Cell sorting
Background
Cell sorting is the ability to separate cells according to their physical properties, such as size, density, electric charge, etc. Cell separation is important for many researchers because it allows for the collection of a single cell type from a bulk heterogeneous solution with little to no contamination. This can be hugely beneficial to patients who may need to be treated with a single type of cell. [1] Some methods of cell sorting include centrifugation, flow cytometry, magnetic cell separating, and Deterministic Lateral Displacement (DLD). Each of these methods and their challenges are further described in the Methods section of this page. Cell sorting is used for a variety of applications. Solid tissues can be broken down into specific cell types, cell cultures are often separated as a means of researching specific cells, and blood is most commonly separated for use in research as well as patient treatment.
Uses
Cell sorting is used for a variety of purposes, but one of the most common uses is the separation of blood cells. Blood separation is used for a variety of causes, including clinical diagnosis, research, and biotechnology. One of the most common examples of blood cell sorting is erythrocyapheresis, which is the method of separating red blood cells (RBC) from whole blood and recirculating the resultant blood back into the patient. Many blood centers use this as a form of blood donation. This allows for the collection of more useful RBCs without collecting any waste. Centrifugation and membrane filtration are often used for this type of blood donation.
Flow cytometry, explained below, allows for viable research in biotechnology. The separation of cells gives researchers the ability to focus their research on specific cells and particles. In protein and cell engineering, cells are separated for a variety of purposes, depending on the area of study. Many studies are focused on screening for protein overproduction. [3]
Methods of Separation
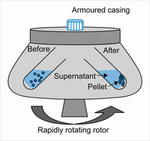

Centrifugation:
Separate cells based on their densities. This method is used in many laboratories and it is also used routinely in clinical settings. Denser and larger particles will sediment faster and collect at the bottom of the tube while lighter particles, cells, and solutions will accumulate above the dense particles. This process can be seen in Figure 1 to the right. The tubes will spin at a high velocity, causing the dense particles to accumulate at the bottom of the tube.
A major challenge with this type of separation is that the collected particles may not be completely pure. This can be accounted for by repeating the spinning process numerous times. Centrifugation is typically used for larger volume solutions. The most common application is the separation of blood cells. Solid tissues can also be broken down using centrifugation. The tissue is dissociated and filtered, then suspended and isolated with the centrifuge. [1]
Flow Cytometry:
Flow cytomtery is a laser based biophysical technology. This method is used to separate cells based on their physical and chemical characteristics. Cells pass through a light source one by one. The physical and chemical characteristics are detected and measured. Based on the light scattering and fluorescent characteristics of each cell, they will flow into different containers. A diagram of this can be seen in Figure 2. This method is similar to FACS, which is described below.
Fluorescence-Activated Cell Sorting and Magnetic-Activated Cell Sorting:
FACS and MACS are techniques most commonly used for antibody binding methods of cell sorting. Both methods raise antibodies against cell surface antigens. FACS utilizes fluorescent labeling of the antibodies, while MACS utilizes the antibodies’ conjugation to microbeads that contain iron oxide. After binding, FACS uses laser excitation to separate the cells. The fluorophores which are bound to the antibodies are excited to a point above the threshold level. This signals certain cells to separate out. MACS puts cells into a magnetic field where unlabeled cells are emitted out and labeled cells remain in the field.
Overall, MACS can be generally less specific than FACS, however, it takes significantly less time to complete. [1]
Lab on a Chip:
Lab on a chip is a method that is used for microscale cell separation. This technique is typically used for the experimental stage of different types of separation. Micro-scale filters are used to separate cells based on size and deformability. Micro-scale pillars are also used for this and this method is referred to as Deterministic Lateral Displacement (DLD). Field Flow Fractionation (FFF) uses parabolic flow as well as an external magnetic or gravitational field to separate cells along the length of a microfluidic channel. Acoustophoresis uses a high-pressure sound wave to deform cells based on their size and density. This wave also pushes cells towards specified areas of the microfluidic channel, which allows them to be separated out from cells of differing size and density. [1]
Benefits of Transitioning to the Microscale
There are numerous advantages to performing experiments on the micro-scale. Some of the more obvious advantages include reduced sample volumes and reduced costs. Others include simplicity of operation and high purity outputs. When working with such small volumes, it becomes easier to obtain complete separation of cells and particles. With the creation of soft lithography, building microfluidic devices which achieve high throughputs and high purity outputs becomes very easy and cost effective. Although there are many advantages that coincide with micro-scale experimentation, there are also drawbacks. On the micro-scale, all flow is laminar flow. This can be limiting in terms of diffusive mixing. The speed at which samples are able to flow also may be an issue. Tiny volumes tend to travel very slowly and delicate devices may require very slow flow rates. These slow flow rates may cause clogging of particles within the microfluidic device. [2]
Deterministic Lateral Displacement (DLD)
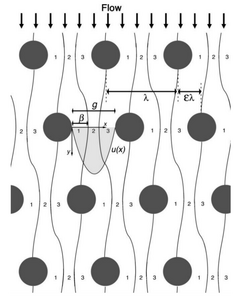
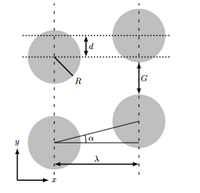
Deterministic Lateral Displacement is a method which uses the physical property of size to separate particles or cells. Posts are strategically placed along the microfluidic channel and the particles flow between them in such a way that would separate the particles based on their size. Based on the theory, cells can travel into three trajectories between posts due to their particle size. As the DLD schematic shows in Figure 3, smaller particles travel within the stream line flowing down to the device, however, the larger particles because the dynamics of it within each posts, they will be bumped through post and form a zigzag trajectory down to the device.
Design parameters:
The device is designed based on the size of the microspheres that are use. The size of the posts and distance between each post are important parameters for designing the device. Both of these parameters are calculated using the size of the microspheres that are to be separated. The gap size between each post can be calculated with the know critical particle diameter, D_c, using the equation: D_c=2 ηgε
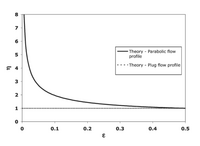
The row shift, ε, comes from the shifting stream for each rows. If 3 streams flow between two posts, then ε is equal to 1/3=0.33. A variable parameter, η, is used for non-uniform flow through posts. The η number chosen is based on the η vs.ε plot found in Figure 5. If ε=0.33, then η=1.3.
Cell density:
There are two ways to count cell densities. The first is to calculate using the equation found in Figure 6. The microspheres per milliliter is calculated with that equation. [11] The second method is to manually count with a microscope and a hemocytometer. A picture of the cells inside of the hemocytometer can be found in Figure 7. The cells in the four corners of the 3x3 square are counted and averaged to find the average cell count for the solution.


Particle Suspension:
Adding a desired amount of microspheres to a vial with 0.1% SDS solution. Note: Usually we do 0.005 g/ml, particles will bind together if concentration is too high. Using centrifuge to wet the spheres for 5-10 minutes. Note: Don’t forget to make a same weight vial as the sample one to balance the centrifuges. [12]
Material Considerations:
Polyethylene Microspheres: Microspheres can be made from a variety of different materials. Some of these include: PMMA, glass, metal or other materials. Polyethylene microspheres are commonly used for microfluidic device experiments. They are generally hydrophobic with high spherocity. Their size can vary between 1um and 1mm. They are colored and fluorescent visibly. All of these properties can help to mimic the real blood cells separation process.
PDMS: PDMS is the most commonly material used to make microfluidic chips. It is a transparent, deformable, and inexpensive elastomer. It molds very easity to other microfluidic materials like glass, silicon or other hard materials. PDMS is permeable can apply for cell culture. [10]
References
[1] 10.1177/2041731412472690
[2] 10.1016/j.nantod.2009.12.001
[3] 10.1186/1475-2859-5-12
[4] Davis, J. A.; Inglis, D. W.; Morton, K. J.; Lawrence, D. A.; Huang, L. R.; Chou, S. Y.; Sturm, J. C.; Austin, R. H. Deterministic Hydrodynamics: Taking Blood Apart. Proceedings of the National Academy of Sciences. 2006, 103, 14779–14784.
[5] Krüger, T.; Holmes, D.; Coveney, P. V. Deformability-Based Red Blood Cell Separation in Deterministic Lateral Displacement Devices—A Simulation Study. Biomicrofluidics. 2014, 8, 054114.
[6] Karabacak, N. M.; Spuhler, P. S.; Fachin, F.; Lim, E. J.; Pai, V.; Ozkumur, E.; Martel, J. M.; Kojic, N.; Smith, K.; Chen, P.-I.; Yang, J.; Hwang, H.; Morgan, B.; Trautwein, J.; Barber, T. A.; Stott, S. L.; Maheswaran, S.; Kapur, R.; Haber, D. A.; Toner, M. Microfluidic, Marker-Free Isolation of Circulating Tumor Cells from Blood Samples. Nat Protoc Nature Protocols. 2014, 9, 694–710.
[7] Inglis, D. W.; Davis, J. A.; Austin, R. H.; Sturm, J. C. Critical Particle Size for Fractionation by Deterministic Lateral Displacement. Lab on a Chip Lab Chip. 2006, 6, 655.
[8] Hou, Han Wei, Ali Asgar S. Bhagat, Wong Cheng Lee, Sha Huang, Jongyoon Han, and Chwee Teck Lim. “Microfluidic Devices for Blood Fractionation.” Micromachines 2, no. 4 (July 20, 2011): 319–343.
[9] Sritharan, D. SIZE-BASED CELL SORTI G BY DETERMI ISTIC LATERAL DISPLACEME T. Dissertation, Northeastern University, 2009.
[10] Ren, K.; Zhou, J.; Wu, H. Materials For Microfluidic Chip Fabrication. Accounts of Chemical Research Acc. Chem. Res. 2013, 46, 2396–2406. DOI: http://pubs.acs.org/doi/full/10.1021/ar300314s
[11] Microspheres & Particles Handling Guide. Polysciences, Inc, http://www.polysciences.com/skin/frontend/default/polysciences/pdf/microparticles guide.pdf.
[12] www.cospheric.com
