CH391L/S2013 Alesha Stewart Jan 30 2013
Double Strand Break (DSB) Repair- User:Alesha Stewart
“Spontaneous DNA breakage in single living Escherichia coli cells” Jeanine M Pennington & Susan M Rosenberg
Background
Genomic maintenance plays an important role in cancer and genetic diseases, as cancerous cells usually have some form of variance in their genome. Double-stranded breaks (DSB) are one of the several instances of genome instability, where a lesion is formed from endogenous or exogenous agents1. The break often stems from ionizing radiation (IR), lesions in DNA replication forks, drugs used for chemotherapy, chromosomal stress, species reactive to oxygen, and general replication and developmental processes 2,5,6. Single-strand DNA lesions (SSL) that are not repaired are another starting point for DSBs 4.
DNA DSBs are crucial as they can result in apoptosis or the loss of genes required for cell survival.3 Repair of the break is also challenging and error prone since the likelihood of chromosomal translocations increases. These instances demonstrate why DBSs are harmful and tend to lead to carcinogenesis2.
Repair
The two methods of repair for DSBs are homologous recombination (HR) and non-homologous end-joining (NHEJ). In HR the nucleotide sequences of two sets of DNA are exchanged to repair the broken strands. For this to occur one of the sets of DNA must be undamaged. The 3’ strand of the DSB invades the undamaged double stranded DNA and partially unwinds it for homologous pairing. The strands form branch migrations and Holiday junctions to restore the DNA with the DSB without any crossing over of the two sets 2.
In NHEJ of a DSB, the broken pieces are ligated together (via the KU heterodimer) without the need of an undamaged, homologous set of DNA. However, there can be degradation at the ends of the break that can result in errors and the loss of DNA sequences2. DSBs caused from IR are usually repaired by this method4. DSBs can also be repaired by single-strand annealing (SSA) if the break occurs between two repeated sequences. The repeated sequences are extended on the ends and annealed to repair the lesion 4. Although each of these mechanisms have repaired DSBs in other organisms, E. coli lacks Ku and HR is the only form of repair Supplementary Note 1. There are no sister chromatids present in E. coli, so it uses its own rapid growth to have a homologous copy of its DNA for HR 11.
Damage response
When there is damage in the DNA the SOS response is induced and proteins that manage DNA damage are readily synthesized 10. Sensors detect the DSB and transduce a signal down the DNA-damage-response pathway. Effector proteins then direct the cell to: be repaired, trigger cell-cycle checkpoint arrest to prevent further damage, or trigger apoptosis 2. The G1/S checkpoint of the cell cycle prevents the replication of damaged DNA into S phase, while the G2/M checkpoint detects damaged chromosomes before mitosis. The G2/M checkpoint, however, is characterized as insensitive and negligent as it can take 10-20 DSBs to signal DNA damage. Consequently, low quantities of DSBs go undetected and the cells multiply evoking mutagenesis and carcinogenesis 6.
Introduction
Although DSBs have proven to be significant contributors to mutagenesis and carcinogenesis, there is still no method to explicitly evaluate these spontaneous DNA breaks in cells. Some forms of DNA damage have inferred “indirect estimates” of the rates in which DSBs occur, but this research intended to provide a more precise quantification 1.
Single living Escherichia coli (E. coli) cells were used to determine the amount of cells relaying the SOS damage response after experiencing spontaneous DNA DSBs. As an outcome of the DSB, there is single-stranded (ss) DNA with an attached RecA protein to cleave and inactivate the LexA transcription repressor. This inactivation intensifies expression of over 40 genes involved in SOS damage response for greater response and detection, to be quantified by a sensitive flow cytometric assay 1.
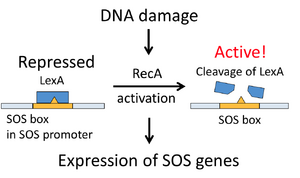
Methods
The researchers inserted a gfp gene into the chromosome to represent the wild-type E. coli in the experiment. Cells with this engineering were GFP+. The gene encoded for green fluorescent protein (GFP) coupled with the LexA-regulated sulA promoter. Engineering the wild type this way propelled the sulA promoter to produce a fluorescent green SOS response after experiencing DNA damage.
The strains of E. coli used were from the K12 strain MG1655. The derivations of the strains were formed via methods of P1 transduction or bacterial linear-replacement. The conditions of the experiments were log phase, low cell density, and strong ventilation. 105 cells were cultured for each individual experiment, and the data represent the averages of a minimum of 3 trials (Supplementary Methods1).
Results

The assay’s sensitivity and responsiveness to induction of the DNA SOS damage response was evaluated. In Figure 1a, positive and negative controls were used to assess the GFP intensity in the cells. The lexA(Def) cells were GFP+ and SOS-inducible (positive control); while the negative controls were the SOS-uninducible lexA(Ind-) and recA non-green cells. The lexA(Def) cells have a high GFP intensity as expected. Figure 1d, shows that ~0.9% of untreated wild-type cells spontaneously produce low fluoresce. The recB cells have the RecBCD enzyme that is required for SOS-induction from DSBs; thus the fraction of response related to the RecBCD enzyme was used to determine that 62% ± 3% of spontaneous SOS-induction was from DSBs. The remainder of the cells were RecA dependent, and fluoresced green due to DNA damage from other types of lesions 1.
Other control experiments were also done to determine if GFP+ cells still fluoresce after exposure to UV light and test spontaneous SOS-induction levels via GFP intensity.
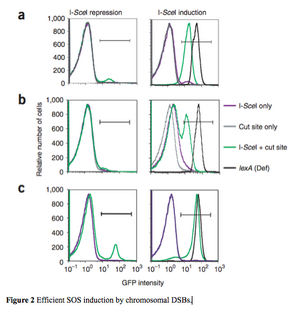
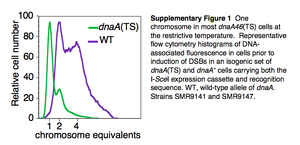
The researchers used I-SceI double-strand endonucleases and chromosomal cut sites to create single, chromosomal DSBs for evaluation of the assay's sensitivity. With these modifications, the cells fluoresced green and 87% ± 5% fell within the GFP+ gate after I-SceI induction Figure 2a. Since the log-phase cells have more than one chromosome, measures to detect a single DSB per cell were taken. A dnaA (TS) mutation was used to produce cells with only one chromosome by preventing replication at the restriction temperature. Without replication the cells divide until they only have one chromosome remaining that can then be cut by I-SceI induction. The chromosome reduction was successful in 61% ± 4% of the cells Supplementary Figure 1 which were then used to induce a single DSB per cell via I-SceI. 84% ± 4% of the single chromosomal DSB were detected to fluoresce within the GFP+ gate, indicating the assay was sensitive and efficient Figure 2c.
Since the cells no longer contained multiple chromosomes, an F' episome carrying ~20 kb of DNA similar to the region of the chromosome with the I-SceI cut site was incorporated within the cells. The DNA was used to repair the cuts DSBs by HR. From this they found that repairable DSBs could only be detected with 27% efficiency, implicating that only a fourth of repaired DSBs were detectable due to rapid repair or low sulA activation. By compiling the aforementioned data (0.9% of growing cells showed spontaneous SOS induction, 62% of log-phase cell showing RecB dependence, and assay detection of 27% of reparable DSBs), it was also determined that 2.1% of cells have 1 or more DSBs at 1% per generation 1 - values decreased by 20-100 fold from previous E. coli studies 9-10.
Determining cell fate after experiencing spontaneous damage to the DNA was another agenda. With fluorescence activated cells (FAC) it was found that 35% of GFP+ cells were culturable, rendering the remaining 65% non-culturable but still viable. This indicated that 35% of the cells were repaired and functional via the SOS-response; whereas 65% were in a response state that inhibited them from producing colonies. The viability of the cells were determined with propidium iodide staining. In conclusion, this data shows a drastically decreased occurrence of DSBs in cells than previously believed. However, 10% of the breaks are repaired mutagenically promoting chromosome rearrangements and genome instability 1.
References
- Pennington, J. M. & Rosenberg, S. M. Spontaneous DNA breakage in single living Escherichia coli cells. Nature Genetics 39, 797 - 802 (2007).
- Khanna, K. K. & Jackson, S. P. DNA double-strand breaks: signaling, repair and the cancer connection. Nature Genetics 27, 247 - 254 (2001).
- Rich, T., Allen, R. L & Wyllie, A. H. Defying death after DNA damage. Nature 407, 777–783 (2000).
- Vilenchik, M. M. & Knudson, A. G. Endogenous DNA double-strand breaks: production, fidelity of repair, and induction of cancer. Proc. Natl. Acad. Sci. USA 100, 12871–12876 (2003).
- O’Driscoll, M. and Jeggo, P. A. The role of double-strand break repair — insights from human genetics. Nature Reviews Genetics 7, 45-54 (2006).
- Löbrich, M. & Jeggo, P. A. The impact of a negligent G2/M checkpoint on genomic instability and cancer induction. Nature Reviews Cancer 7, 861-869 (2007).
- Friedberg, E.C. et al. DNA Repair and Mutagenesis. (American Society for Microbiology, Washington, D.C., 2005).
- Cox, M.M. et al. The importance of repairing stalled replication forks. Nature 404, 37–41 (2000).
- Kuzminov, A. Recombinational repair of DNA damage in Escherichia coli and bacter- iophage lambda. Microbiol. Mol. Biol. Rev. 63, 751–813 (1999).
- H. Lu et al., “Deinococcus radiodurans PprI switches on DNA damage response and cellular survival networks after radiation damage,” Molecular & Cellular Proteomics: MCP, vol. 8, no. 3, pp. 481-494, Mar. 2009.
- Kowalczykowski SC, Dixon DA, Eggleston AK, Lauder SD, Rehrauer WM (1994) Biochemistry of homologous recombination in Escherichia coli. Microbiol Rev 58: 401–465.