CH391L/S13/Metagenomics & Bioprospecting
Introduction
Metagenomics and bioprospecting are two 'umbrella' terms that were independently coined in the 1990's. These terms can create some confusion given shared characteristics. Whereas metagenomics is indeed a field of research, bioprospecting is more akin to a strategy or process encompassing many techniques.
Metagenomics and bioprospecting could be likened to "different sides of the same coin". More specifically, metagenomics is the sum of all genetic information present in a given environmental sample. Conversely, bioprospecting is simply application-driven research that seeks to discover commercially relevant compounds and biomaterials.
Bioprospecting: Hunting for Utility in Nature
The Roots of Bioprospecting
Of these concepts, bioprospecting is the elder and grew out of the field of chemical ecology. Dating back the 1950's, chemical ecology is the study of chemicals involved in interactions between organisms and their environment [1]. Increasing natural products research led to the active pursuit of commercially-relevant compounds in nature; also known as 'chemical prospecting'. While similar in principle, chemical prospecting ultimately relied upon chemical synthesis of newly discovered, commercially-relevant compounds. Many proponents for chemical prospecting were ardent conservationists arguing for the protection of our planets biodiversity [2]. Interestingly, ethical issues arose regarding exploitation or 'bio-piracy' with the INBio-Merck Argeement as perhaps the most famous instance [3].
The advent of the genomic era changed things yet again. DNA sequencing and recombinant techniques soon permitted biosynthetic production of natural products, which for some organic synthesis proved intractable. Many of those same technological advances would also lead to the development of metagenomics.
Therapeutics & Drug Discovery

As expected, there are many examples of bioprospecting for the purpose of drug discovery. As an outgrowth from chemical prospecting, considerable bioprospecting efforts - both past and present - have focused on plant secondary metabolites. One potent chemotherapy drug, paclitaxel (i.e. taxol) serves as an excellent example of this transition from chemical prospecting to bioprospecting. This isoprenoid compound was discovered in the bark of the Pacific Yew tree. Before the adoption of semi-synthetic production in 1988, therapeutic paclitaxel production relied upon low yield chemical extraction [4]. Using metabolic engineering techniques, researchers created transgenic Arabidopsis thaliana capable of producing taxidene, the first committed step in paclitaxel biosynthesis [5]. Since then, additional research led to production via plant cell fermentation. More recently, researchers engineered strains of E. coli and yeast with the capacity to produce taxidene and other isoprenoid compounds [4][6]. This was accomplished following the introduction of isoprenoid biosynthesis pathways. The tale of paclitaxel is principally considered a feat in the field of metabolic engineering. Still, those engineered strains of E. coli and yeast serve as platform technologies for tractable expression of other newly discovered enzymes.
Although terrestrial plants remain an important aspect of bioprospecting, increasing attention is being paid to marine biodiversity in the search for new therapies. Study of tunicates has led to the discovery of numerous cytotoxic compounds with potential anticancer properties [7]. Commonly known as seaweed, macroalgae offer another opportunity for bioprospecting [8].
Biofuels
Regarding second-generation or advanced biofuels, bioprospecting techniques are becoming an increasingly important strategy for biochemical pathway engineering and overall optimization. In a 2010 publication, LS9, Inc. reported the discovery of alkane biosynthesis pathways in a diverse set of cyanobacteria. Those enzymes were subsequently expressed in E. coli for the production of higher-value biofuel products [9]. That body of work provides an excellent demonstration of various bioprospecting techniques.
Research Tools
Fluorescent proteins are likely one of the most famous research tools derived from bioprospecting. Examples include dsRed as well as GFP and its many derivatives, which have been utilized throughout biological research [10]. Interestingly, these fluorescent proteins are finding new purpose in medicine as visual guides during surgery. Before tumorectomy, a mouse with internal tumors is injected with a recombinant form of GFP, which is targeted to and accumulates on the cells of blood vessels. During surgical removal of the tumor, the introduced GFP provides a surgeon with a strong visual queue of nearby blood vessels greatly reducing the risk of blood vessel lacerations [11].
DNA and RNA polymerases are the workhorses of modern biotechnology. Almost every aspect of modern biological research depends upon nucleic acid polymerases in one way or another. Recombinant cloning techniques, Sanger sequencing, and qPCR cover a few of the most common uses. These examples also highlight the shared importance of nucleic acid polymerases and Polymerase Chain Reaction (PCR). It was the development of PCR using Taq polymerase that began the drive for bioprospecting of DNA and RNA polymerases. Over the years, several other polymerases of thermophilic origin have been discovered and rapidly commercialized such as 'Vent polymerase' [12]. One area of considerable interest is the discovery or development of high-fidelity, thermostable reverse transcriptases. Using bioprospecting techniques, one research group isolated and cultured a novel thermophilic bacterium from a hot spring. That bacterium's DNA polymerase I gene was subsequently cloned and engineered to alter its specificity from DNA to RNA. In this manner, the researchers mutated the DNA-dependent DNA polymerase into an RNA-dependent DNA polymerase (i.e. a reverse transcriptase) [13]. This work may represent one step toward a thermostable reverse transcriptase with proofreading activity.
Metagenomics: Biocoenosis Data Mining
Consideration of biological organization greatly assists understanding the meaning of metagenomics. Within that conceptual framework, metagenomics is akin to the tier representing an interactive community of organisms; also known as a biocoenosis. In brief, metagenomics refers to the sum of all genetic information present in an environmental sample. The term itself was coined in 1998 [14]. Shortly thereafter, researchers characterized the first bacterial rhodopsin protein, which was isolated from seawater genomic DNA fragments [15].
Craig Venter and his Yacht
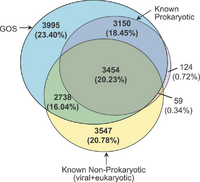
In the early 2000's, the J. Craig Venter Institute set as one of its goals to sequence the genomic diversity in the oceans. Craig Venter used his personal yacht, the Sorcerer II, to traverse the Earth's oceans, taking samples of oceanic life and sequencing using Whole Genome Shotgun Sequencing. From this adventure, they uncovered 6 million proteins (double the current database), which consisted of 1,700 clusters of gene families with no known homology. The data also revealed homology for 6,000 unknown ORF families (ORFan). They found that a very high proportion of new genes belonged to viruses (likely marine phage), which current databases had underrepresented [16].
The Benefit and Cost of Sequencing Technology
Since the turn of the century, metagenomics has bosomed as a field. Decreasing sequencing costs have greatly increased the number of metagenomic research projects. The April 2012 release of the UniProt database comprised an impressive 20.6 million protein sequences. However, only 2.8% of those protein sequences were confirmed at the protein and or transcript level. The matter is further complicated as the probability of feature identification is proportional to read length. Hence, there is a significant difference between the information gathered from pyrosequencing and Sanger sequencing [17].
Targeted Metagenomics: Bioprospecting with Metagenomics
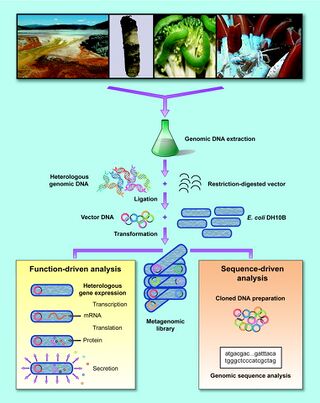
Targeted metagenomics is a useful combination of bioprospecting and metagenomics. This technique can be used to identify novel genes by screening ORFs derived from a metagenomic library. It is possible to conduct an initial screen computationally by first parsing identified ORFs for a desired homology. Following cloning, a functional screen is used to identify and recovery desired genes from the metagenomic library. Targeted metagenomic screens have led to the discover of new antibiotic resistance genes, cold-adaptive rRNAs, and cellulose degrading enzymes [18].
Typical Targeted Metagenomic Pipeline'
- Isolate nucleic acids (i.e. DNA, RNA) from an environmental sample
- Next-Gen sequencing
- Computational analysis: ORF and sequence homology
- Gene synthesis with possible codon optimization
- Heterologous expression of ORF library followed by functional screening
For a review on methods for constructing metagenomic libraries, see [19].
List of Parts found by Metagenomics
- glycosyl hydrolases (cellulose degradation)
- antibiotic resistance (tetracycline/bleomycin)
- extradiol dioxygenases (aromatic carbon usage)
- sulfate reductases
- deaminases
- Zn-dependent carboxypeptidases
- RNA-binding proteins
- many unique ORFans
Cellulosic Biomass degrading genes found in Cow Rumen
Plant polysaccharides such as cellulose are not broken down with enyzmes found in mammals, but species such as ruminents (cows) carry symbiotic bacteria that perform this job. These microbes cannot be cultured in lab. However, acquisition of the enzymes used to break down cellulose could be used to generate biofuel from easily grown plants like grass. Here, Mattias Hess and colleagues used metagenomic analysis to identify 51 enzymes active in breaking down polysaccharides [20]. The authors isolated microbes from a nylon bag filled with switchgrass placed inside a fistula created into a cow's rumen. Various next-gen sequencing technologies were used to generate 268 giga-basepairs of sequences. From the various organisms, they predicted 27,755 putative polysaccharide enyzymes, of which 43% had less than 50% similarity to any known sequence. The authors expressed and tested 90 of these genes based on similarity to glycosyl hyrdolase domains, which identified 51 active enzymes. The enyzmes were active against various substrates used as biofuel crops. Finally, the authors generated 15 draft genomes of new microbial species. A number of earlier studies have also attempted to identify glycosyl hydrolases in species such as termites and panda [21][18].
Current Status and the Future
The Human Microbiome Project
In a 2007 Nature article, researchers outlined the logistics of and rationale for amassing a human-microbe metagenome. Those authors described a human as a conglomerate of both human and microbial cells. Accordingly, they went on to postulate that this project would lead to an understanding of an individual's micro-evolution, that human's health, and disease predisposition. The author's further postulated that the resulting database would deepen the understanding of diagnostic biomarkers while having potential ramifications in industry through novel enzyme discovery [22].
Virology
Development of metagenomic techniques and protocols has also stimulated the field of virology. Viral metagenomic studies have led to the discovery of many previously unknown viruses. These endeavors have generated a vast amount of viral sequences for which the majority of sequences are reported as unknown [23]. It is worth noting that viral metagenomics has its own unique difficulties. Some viruses employ modified nucleotides as one part of their infection strategy. Additionally, many viruses employ lytic genes that could kill bacteria used during routine cloning. These and other factors necessitate techniques such as LASLs or link-amplified shotgun libraries [24].
References
-
Hartmann, Thomas. The lost origin of chemical ecology in the late 19th century. PNAS, 2008.
-
Eisner, Thomas. Prospecting for nature's chemical riches. Chemoecology, 1990.
-
Eisner, Thomas. Chemical Prospecting: A Global Imperative. Proceedings of the American Philosophical Society, 1994.
-
Boghigian, Brett A. Simultaneous production and partitioning of heterologous polyketide and isoprenoid natural products in an Escherichia coli two-phase bioprocess. J Ind Microbiol Biotechnol, 2011.
-
Besumbes, Oscar. Metabolic engineering of isoprenoid biosynthesis in Arabidopsis for the production of taxadiene, the first committed precursor of Taxol. Biotechnol Bioeng, 2004.
-
Engels, Benedikt. Metabolic engineering of taxadiene biosynthesis in yeast as a first step towards Taxol (Paclitaxel) production. Metab Eng, 2008.
-
Rinehart, K.L. Antitumor Compounds from Tunicates. Med Res Rev, 1999.
-
Pereira, Renato C. Bioprospecting for bioactives from seaweeds: potential, obstacles and alternatives. Braz J Pharmacogn, 2012.
-
Schirmer, Andreas. Microbial Biosynthesis of Alkanes. Science, 2010.
-
Tsien Roger. The green fluorescent protein. Annu. Rev. Biochem., 1998.
-
Nguyen, Quyen T. Surgery with molecular fluorescence imaging using activatable cell-penetrating peptides decreases residual cancer and improves survival. PNAS, 2011.
-
Leary, David. Marine genetic resources: A review of scientific and commercial interest. Marine Policy, 2009.
-
Sano, Sotaro. Mutations to create thermostable reverse transcriptase with bacterial family A DNA polymerase from Thermotoga petrophila K4. J Biosci Bioeng, 2012.
-
Handelsman, Jo. Molecular biology access to the chemistry of unknown soil microbes: a new frontier for natural products. Chemistry & Biology, 1998.
-
Beja, Oded. Bacterial Rhodopsin: Evidence for a New Type of Phototrophy in the Sea. Science, 2000.
-
Yooseph, Shibu. The Sorcerer II Global Ocean Sampling expedition: expanding the universe of protein families. PLoS Biol, 2007.
-
Temperton, Ben. Metagenomics: microbial diversity through a scratched lens. Curr Opin Microbiol, 2012.
-
Suenaga, Hikaru. Targeted metagenomics: a high-resolution metagenomics approach for specific gene clusters in complex microbial communities. Environ Microbiol, 2012.
-
Daniel, Rolf. The soil metagenome--a rich resource for the discovery of novel natural products. Curr Opin Biotechnol, 2004.
-
Hess, Matthius. Metagenomic discovery of biomass-degrading genes and genomes from cow rumen. Science, 2011.
-
Chistoserdova, Ludmila. Recent progress and new challenges in metagenomics for biotechnology. Biotechnol Lett, 2010.
-
Turnbaugh, Peter J. The Human Microbiome Project. Nature, 2007.
-
Rosario, Karyna. Exploring the world through viral metagenomics. Curr Opin Virol, 2011.
-
Breitbart, Mya. Genomic analysis of uncultured marine viral communities. PNAS, 2002.
-
Wilkins, Michael J. Proteogenomic monitoring of Geobacter physiology during stimulated uranium bioremediation. Appl Environ Microbiol, 2009.