CHIP:Research
Research Interests
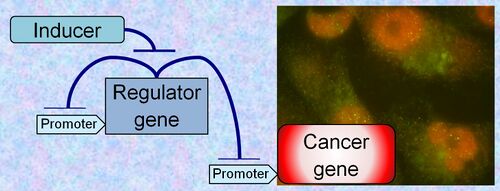
"Using synthetic gene circuits and genome editing to control cells and study cellular evolution, development, and cancer."
- Project #1. We study by experiment and computational modeling the effect of biological noise (nongenetic cellular diversity) on (i) survival during drug treatment [1] and the evolution of drug/stress resistance [22,47,52]; (ii) cellular behaviors related to metastasis [23]. We showed that noise can aid cell survival after exposure to stress (drug treatment) [1]. We introduced the concept of fitness noise [19], which arises when noisy protein levels affect cell doubling times. We have studied the effect of regulatory network architecture on fitness noise, and its contribution to the emergence of non-genetic resistance/tolerance, and later, genetic drug resistance [47,52].
- Project #2. We combine (i) genome editing with (ii) synthetic biology for single-copy integration of synthetic gene circuits that control the distribution of protein levels within a cell population [1,7,19,22,47,48]. We use various chemicals and light [48] as control signals. This surpasses the typical on/off control of only the cell population mean. For example, we can now independently adjust the mean and noise of a target gene's expression in yeast [1,7] and mammalian cells [47]. We have built "linearizer" gene circuits in yeast [7] and various rodent [47] and human cells [22,48] to tune the expression of a target gene linearly and precisely, with minimal noise, by adjusting the concentrations of various external chemicals and light intensity. We are "talking" to cells and "watch" their responses by single-cell measurements.
- Project #3. We study the responses of large-scale gene regulatory networks of infectious microbes and cancer cells to external stress using public gene expression data and regulatory networks [6,18,30]. We identify distinct sets of transcriptional subnetworks that are affected following exposure to stress. These results pave the path towards a systems-level understanding of the response of infectious microbes or cancer cells to stress, providing insights into their drug tolerance or drug resistance. We are interested in using synthetic gene circuits as "control knobs" to control various nodes of natural gene networks, thereby affecting cellular decisions such as epithelial/mesenchymal states or cell cycle exit, and comparing the results with mathematical predictions.
- Project #4. We study genetic and environmental causes of pattern formation in yeast and cancer cells by applying precisely controlled environments [26] and gene expression. We study by mathematical modeling how physical factors (strain, pressure, friction) interact with biological aspects (growth rate, cell-cell and cell-substrate attachment) to give rise to patterns. We are also interested building physical models and experimental systems to test how environmental factors (such as temperature or various chemical concentrations) affect the gene expression patterns in cell populations.
References since 2006 when GB established his lab -
(* indicates a "core" paper recommended for anyone interested in joining the lab):
1*. Blake WJ, Balázsi G, Kohanski MA, Isaacs FJ, Murphy KF, Kuang Y, Cantor CR, Walt DR, Collins JJ. Phenotypic consequences of promoter-mediated transcriptional noise. Mol. Cell 24(6):853-865 (2006). [1]
2. Murphy, KF, Balázsi G, Collins JJ. Combinatorial promoter design for engineering noisy gene expression. Proc. Nat. Acad. Sci. 104(31):12726-12731 (2007).
3. Balázsi G, Collins JJ. Sensing Your Surroundings: Taking the inventory inside single cells. News and Views. Nature Chemical Biology 3(3):141-142 (2007).
4. Strickler JR, Balázsi G. Planktonic copepods reacting selectively to disturbances. Phil. Trans. R. Soc. B. (2007)
5. Ernst J, Beg QK, Kay KA, Balázsi G, Oltvai ZN, Bar-Joseph Z. A semi-supervised method for predicting transcription factor-gene interactions in Escherichia coli. PLoS Comput Biol. 2008 Mar 28; 4(3):e1000044.
5. Heath AP, Kavraki L, Balázsi G, Bipolarity of the Saccharomyces Cerevisiae Genome. IEEE 2nd Intl. Conf. Bioinformatics and Biomedical Engineering, 330-333 (2008).
6. Balázsi G, Heath A, Shi L, Gennaro ML. The temporal response of the Mycobacterium tuberculosis gene regulatory network during growth arrest. Mol. Systems Biol. 4:225 (2008).
7*. Nevozhay D, Adams R, Murphy K, Josic K, Balázsi G. Negative autoregulation linearizes the dose response and suppresses the heterogeneity of gene expression. Proc. Nat. Acad. Sci. 106(13), 5123-5128 (2009). [2]
8. Irimia D, Balázsi G, Agrawal N, Toner M, Adaptive-Control Model for Neutrophil Orientation in the Direction of Chemical Gradients. Biophys. J. 96(10), 3897-3916 (2009).
9. Veiga DFT, Dutta B, Balázsi G, Network inference and network response identification: moving genome-scale data to the next level of biological discovery. Mol. Biosyst. 6(3), 469-480 (2010).
10. Murphy KF, Adams R, Wang, X, Balázsi G, Collins JJ, Tuning and controlling gene expression noise in synthetic gene networks. Nucleic Acids Res. 38(8), 2712-2726 (2010).
11. Balázsi G, Network reconstruction reveals new links between aging and calorie restriction in yeast. HFSP Journal 4(3), 94-99 (2010).
12. Tiwari A, Balázsi G, Gennaro M, and Igoshin OA, Interplay of multiple feedbacks with post-translational kinetics results in bistability of mycobacterial stress-response. Phys. Biology 7(3), 036005 (2010).
13. Nevozhay D, Adams R, Balázsi G, Linearizer Gene Circuits with Negative Feedback Regulation. Methods Mol Biol. 734, 81-100 (2011).
14. Datta P, Shi L, Bibi N, Balázsi G, Gennaro ML, Regulation of central metabolism genes of Mycobacterium tuberculosis by parallel feed-forward loops controlled by sigma factor E (σ(E)). J Bacteriol. 193(5), 1154-60 (2011).
15*. Balázsi G, van Oudenaarden A, Collins JJ, Cellular decision making and biological noise: from microbes to mammals. Cell 144(6), 910-925 (2011). [3]
16. Stamatakis M, Adams RM, Balázsi G, A common repressor pool results in indeterminacy of extrinsic noise. Chaos 21(4), 047523 (2011).
17. Quan S, Ray JC, Kwota Z, Duong T, Balázsi G, Cooper TF, Monds RD. Adaptive Evolution of the Lactose Utilization Network in Experimentally Evolved Populations of Escherichia coli. PLoS Genet. 8(1), e1002444 (2012).
18. Dutta B, Pusztai L, Qi Y, André F, Lazar V, Bianchini G, Ueno N, Agarwal R, Wang B, Shiang CY, Hortobagyi GN, Mills GB, Symmans WF, Balázsi G, A network-based, integrative study to identify core biological pathways that drive breast cancer clinical subtypes. Br J Cancer 106(6):1107-16 (2012).
19*. Nevozhay D, Adams RM, Van Itallie E, Bennett MR, Balázsi G, Mapping the environmental fitness landscape of a synthetic gene circuit. PLoS Comput. Biol. 8(4):e1002480 (2012). [4]
20. Rohde KH, Veiga DF, Caldwell S, Balázsi G, Russell DG, Linking the Transcriptional Profiles and the Physiological States of Mycobacterium tuberculosis during an Extended Intracellular Infection. PLoS Pathog. 8(6):e1002769 (2012.)
21. Claerhout S, Dutta B, Bossuyt W, Zhang F, Nguyen-Charles C, Dennison JB, Yu Q, Yu S, Balázsi G, Lu Y, Mills GB, Abortive autophagy induces endoplasmic reticulum stress and cell death in cancer cells. PLoS One 7(6):e39400 (2012).
22*. Nevozhay D, Zal T, Balázsi G, Transferring a synthetic gene circuit from yeast to mammalian cells. Nat Commun. 4:1451 (2013). [5]
23*. Lee J, Lee J, Farquhar KS, Yun J, Frankenberger CA, Bevilacqua E, Yeung K, Kim EJ, Balázsi G, Rosner MR, Network of mutually repressive metastasis regulators can promote cell heterogeneity and metastatic transitions. Proc. Natl. Acad. Sci., 111(3):E364-73 (2014). [6]
24. Lee J, Tiwari A, Shum V, Mills GB, Mancini MA, Igoshin OA, Balázsi G, Unraveling the regulatory connections between two controllers of breast cancer cell fate. Nucleic Acids Res., 42(11):6839-49 (2014).
25. Charlebois DA, Balázsi G, Kaern M, Coherent feedforward transcriptional regulatory motifs enhance drug resistance. Phys. Rev. E. 89:052708 (2014). [7]
26. Chen L, Noorbakhsh J, Adams RM, Samaniego-Evans J, Agollah G, Nevozhay D, Kuzdzal-Fick J, Mehta P, Balázsi G, Two-Dimensionality of Yeast Colony Expansion Accompanied by Pattern Formation. PLoS Comput. Biol. 10(12):e1003979 (2014). [8]
27*. González C, Ray JC, Manhart M, Adams RM, Nevozhay D, Morozov AV, Balázsi G, Stress-response balance drives the evolution of a network module and its host genome. Mol. Syst. Biol. 11(8):827 (2015). [9]
28. Belete MK, Balázsi G, Optimality and adaptation of phenotypically switching cells in fluctuating environments. Phys. Rev. E 92:062716 (2015). [10]
29. Ray JC, Wickersheim ML, Jalihal AP, Adeshina YO, Cooper TF, Balázsi G, Cellular Growth Arrest and Persistence from Enzyme Saturation. PLoS Comput. Biol. 12(3):e1004825 (2016). [11]
30. Chauhan R, Ravi J, Datta P, Chen T, Schnappinger D, Bassler KE, Balázsi G, Gennaro ML, Reconstruction and topological characterization of the sigma factor regulatory network of Mycobacterium tuberculosis. Nat. Commun. 7:11062 (2016). [12]
31. Diao J, Charlebois DA, Nevozhay D, Bódi Z, Pál C, Balázsi G, Efflux Pump Control Alters Synthetic Gene Circuit Function. ACS Synth. Biol. 5(7):619-631 (2016) [13].
32. Bien H, Balázsi G, Book review on "Biomolecular Feedback Systems" by Del Vecchio and Murray, Quart. Rev. Biol. 91(2), 220-221 (2016).
33. Charlebois DA, Balázsi G, Frequency-dependent selection: a diversifying force in microbial populations, Mol. Syst. Biol. 12(8):880 (2016) [14].
34. Shao Q, Trinh JT, McIntosh CS, Christenson B, Balázsi G, Zeng L, Lysis-lysogeny coexistence: prophage integration during lytic development, MicrobiologyOpen, (2016) [15].
35. Trinh J, Székely T, Shao Q, Balázsi G, Zeng L, Cell Fate Decisions Emerge as Phages Cooperate or Compete Inside their Host. Nat. Commun. 8:14341 (2017). [16]
36. Bódi Z, Farkas Z, Nevozhay D, Kalapis D, Lázár V, Csörgő B, Nyerges Á, Szamecz B, Fekete G, Papp B, Oliveira JL, Moura G, Santos MAS, Székely T Jr., Balázsi G, Pál Cs. Phenotypic heterogeneity promotes adaptive evolution PloS Biol. 15(5):e2000644 (2017). [17]
37. Bouklas T, Alonso-Crisóstomo L, Székely T Jr., Diago-Navarro E, Orner EP, Smith K, Munshi MA, Del Poeta M, Balázsi G, Fries BC. Generational distribution of a Candida glabrata population: Resilient old cells prevail, while younger cells dominate in the vulnerable host. PloS Path. 13(5):e1006355 (2017). [18]
38. Cortes M, Trinh J, Zeng L, Balázsi G. Late-arriving signals contribute less to cell fate decisions. Biophys. J. 113(9):2110-2120 (2017). [19]
39. Charlebois DA*, Diao J*, Nevozhay D, Balázsi G. Negative Regulation Gene Circuits for Efflux Pump Control. Methods in Molecular Biology, 1772:25-42 (2018). *Equal contribution.
40. Székely T Jr, Balázsi G. Beyond Promoters: How Genes Tweak Their Own Expression. Trends Genet. 34(10):733-735 (2018). [20]
41. Li C, Balazsi G. A landscape view on the interplay between EMT and cancer metastasis. NPJ Syst. Biol. Appl. 4:34 (2018). [21]
42. Shao Q, Cortes MG, Trinh JT, Guan J, Balázsi G, Zeng L. Coupling of DNA Replication and Negative Feedback Controls Gene Expression for Cell-Fate Decisions. iScience 6:1-12 (2018). [22]
43. Andrews Steven S, Brent R, Balázsi G. Signaling systems: Undistorted signaling. eLife 7:e41894 (2018). [23]
44*. Charlebois DA, Hauser K, Marshall S, Balázsi G. Multiscale effects of heating and cooling on genes and gene networks. Proc. Natl. Acad. Sci. 115(45), E10797-E10806 (2018). [24]
45. Charlebois DA, Balázsi G. Modeling cell population dynamics. In Silico Biology 13:21-23 (2019). [25]
46. Gómez Tejeda Zañudo J, Guinn MT, Farquhar K, Szenk M, Steinway SN, Balázsi G, Albert R. Towards control of cellular decision-making networks in the epithelial-to-mesenchymal transition. Phys. Biol. 16(3):031002 (2019). [26]
47*. Farquhar KS, Charlebois DA, Szenk M, Cohen J, Nevozhay D, Balázsi G. Role of network-mediated stochasticity in mammalian drug resistance. Nat Commun. 10(1):2766 (2019). [27]
48*. Guinn MT, Balázsi G. Noise-reducing optogenetic negative-feedback gene circuits in human cells. Nucleic Acids Res. pii: gkz556 (2019). [28]
49. Kuzdzal-Fick JJ, Chen L, Balázsi G. Disadvantages and benefits of evolved unicellularity versus multicellularity in budding yeast. Ecology & Evolution 9(15):8509-8523 (2019). [29]
50. Cortes MG, Krog J, Balázsi G. Optimality of the spontaneous prophage induction rate. J Theor Biol. 483:110005 (2019). [30]
51. Phillips K, Widmann S, Lai HY, Nguyen J, Ray JC, Balázsi G, Cooper TF. Diversity in lac operon regulation among diverse Escherichia coli isolates depends on the broader genetic background but is not explained by genetic relatedness. mBio 10(6), e02232-19 (2019). [31]
52*. Kheir Gouda M, Manhart M, Balázsi G. Evolutionary regain of lost gene circuit function. Proc. Natl. Acad. Sci. 116(50):25162-25171 (2019). [32]
53. Szenk M, Yim T, Balázsi G. Multiplexed Gene Expression Tuning with Orthogonal Synthetic Gene Circuits. ACS Synth. Biol. 9(4):930-939 (2020). [33]
54. Agozzino L, Balázsi G, Wang J, Dill KA. How Do Cells Adapt? Stories Told in Landscapes. Annu. Rev. Chem. Biomol. Eng. 11:155-182 (2020). [34]
55. Guinn MT, Wan Y, Levovitz S, Yang D, Rosner MR, Balázsi G. Observation and Control of Gene Expression Noise: Barrier Crossing Analogies between Drug Resistance and Metastasis. Front. Gen. 11:586726 (2020). [35]
56. Gama LR, Giovanini G, Balázsi G, Ramos AF. Binary Expression Enhances Reliability of Messaging in Gene Networks. Entropy 22(4):479 (2020).
57. Balázsi G. Discovering evolutionary hidden treasures. Nature Comp. Sci. 1:18–19 (2021). [36]
58. Cortes MG, Lin Y, Zeng L, Balázsi G. From bench to keyboard and back again: A brief history of lambda phage modeling. Ann. Rev. Biophysics 50:117-134 (2021). [37]
59. Krzysztoń R, Wan Y, Petreczky J, Balázsi G. Gene-circuit therapy on the horizon: synthetic biology tools for engineered therapeutics. Acta Biochim. Pol. 68(3):377-383. (2021). [38]
60. Balázsi G. Multiscale stress response. What differentiates a stress response from responsiveness in general? (Voices) Cell Systems 13(3):P195-200 (2022). [39]
61. Guinn L, Lo E, Balázsi G. Drug-dependent growth curve reshaping reveals mechanisms of antifungal resistance in Saccharomyces cerevisiae. Communications Biology 5:292 (2022). [40]
62. Balázsi G. New antivirals exploit viral feedback tricks for a cure without resistance. Cell 185(13):2210-2212 (2022). [41]
63. Tshering LF, Luo F, Russ S, Szenk M, Rubel D, Tutuska K, Rail JG, Balázsi G, Shen MM, Talos F. Immune mechanisms shape the clonal landscape during early progression of prostate cancer. Dev. Cell 58(12):1071-1086.e8 (2023). [42]
64. Torres A, Cockerell S, Phillips M, Balázsi G, Ghosh K. MaxCal can infer models from coupled stochastic trajectories of gene expression and cell division. Biophys J. 122(13):2623-2635 (2023). [43]
65*. Wan Y, Cohen J, Szenk M, Farquhar KS, Coraci D, Krzysztoń R, Azukas J, Van Nest N, Smashnov A, Chern Y-J, De Martino D, Nguyen LC, Bien H, Bravo-Cordero JJ, Chan C-H, Rosner MR, Balázsi G. Nonmonotone invasion landscape by noise-aware control of metastasis activator levels. Nature Chem. Biol. 19(7):887-899 (2023). [44]
66. Delamonica B, Balázsi G, Shub M. Cusp bifurcation in a metastatic regulatory network. J. Theor. Biol. 5:111630 (2023). [45]
67*. Wan Y, Mu Q, Krzysztoń R, Cohen J, Coraci D, Helenek C, Tompkins C, Lin A, Farquhar K, Cross E, Wang J, Balázsi G. Adaptive DNA amplification of synthetic gene circuit opens a way to overcome cancer chemoresistance. Proc. Natl. Acad. Sci. 120(49):e2303114120 (2023). [46]
68. Helenek C, Krzysztoń R, Petreczky J, Wan Y, Cabral M, Coraci D, Balázsi G. Synthetic gene circuit evolution: Insights and opportunities at the mid-scale. Cell Chem. Biol. 31(8):1447-1459 (2024). [47]
69. Matsuzaki T, Weistuch C, de Graff A, Dill KA, Balázsi G. Transcriptional drift in aging cells: A global decontroller. Proc. Natl. Acad. Sci. 121(30):e2401830121 (2024). [48]
70. Wan Y, Helenek C, Coraci D, Balázsi G. Optimizing a CRISPR-Cas13d Gene Circuit for Tunable Target RNA Downregulation with Minimal Collateral RNA Cutting. ACS Synth. Biol. 13(10):3212-3230 (2024). [49]
71. Panagoda NT, Balázsi G, Sampson NS. Mycobacterium tuberculosis Mce3R TetR-like Repressor Forms an Asymmetric Four-Helix Bundle and Binds a Nonpalindrome Sequence. ACS Chem. Biol. 19(12):2580-2592 (2024). [50]
72. Azukas J, Krzysztoń R, Helenek C, Carter S, Li SX, Balázsi G, Strey HH. Single-Cell Parameter Inference Reveals Kinetic Heterogeneity in Synthetic Mammalian Gene Expression. Biophys. J. (in press, 2026). [51]
More information may be found on three other websites:
1) The Louis & Beatrice Laufer Center for Physical & Quantitative Biology: [52] and [53]
2) Biomedical Engineering Department: [54].
3) Stony Brook Cancer Center: [55]
Back to the main page: CHIP