Endy:High-efficiency electroporation
From OpenWetWare
Jump to navigationJump to search
some notes:
- ligase supposedly inhibits transformation of dna, so it should always be heat-inactivated at 65 C for 10 min
- NEB sells T4 DNA ligase at two concentrations: 400,000 units/ml and 2,000,000 units/ml. Jen Braff was using the 2,000,000 units/ml ligase and found that a 1:5 dilution resulted in about 30x more colonies. Also mentions phenol extraction/precipitation to improve transformation efficiency, from JGI library protocols (?). (all in email to Jason)
- NEB offers a Quick Ligation Kit, with a buffer containing PEG. PEG when heat-inactivated inhibits transformation, so the ligation has to be purified instead. Endy Lab has the normal ligation kit without PEG, so shouldn't have to worry about this.
- comparison of undiluted vs. 1:10 dilution of ligase (using 400,000 units/ml ligase stock) for one of my libraries showed about 3.5x more colonies for the undiluted ligase (see my notes for August 23)
- amount of DNA
- NEB recommends between 1-10 ug/ml of DNA in a ligation reaction, so originally I was trying to shoot for somewhere in the middle. I wasn't getting very many colonies (a few at most), so I tried digesting more DNA and ligating higher concentrations, which improved things a little. Found that DNA recovery efficiencies from gel extractions were much lower than expected.
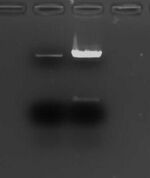
on the left is 0.6 ul (out of 30 ul elution) of overnight digested, gel extracted backbone (10 ug); on the right is 1 ul (out of 50 ul digest reaction) of 1 hr digest (10 ug) - Ended up just PCR cleaning the digested backbone (P1010 insert) and the digested PCR'ed insert and using as much as possible in a 20-ul ligation reaction (~2.8 ug backbone + ~0.85 ug insert, way over maximum NEB recommendation) and ended up with libraries between 1e3 and 4e4, which was comparable to what it was using gel-extracted backbone
- NEB recommends between 1-10 ug/ml of DNA in a ligation reaction, so originally I was trying to shoot for somewhere in the middle. I wasn't getting very many colonies (a few at most), so I tried digesting more DNA and ligating higher concentrations, which improved things a little. Found that DNA recovery efficiencies from gel extractions were much lower than expected.