ESSCOSMOS/2009:Enzymes
Based on Steve Allison's Colormetric Enzyme Assay Protocol
Overview
White-rot fungi produce oxidative enzymes that are capable of decomposing a wide range of aromatic compounds and may improve digestion in animals. These enzymes may be abundant in the substrate on which edible white-rot fungi are cultivated. If so, the used substrate that is left after harvest could be a potential source of these enzymes for bioremediation and nutrition.
Paul Stamets provides an overview of this and other potential uses of white rot fungi in his ambitious lecture "Six ways mushrooms can save the world"
Objectives
The specific objectives of this laboratory exercise are to:
- Determine the activity of two oxidative enzymes, PPO and peroxidase, in substrate that is produced as a by-product of mushroom cultivation.
- Compare the production of these enzymes by three different mushroom species.
- Develop ideas about how spent substrate could be used in bioremediation.
Background
Materials
- Cuvettes: three per sample
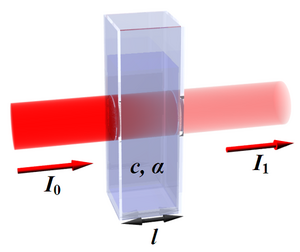
Absorption of a beam of light traveling through a cuvette with width ℓ. [math]\displaystyle{ A = -\log_{10} \left( \frac{I}{I_0} \right) }[/math] - Spectrophotometer
- Pipettes
- Mushroom substrate from three mushrooms: Pleurotus eryngii, Hypsizygus marmoreus, and Grifola frondosa that was collected on 7/18/2009 from Golden Gourmet Mushrooms / Hokto Kinoko Company in San Marcos, CA.
Reagents
Buffer
50 mM sodium acetate, pH 5.0
Enzyme Substrate
25 mM L-dihydroxyphenylalanine (L-DOPA)
WARNING: L-DOPA is hazardous; wear gloves.
Homogenate
For each mushroom species:
- Combine 2 g sample and 100 mL buffer in a blender and blend for one minute; pour homogenate into a flask labeled with the mushroom species name.
Hydrogen Peroxide
0.3% H2O2
Methods
- Make one substrate control
- For each species, make three cuvettes, one for each assay and one for the homogenate control (click link or see below for details):
- homogenate control for homogenate absorbance
- substrate control for L-DOPA absorbance
- assay for polyphenol oxidase
- assay for peroxidase
- For assay cuvettes (3,4), record the time at which homogenate and L-DOPA are combined.
- Cover cuvettes and invert to mix every fifteen minutes
- While you are waiting, research the different species of mushroom, and predict which mushroom species will have the highest enzyme activities based on what you find out about their habitat and physiology.
- After 1-2 hours, samples will be ready to read
- Record absorbance at 450 nm using spectrometer
Homogenate control
n=3 cuvettes
Label first cuvette "Ho" + species id, add:
- 0.5 mL homogenate
- 1.5 mL buffer
Enzyme substrate control
n=1
Label second cuvette "Su" (enzyme substrate), add:
Polyphenol Oxidase Assay
n=3
Label third cuvette "PPO" + species id, add:
- 0.5 mL homogenate
- 1.5 mL L-DOPA
Peroxidase Assay
n=3
Label third cuvette "Per" + species id, add:
- 0.5 mL homogenate
- 1.5 mL L-DOPA
- 0.1 mL Hydrogen Peroxide, 0.3%
Results
1. Enter your data in this form
2. Compute activity as μmol L-DOPA converted per hour per g sample as follows:
Activity (μmol h-1 g-1) =
OD/[(EC/μmol/ml)/(2.0 ml/assay)(incubation, hr)(g Sample/ ml sample homogenate)(0.5mL homogenate/assay)]
where:
- OD=Sample Absorbance - (Substrate Absorbance + Homogenate Absorbance)
= PPO - (Ho + Su) or = Per - (Ho + Su)
- Extinction Coefficient, EC = 0.403
3. Peroxidase activity is the difference in activity between the PPO and peroxidase assay.
Analysis
- Which species has the highest PPO activity?
- Which species has the highest Peroxidase activity?
