CH391L/S12/HostOrganisms
Host Organism Chassis
Introduction
Synthetic biology tends to focus on the assembly of defined parts and modules into novel circuits that perform some interesting or useful task. However, with the exception of in vitro transcription/translation systems, none of these complex constructions exist on their own, but rather in the context of a host cell and all of its cellular machinery and components. As a result, genetically identical genes and constructs will display different behavior in the context of different host cells. For this reason, understanding, constructing, and evolving host strains (described as "chassis" in synthetic biology jargon),is crucial to the development of reliably functioning biological systems.
Chassis Organisms
Escherichia coli: Escherichia coli is a gram-negative bacterium found in the human gut and a model organism in biological engineering research. It is the most intensively studied prokaryotic model organism, is easily transformed, has a rapid doubling time, and is a biosafety level 1 organism. All of these characteristics make it ideal for genetic engineering.
Bacillus subtilis: B. subtilis is a gram-positive model organism. It is easily transformable and its genome has been sequenced. It forms spores when exposed to stressful environments, which can make it difficult to kill with strategies often employed to kill bacteria.
Acinetobacter: Acinetobacter is a genus of gram-negative bacteria often found in soil and implicated in many infections of debilitated individuals. It is known for its exceptionally high natural competence, or ability to take up and incorporate foreign DNA into its genome. These properties make it ideal for studies of horizontal gene transfer, and some argue as a model organism for genetic engineering. [1]
Saccharomyces cerivisiae S. cerivisiae is a species of yeast used commonly in brewing and baking. It is eukaryotic, has a diploid and haploid stage of its life cycle, and is among the most intensively studied single-celled eukaryotic model systems. [2]
Mycoplasma genitalium: Mycoplasma genitalium is a small parasitic bacterium that lives on the ciliated epithelial cells of the primate genital and respiratory tracts. M. genitalium is the smallest known genome that can constitute a cell, which makes it an interesting subject for the study of minimal genomes.
E. coli Strains
Cloning
TOP10 E. coli can be purchased from vendors like Invitrogen, typically come at a transformation efficiency of 1 x 10e9 cfu/µg supercoiled DNA, and are ideal for high-efficiency cloning and plasmid propagation. They allow stable replication of high-copy number plasmids. Invitrogen TOP10 cells
The genotype of TOP10 cells is F- mcrA Δ(mrr-hsdRMS-mcrBC) φ80lacZΔM15 ΔlacΧ74 recA1 araD139 Δ(ara-leu) 7697 galU galK rpsL (StrR) endA1 nupG λ-
- hsdR for efficient transformation of unmethylated DNA from PCR amplifications
- mcrA for efficient transformation of methylated DNA from genomic preparations
- lacZΔM15 for blue/white color screening of recombinant clones
- endA1 for cleaner preparations of DNA and better results in downstream applications due to the elimination of non-specific digestion by Endonuclease I
- recA1 for reduced occurrence of non-specific recombination in cloned DNA
Protein Expression
BL21 is a common protein expression strain. Chemically competent cells can be acquired from vendors such as NEB at a transformation efficiency of 1–5 x 10e7 cfu/µg pUC19 DNA (note: much lower than TOP10). NEB BL21 cells
The genotype of BL21 is : fhuA2 [lon] ompT gal (λ DE3) [dcm] ∆hsdS λ DE3 = λ sBamHIo ∆EcoRI-B int::(lacI::PlacUV5::T7 gene1) i21 ∆nin5
T7 RNA Polymerase: (T7 gene1) is encoded by the lambda DE3 prophage present within the chromosome. T7 RNA polymerase is expressed from the lacUV5 promoter, which is less sensitive to catabolite repression than the wt lac promoter. Thus DE3 strains may exhibit uninduced target protein expression. Although λDE3 is normally dormant in the host chromosome, the induction of the SOS cascade can occur as the result of expressing proteins that damage the E. coli chromosome, either directly or indirectly. This may lead to cell lysis. T7 Express strains do not carry the DE3 prophage and better tolerate an SOS response.
Protease Deficient ([lon] ompT): E. coli B strains are “naturally” deficient in the lon protease which in K-12 strains serves to degrade misfolded proteins and to prevent some cell cycle-specific proteins from accumulating. The OmpT protease resides at the surface of wild type E. coli in both K-12 and B strains, presumably helping the cells to derive amino acids from their external environment. Cells deficient in both these proteases are much more amenable to the production of proteins from cloned genes.
T1 Phage Resistant (fhuA2): T1, an extremely virulent phage requires the E. coli ferric hydroxamate uptake receptor for infectivity. Deletion of this gene confers resistance to this type of phage, but does not significantly affect the transformation or growth characteristics of the cell.
Orthogonality
One strategy to minimize undesirable interaction of a synthetic construct with host machinery is to use orthogonal components. "Orthogonal" comes from the Greek orthos, meaning "straight", and gonia, meaning "angle". It has somewhat different meanings depending on the context, but most involve the idea of perpendicular, non-overlapping, varying independently, or uncorrelated. In this context, it means using components in your systems that do not come from the host cell and presumably interact minimally with the host cell.
One example is the use of genes that come from very unrelated organisms, presumably minimizing interactions within host cells. Another more extreme example is the introduction of the T7 RNA polymerase and an orthogonal ribosome to Escherichia coli, and using these components for the expression of the rest of your gene circuit. [3] This strategy somewhat uncouples circuit behavior from the endogenous ribosomes and RNA polymerase.
Genome Reduction
One approach to creating more reliable, efficient host organisms for synthetic constructs is the reduction of the genome to eliminate extraneous genes, mutagenic mobile elements, and other unnecessary or destabilizing factors. This can be viewed as a form of reverse engineering of extant strains.
Systematic Genome Reduction
One natural approach to engineering strains with a reduced genome is to systematically identify and delete regions of the genome not necessary for host cell survival. Posfai et al. created the MDS strains (multiple deletion strains) by aligning the genomes of multiple genomes of E. coli, identifying regions which were absent in multiple strains, and deleting them via Lambda Red recombination. All IS elements were removed as well, lowering the mutation rate and increasing the stability of genetic constructs introduced into the cell. The strain had comparable growth rate compared to wild type.[4][5]
Ara et al. constructed a minimal version of the B. subtilis genome in 2007. [6] This strain had slightly decreased growth rate compared to wild-type, but displayed normal morphology and similar protein production capabilities.
Selection for Reduced Genome
Long-term evolution of strains under the correct conditions could select for a genome of minimal size. Such conditions may include growth in rich media lacking sugars to favor the loss of biosynthetic pathways or sugar metabolism operons, growth in structured environments which favor a smaller cell, or growth under other conditions which favor the loss of unnecessary genes.
Minimal Genome Synthesis
Another approach is the synthesis of a minimal, designed genome from scratch using DNA synthesis technology and the transformation of this genome into cells to create a viable, novel, synthetic organism. This approach can be viewed as forward engineering of a novel organism, but would likely be informed by studies which determine the minimal set of genes necessary for a living organism.
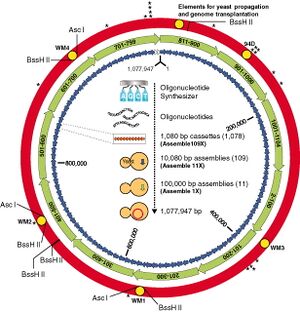
Mycoplasma mycoides Synthesis
Gibson et al synthesized the first artificial cell by generating the Mycoplasma mycoides genome from digitized genome information and transforming it into Mycoplasma capricolum cells devoid of genomic information. These cells were capable of continuous self-replication and were identified by "watermarks" inserted in the genome. This technology could be utilized in the future to create cells with novel and useful properties from scratch.[7][8]
Yeast as a Model Organism
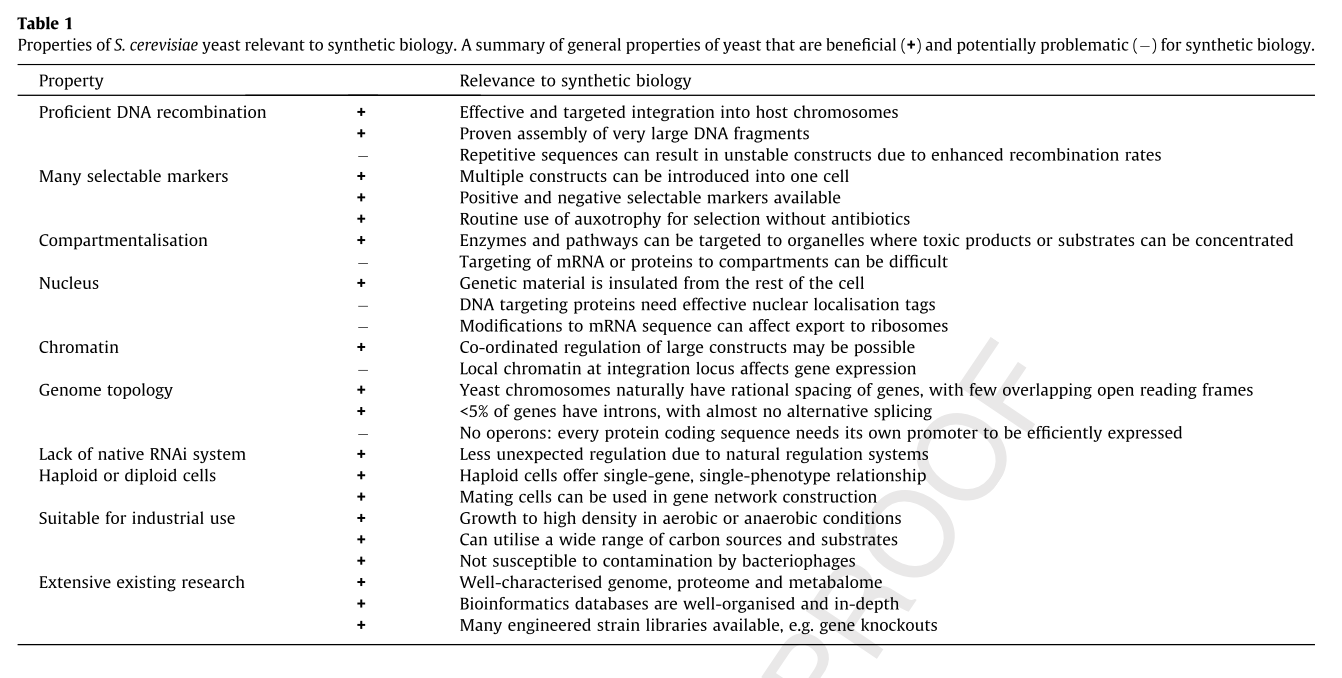
The following table summarizes some of the advantages and disadvantages of using yeast as a model organism for synthetic biology. [2]
References
- Metzgar D, Bacher JM, Pezo V, Reader J, Döring V, Schimmel P, Marlière P, and de Crécy-Lagard V. Acinetobacter sp. ADP1: an ideal model organism for genetic analysis and genome engineering. Nucleic Acids Res. 2004;32(19):5780-90. DOI:10.1093/nar/gkh881 |
- Blount BA, Weenink T, and Ellis T. Construction of synthetic regulatory networks in yeast. FEBS Lett. 2012 Jul 16;586(15):2112-21. DOI:10.1016/j.febslet.2012.01.053 |
- An W and Chin JW. Synthesis of orthogonal transcription-translation networks. Proc Natl Acad Sci U S A. 2009 May 26;106(21):8477-82. DOI:10.1073/pnas.0900267106 |
- Kolisnychenko V, Plunkett G 3rd, Herring CD, Fehér T, Pósfai J, Blattner FR, and Pósfai G. Engineering a reduced Escherichia coli genome. Genome Res. 2002 Apr;12(4):640-7. DOI:10.1101/gr.217202 |
- Pósfai G, Plunkett G 3rd, Fehér T, Frisch D, Keil GM, Umenhoffer K, Kolisnychenko V, Stahl B, Sharma SS, de Arruda M, Burland V, Harcum SW, and Blattner FR. Emergent properties of reduced-genome Escherichia coli. Science. 2006 May 19;312(5776):1044-6. DOI:10.1126/science.1126439 |
- Ara K, Ozaki K, Nakamura K, Yamane K, Sekiguchi J, and Ogasawara N. Bacillus minimum genome factory: effective utilization of microbial genome information. Biotechnol Appl Biochem. 2007 Mar;46(Pt 3):169-78. DOI:10.1042/BA20060111 |
- Gibson DG, Benders GA, Andrews-Pfannkoch C, Denisova EA, Baden-Tillson H, Zaveri J, Stockwell TB, Brownley A, Thomas DW, Algire MA, Merryman C, Young L, Noskov VN, Glass JI, Venter JC, Hutchison CA 3rd, and Smith HO. Complete chemical synthesis, assembly, and cloning of a Mycoplasma genitalium genome. Science. 2008 Feb 29;319(5867):1215-20. DOI:10.1126/science.1151721 |
- Gibson DG, Glass JI, Lartigue C, Noskov VN, Chuang RY, Algire MA, Benders GA, Montague MG, Ma L, Moodie MM, Merryman C, Vashee S, Krishnakumar R, Assad-Garcia N, Andrews-Pfannkoch C, Denisova EA, Young L, Qi ZQ, Segall-Shapiro TH, Calvey CH, Parmar PP, Hutchison CA 3rd, Smith HO, and Venter JC. Creation of a bacterial cell controlled by a chemically synthesized genome. Science. 2010 Jul 2;329(5987):52-6. DOI:10.1126/science.1190719 |