Davidson:The effect of plasmid copy number
From OpenWetWare
Jump to navigationJump to search
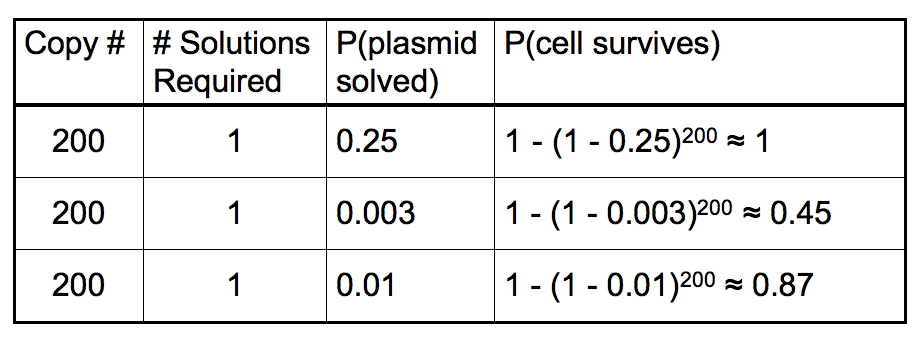
Cell survival in our selection process will require one or more plasmids to be sorted in the correct order and orientation. We used the binomial probability distribution to model cell survival as a function of number of plasmids and the probability of each plasmid being sorted. This model assumes that the flipping and sorting of each plasmid is independent of the others in the cell. In this model, the probability of cell survival is given by:
 where
where
- n is the plasmid copy number
- p is the probability of a single plasmid being sorted in the correct order and orientation
- x is the number of plasmids that must be sorted for the cell to survive the antibiotic selection
Although we do not know in advance whether a single plasmid being sorted is sufficient for cell survival, we will be able to try different values of x in the binomial equation, and compare our cell survival rate to the corresponding probabilities.