CH391L/S12/Pattern Formation
Synthetic Pattern Formation
Organisms have evolved to receive signals, transduce the message, and elicit an appropriate response. Patterns are formed by cells deferentially responding to signals, whether it be from signals generated from other cells or external features such as light. In recent years, biologists are beginning to create synthetic pattern formation algorithms which allow for manipulation of biology on a more complex level. This page will specifically focus on synthetic pattern formation in E.coli.
Synthetic Pattern Formation Driven by Light
The synthetic biology community has been using light as a tool for pattern formation in microbes since 2002. [1] Light has been utilized for pattern formation in E. coli for the first ever organismal photograph[2]. Since then that system was modified to be an "edge detector" of light[3]. The focus of this section will be on the paper Multichromatic Control of Gene Expression in Escherichia coli[4].
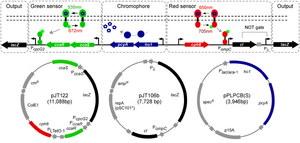
Having multiple light induction systems is great for having high resolution spatio-temporal control of gene expression.This paper essentially describes a two wavelength (RED and GREEN) gene expression system. Essentially, gene expression can be turned on via RED light or GREEN light in a specific and orthogonal manner. The RED system was borrowed from the initial photograph forming E.coli. Briefly, the RED system uses an engineered phytochrome, cph8 ,coupled to a EnvZ/OmpR pathway to activate gene expression. In the gene circuit cph8 is produced in the phosphorylated "ON" state, which upon induction of RED light switches to the unphosphorylated "OFF" state. In order for the RED light to produce LacZ, the cph8 regulates an inhibitor (C1) that binds to the LacZ reporter. In the dark cph8 is "ON" which will make C1 and keep the LacZ "OFF." But when RED light is present the cph8 no longer expresses the C1, thus allowing LacZ production. The cph8 system can be considered a NOT gate, or genetic inverter.
The RED system was tested by the Miller assay (a quantitative LacZ expression assay). In Dark conditions 0.58MU were produced as compared to RED conditions 1.41MU. This corresponds to an induction of 2.4 fold, which is similar to the green system. The RED system is dependent on the PCB chromophore, which is produced from heme by two genes on a plasmid. For proper induction under the corresponding light-wavelengths, transfer functions of induction were tested. This essentially tests the induction of LacZ expression across the spectrum of light. As it turns out, the RED system is turned on partially at the Green induction wavelength (532nM).
The GREEN system is a two component system involving a membrane associated histidine kinase CcaS and response regulator CcaR. Green light increases the rate of CcaS autophosphorylation, phosphotransfer to CcaR, and transcription by promoter cpcG2 upon CcaR binding. This is different than the RED system, which turns expression OFF in Red light, because it directly turns ON gene expression under green light conditions. The Green light System was tested through Miller Assays. In Green light the cells produced 50MUs of LacZ while in Dark Conditions made 25MU. (2-Fold Induction). Cells grown in Red light slightly decreased LacZ production. Experiments were also done in cells that lacked the PCB forming vector, which resulted in the same "No Response" LacZ production in each test case (Green, Red or Dark) and at roughly 10MU production of LacZ.
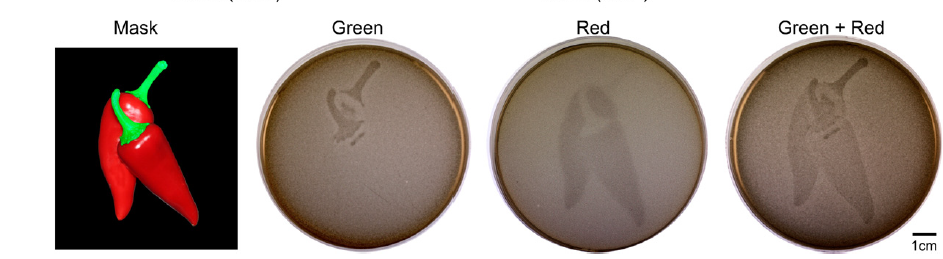
Both of these systems were use in parallel to demonstrate compatibility and orthogonality. The engineered cells were shown a mask of a chili pepper, which has a GREEN stem and RED core. The system responded correctly to each of the input signals after some tinkering with expression levels and the intensity of light. This tinkering was necessary because the RED system was turned on slightly at the GREEN wavelength (532nM). This effect was limited by using less intense light (which decreased sensitivity to the wrong wavelength) and by decreasing the strength of the ribosome binding site.
Synthetic Multicellularity
Pattern formation is critical for coordinated cellular behavior. This is achieved through cell-cell communication and intracellular signal processing. Pattern formation behavior was synthetically programmed into E. coli by the creation of two cell types, "senders" and "recievers."[5].
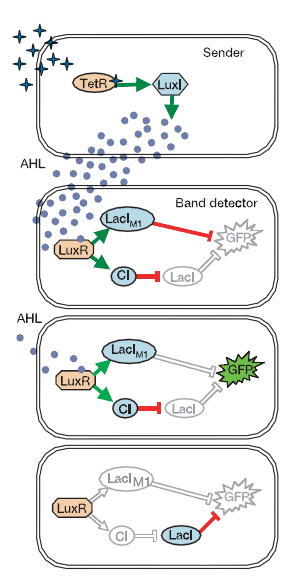

The sender cells produce the quorum sensing molecule, AHL, through expression of gene LuxI. The orientation of the sender cells produces a gradient of AHL around them; more concentrated AHL the more proximal to the sender cells. The concentration of AHL determines the transcriptional activation of C1 and LacI mediated by the LuxR transcriptional activator. Cells receiving the most AHL produce the highest levels of both C1 and LacI (Lux Mediated). The reporter for the system, GFP, is under control of the Lac promoter (which is inhibited by LacI). Therefore, receiver cells that have high concentrations of AHL produce large volumes of LacI and shut off GFP expression. If cells receive NO AHL then GFP production is shut off by the non-LuxR mediated LacI. However, when a cell is in a "sweet spot" receiving medium levels of AHL, then the LuxR-LacI is expressed poorly such that it cannot inhibit GFP expression, and the C1 repressor is produced enough to shut off the non-LuxI mediated LacI; turning ON GFP.
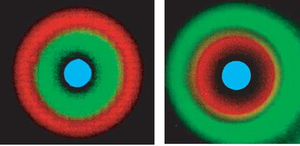
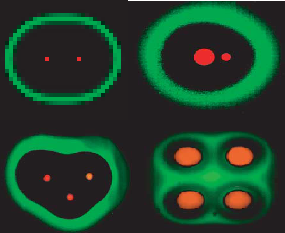
By varying the response of LuxR to AHL and the plasmid copy number, different responses to AHL concentrations are observed. This keeps the same banding pattern, but changes the response localization along the AHL gradient. This effect can be seen by using two cell types, with different responses and reporter gene. The result is a bullseye pattern, as the differing AHL response created different "sweet spots" between the two cell types. Interestingly, a dynamic "sweet spot" was observed when an unstable variant of LacI is used in place of the wild-type LacI. The response is due entirely the the spatial orientation of the sender cells. This was demonstrated by placing sender cells in various orientations and sizes. The placement of the senders on the plate created differing patterns. The fact that these programmed cells could create many complex patterns though a limited set of genetic circuitry is fascinating and gives insights on the origins and evolution of multicellular organisms.
Synthetic Pattern Formation via Digital Computing
In Tamsir et al.[6] electronic digital inspired the creation of its biological counterpart.Each cell will perform simple genetic algorithms but when placed in a cell-to-cell format more complex functions can arise. E. coli has been engineered to harbor synthetic Boolean logic including AND, OR and NOT functions. What is unique about this circuit is the simple circuits are compartmentalized from the logic program as a whole and communicate through quorum sensing molecules. This allows for robustness by allowing colonies of cells to give a population averaged response. Also, the same genetic parts can be reused orthogonally. An OR gate was created simply by placing two promoters in tandem to each other, which respond to exogenous small molecules. The activation of either promoter will turn on the reporter gene that is directly downstream of the promoters.
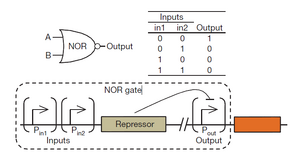
A simple NOR gate was created by placing two input promoters in tandem upstream of a repressor protein. The repressor will inhibit the reporter gene expression. Such that when either of the two inputs is in the ON state, the reporter gene is turned off.
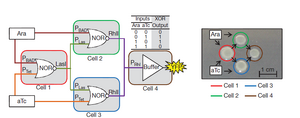
These logic functions can be put together in a more complicated array to compute higher order operations. The exemplary complex circuit herein was the XOR gate function. This function will have an output if either of the two signals is present, but not both signals simultaneously. This works by situating three NOR gates, to an output cell. It is easy to see why the signal is off when both stimuli are present (the Ara and aTc), because they are both linked to the NOR gates (in cells 2 and 3) that interact with the reporter cell (#4) through the RhII quorum producer. It is also simple to see why a single input will turn the reporter on. For instance, if Ara is present then cell 2's RhII will be OFF but cell 3 will be ON. The trick in this instance is the presence of Cell 1. Cell 1 will produce LasI, which will turn the NOR gates OFF in cell's 2 and 3, unless it is turned OFF by either the Ara or aTc.
References
- Levskaya A, Chevalier AA, Tabor JJ, Simpson ZB, Lavery LA, Levy M, Davidson EA, Scouras A, Ellington AD, Marcotte EM, and Voigt CA. Synthetic biology: engineering Escherichia coli to see light. Nature. 2005 Nov 24;438(7067):441-2. DOI:10.1038/nature04405 |
- Tabor JJ, Salis HM, Simpson ZB, Chevalier AA, Levskaya A, Marcotte EM, Voigt CA, and Ellington AD. A synthetic genetic edge detection program. Cell. 2009 Jun 26;137(7):1272-81. DOI:10.1016/j.cell.2009.04.048 |
- Tabor JJ, Levskaya A, and Voigt CA. Multichromatic control of gene expression in Escherichia coli. J Mol Biol. 2011 Jan 14;405(2):315-24. DOI:10.1016/j.jmb.2010.10.038 |
- Basu S, Gerchman Y, Collins CH, Arnold FH, and Weiss R. A synthetic multicellular system for programmed pattern formation. Nature. 2005 Apr 28;434(7037):1130-4. DOI:10.1038/nature03461 |
- Tamsir A, Tabor JJ, and Voigt CA. Robust multicellular computing using genetically encoded NOR gates and chemical 'wires'. Nature. 2011 Jan 13;469(7329):212-5. DOI:10.1038/nature09565 |
- Shimizu-Sato S, Huq E, Tepperman JM, and Quail PH. A light-switchable gene promoter system. Nat Biotechnol. 2002 Oct;20(10):1041-4. DOI:10.1038/nbt734 |
- Milias-Argeitis A, Summers S, Stewart-Ornstein J, Zuleta I, Pincus D, El-Samad H, Khammash M, and Lygeros J. In silico feedback for in vivo regulation of a gene expression circuit. Nat Biotechnol. 2011 Nov 6;29(12):1114-6. DOI:10.1038/nbt.2018 |