Procedure
- Thaw samples and transfer 1 mL to a 1.5 mL centrifuge tube.
- Return samples to -80°C freezer.
- Spin down samples (max RPM for 3 minutes).
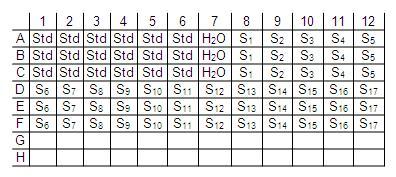
- Add 20 μL of standard solution (in triplicate) to the 96-well PCR plate.
- Add 20 μL H2O in triplicate as a zero reference.
- Add samples in triplicate.
- Add 40 μL of buffer to each well used.
- Add 120 μL of DNS reagent to each well.
- Incubate plate at 95°C for 5 minutes (use thermocycler).
- Cool on ice for ~5 minutes.
- Add 36 μL of sample to 160 μL H2O on the spec. plate.
- Read the plate at 534 nm.
The data cannot be easily copied from the imaging software into Excel, so it's better to print out the data page and manually enter each point. Calculate the standard curve of g/L free sugars per fluorescent unit. Multiply this value by the average of each sample triplicate to calculate the g/L free sugars at that sample point. Uptake rate should be calculated in mid-log phase.