Davidson:Davidson 2005
Davidson College Synth-Aces
Link to Excel File: Nomenclature Scheme for all Davidson Parts[1]
Link to Excel File: List and description of Davidson Parts[2]
Davidson:Davidson Part Nomenclature
The Project
Students
- Nick Cain
- Mac Cowell
- Kristen DeCelle
- Andrew Drysdale
- Matt Gemberling
- Oscar Hernandez
- Tamar Odle
Advisors
- Malcolm Campbell
- Laurie Heyer
Our iGEM 2005 Presentation [3]
Goals
- Design and Creation of Four Fluorescent Proteins with LVA degradation tags
- Enhanced Yellow Fluorescent Protein
- Enhanced Red Fluorescent Protein
- Enhanced Green Fluorescent Protein
- Enhanced Cyan Fluorescent Protein
- Degradation tags are necessary on each color in order to test the effectiveness of our RNA protein regulating molecules
- Design RNA molecules to control protein the production of the Enhanced fluorescent proteins
- Anti-switches
- We have adapted Smolke and Bayer (2004) design to target Enhanced Yellow fluorescent protein and have been trying to test these designs. However we have not recieved out latest shipment from Blue Heron and do not plan on recieving it until after the competition. We plan to try and incorporate another aptamer for malachite green (found not to be toxic in low concentrations).
- Riboswitches
- Riboswitches are segments of mRNA which fold on themselves to inhibit the binding of the ribosome. The riboswitches we constructed contain an aptamer specific to the ligand theophylline. Upon the binding of the theophylline molecule, the conformation of the mRNA changes to inhibit or allow the binding of the ribosome. Utilizing the theophylline aptamer mTCT8-4 from the paper {R. Jenison, S. Gill, A. Pardi, B. Polisky, Science. Vol. 263, 1426 (1994).} we got Blue Heron to construct 3 different riboswitches designed to regulate the production of yellow fluorescent protein. All finished plasmids contain a promoter (R0010), RBS (B0034), YFP (E0032), and terminator (B0015) as well as a theophylline aptamer before or after the RBS. The article by Gallivan and Desai describes the use of a larger riboswitch which includes the mTCT8-4 aptamer within it. The function of the additional bases which Gallivan and Desai added to the aptamer is not known. Our team explored whether just the aptamer and 8 additional bases was enough to have a regulatory affect on translation of YFP.
- External Guide Sequences
- External Guide Sequences are molecules that are designed to target mRNA. The shape of the external guide sequence is recognized by RNase P and the mRNA is cleaved. We are currently designing a EGS that would contain an aptamer placed in the V-loop, between the T-loop and Anti-codon loop. The hope is that when the ligand binds a shape change will take place that RNase P will not recognize, thus turning the transcription of the EYFP "on".
- Anti-switches
The Device
Our project goal was to create a device for detecting differences between eight different combinations of three chemicals. We propose to do this through a system of Anti-switches that are controlled by 3 different aptamers. The device would give a different visual output for any one of 8 different combinations of inputs.
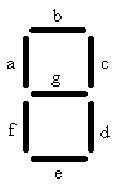
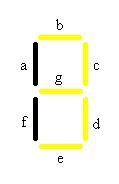
The decoder is represented by 7 strips that have the potential to form any number from 0-7, symbolizing the 8 different possible combinations of chemicals. In order to accomplish this goal we will need 8 different cell types. Each cell type number 0-7 will contain 3 different anti-switches, in one of 8 different combinations of “on” and “off” switches.
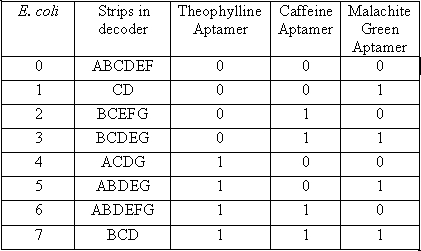
In the table a 0 represents the presence of an “off” switch and 1 represent an “on” switch. For instance when the only chemical present is malachite green, only the stripes in the decoder labeled C and D will glow, giving the visual image of one. When the number one appears it corresponds to only malachite green being present.
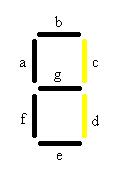
When both malachite green and Caffeine are present the number 3 will be visible will be visible. At first this may seem like a problem because both 1 and 2 should also show up in the presence of these chemicals, but remember that there are also off switches in both of these cell types. Therefore the second chemical in each case will inhibit the production of the fluorescent protein only allowing the 3 to be visible.
By following this same logic all 8 different combinations of these three chemicals can be discerned using this decoder, in a visual manner.
Each strip will be made using a combination of each of the cell type necessary to induce YFP production under each individual condition. For instance strip A will have a mixture of cells # 0, 4, 5, and 6.
Images uploaded 12-5-05 by WTH
We had reported a variability in i6060 fluorescence intensity but could not figure out why it varied. Kristen took samples of bacteria from a single plate that had different intensities, grew them, measured their fluorescence, then minipreped the DNA. As you can see from the gel, there is a good correlation between the amount of plasmid and the amount of fluorescence. All cells were grown in media from a common stock bottle, so we are not sure why they varied.
It is interesting to think that different colonies can carry different loads of plasmid and this is responsible for the difference in fluorescence.
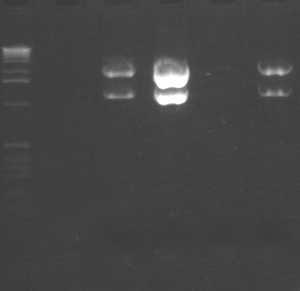
Miniprep DNA from isolates of bacteria carrying i6060 plasmid.
Lanes: 1 = MW; 2 = empty; 3 = spot 1; 4 = spot 3; 5 = spot 4; 6 = spot 6.
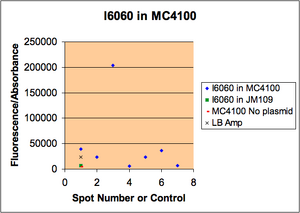
Spot numbers correspond to the spot numbers for the photo of the gel electrophoresis.