CH391L/S12/Bioprospecting
What's Bioprospecting?
Bioprospecting is a catch-all term for activities including discovery, acquisition, and utilization of novel biomaterials. This has historically been a controversial activity, often leading to unregulated commercialization of fauna (e.g., plants and medicinals) from third world countries for the benefit of commercial interests [Pros/Cons of Bioprospecting]. However, as a term in Molecular Biology, it reflects the growing need to discover new types of protein and nucleic acid parts, which can be used in biotechnology and basic research. The advent of multiple Next-Generation Sequencing technologies since 2006 now provides depth of information into the entire genomes (Metagenomics) of species previously inaccessible to basic research. [1]
Examples of Genes Identified via Bioprospecting
GFP
Although not planned, one of the great examples of Bioprospecting is the story of Green Fluorescent Protein (GFP), a protein that has had a profound impact on every major field in modern biology. Originally isolated and characterized by Osamu Shimomura in the 1960's and 1970's from jellyfish and sea pansies, it was a mere oddity that conferred the eery bioluminescence of certain deep sea creatures. However, the subsequent cloning of the gene by Martin Chalfie and improvement into enhanced GFP by Roger Tsien made it into one of the modern workhorses in biology. This 40-year journey earned Shimomura, Chalfie, and Tsien the 2008 Nobel Prize in Chemistry. [History of GFP]
Polymerases
Polymerases such as the Klenow fragment and more importantly Taq polymerase, have permitted the synthesis of DNA fragments. For instance, Taq was discovered in a thermophilic bacterium, and because it can withstand extreme heat (~95 celcius) without losing activity permitted the use of thermal cycling in the Polymerase Chain Reaction (PCR). This allows for denaturation and subsequent reannealing of DNA strands used in the exponential amplification of target sequences. PCR earned Mullis and Smith the 1993 Nobel Prize. Polymerases also include the important phage RNA polymerases such T7, T3, or SP6 which have permitted the in vitro transcription of DNA templates into RNA.
Reverse Transcriptases
The discovery of Reverse-Transcriptases, essentially polymerases that copy RNA into DNA, have allowed for the study of RNA via generation of cDNA's. These proteins are found in RNA viruses, mobile genetic elements, and mammalian telomerase. The work led to the 1974 Nobel Prize for David Baltimore and Howard Temin.
Restriction Endonucleases
The discovery of Restriction Endonucleases in the 1970's fueled the Molecular Biology revolution and the advent of genetic engineering. Found in bacteria and archaea, they act to degrade foreign DNA by cleaving at specific palindromic sequences. This work led to the 1978 Nobel prize to Nathans, Arber, and Smith.
Metagenomics
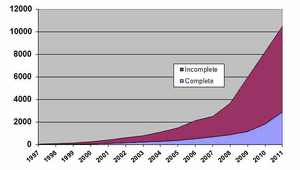
Metagenomics uses Next Generation Sequencing Technologies (e.g., Whole Genome Shotgun Sequencing (WGS), Roche 454, Illumina, ABI Solid) or Protein analysis (Mass Spectrometry) to completely sample the genomes of mixed microbial communities, generating an unbiased view of genomic sequence space. Estimates have suggested that greater than 99% of all microbes are unculturable in the lab and inaccessable to traditional laboratory analysis. Thus, these Next Generation Sequencing approaches allow for analysis of microbes that are small percentages of a microbial community. The current explosion in various Metagenomic projects (340 current projects, 1990 samples [GOLD database]) permits for entirely in silico approaches to identifying new gene families, with potential as parts in Synthetic Biology.
Craig Venter and his Yacht
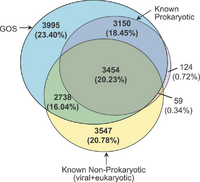
In the early 2000's, the J. Craig Venter Institute set as one of its goals to sequence the genomic diversity in the oceans. Craig Venter used his personal yacht, the Sorcerer II, to traverse the Earth's oceans, taking samples of oceanic life and sequencing using Whole Genome Shotgun Sequencing. From this adventure, they uncovered 6 million proteins (double the current database), which consisted of 1,700 clusters of gene families with no known homology. The data also revealed homology for 6,000 unknown ORF families (ORFan). They found that a very high proportion of new genes belonged to viruses (likely marine phage), which current databases had underrepresented. [2]
Examples of Bioprospecting using Metagenomics (Targeted Metagenomics)
A useful approach to Bioprospecting new genes involves either functional screening or pure sequence screening in what is called Targeted Metagenomics. This involves either challenging microbiota to a particular activity, or looking for specific families of genes. Both types of Targeted Metagenomic screens have led to new antibiotic resistance genes, cold-adaptive rRNA's, and cellulosic enzymes, to name just a few. [3].
- ----Typical Targeted Metagenomic Pipeline----
- Extract (DNA, RNA, or Protein) from Environmental Sample
- Next Gen Sequencing or Mass Spec
- Computational analysis for ORFs and homology searches
- Heterologous Expression and Testing for function
Cellulosic Biomass degrading genes found in Cow Rumen
Plant polysaccharides such as cellulose are not broken down with enyzmes found in mammals, but species such as Ruminents (cows) carry symbiotic bacteria that perform this job. These microbes cannot be cultured in lab. However, acquisition of the enzymes used to break down cellulose could be used to generate biofuel from easily grown plants like grass. Here, Mattias Hess and colleagues used Metagenomic analysis to identify 51 enzymes active in breaking down polysaccharides [4]. The authors isolated microbes from a nylon bag filled with switchgrass placed inside a fistula created into a cow's rumen. Various Next-Gen Sequencing technologies were used to generate 268 Giga-basepairs of sequences. From the various organisms, they predicted 27,755 putative polysaccharide enyzymes, of which 43% had less than 50% similarity to any known sequence. The authors expressed and tested 90 of these genes based on similarity to glycosyl hyrdolase domains, which identified 51 active enzymes. The enyzmes were active against various substrates used as biofuel crops. Finally, the authors generated 15 draft genomes of new microbial species. A number of earlier studies have also attempted to identify glycosyl hydrolases in species such as termites and panda [1][3]
Uranium Bioremediation
Bioremediation of uranium waste is an important industrial process. Here, the authors used a proteomics based approach to identify proteins in iron-reducing (FeIII) microbial species used in the reduction of soluble uranium to insoluble uranium [5]. Although the study here does identify new genes, it identifies new metabolic pathways. The authors started with 3 known Geobacter species stimulated with acetate and used in a bioremediation project and used the known proteins as a reference for performming 2D liquid chromatography tandem mass spectrometry. They identified over 13,000 peptides and 2,500 proteins. Not surprisingly, the authors show that acetate utilization increases via the TCA cycles (acetyl-CoA enzymes) presumably for increased growth as a fuel source. They also note high abundances of pyruvate ferrodoxin oxidoreductase, suggesting the bacteria are undergoing high carbon fixation processes.
Current status and the future
New genes found from Ruminant microbes are being used for generating fuel from biomass, proteins which have thus far been unknown to man. Genes identified in more extromophilic bacteria and archaea may be useful in metabolism of inorganic compounds. They may be useful in new genetic circuits in biotech applications. Finally, the genes will ultimately be useful as scaffolds for directed evolution studies, to generate new functions.
List of Parts found by Metagenomics
- glycosyl hydrolases (cellulose degradation)
- antibiotic resistance (tetracycline/bleomycin)
- extradiol dioxygenases (aromatic carbon usage)
- sulfate reductases
- deaminases
- Zn-dependent carboxypeptidases
- RNA-binding proteins
- many unique ORFans
For a review on methods of constructing metagenomic libraries to screen for useful genes as well as other useful genes isolated from the metagenome, see [6].
Limitations
Diversity
Often, the search for novel genes amongst divergent microbial species is limited to genes with small identical regions of DNA for cloning. Cloning of genes from a diverse sample can be helped with prior knowledge of the target gene family. Various domains of a given gene family can be targeted with degenerate primers to isolate similar genes in a microbial community sample. On the other hand, even the use of degenerate primers may not capture the entirety of gene families in a diverse sample. Alternatively, phylogenetic analysis of regions with high similarity (low degeneracy) can help for designing primers that better capture the genes.
Many of the current Next-Gen Sequencers are limited by short read lengths, which can prove problematic in the de novo construction of a genome. In addition, many protocols utilize emPCR after nebulization of DNA, which can introduce sequencing coverage bias. Nebulization randomly fragments sample genomic DNA into various lengths, which can be of varying GC content and secondary structure. Since emPCR amplifies single fragments, this can often lead to underrepresentation of difficult to PCR fragments. For low copy templates, this can often mean low sequence coverage in important areas.
The coming introduction of single-molecule long read sequencers, such as that by Pacific Biosciences may alleviate some of these limitations. Finally, it is unlikely that genes found in nature will cover all the uses humanity may come up with. Since nature settles for genes that function "well-enough", this may be inadequate since humans require efficiency.
Rare Genomes and Low Density Environments
Another issue is filtering out the interesting bacteria from the less interesting bacteria in a metagenome. When creating a library, it seems like it would be hard to maintain enough complexity to capture the plausibly rare and beneficial genes you're looking for. One solution to this might be to look into metagenomes where there is a huge selection pressure to have the genes you're looking for. Such an environment would have to be particularly hostile - something with very high concentrations of a given contaminant. While it doesn't allow you to capture the genomes of rare bacteria any better, it would increase the relative population of a bacteria with the physiology of interest. An interesting paper that talks about some of the evolutionary dynamics of such an environment has been recently published [7]. The situation does however create sort of a catch-22. Harsher conditions that lead to over representation of useful genes to cope with the environment also can have significantly lower overall concentrations of cells - on the order of 10,000/g of soil. By traditional library construction methods, this would require you to harvest dozens of kilograms of soil for extraction. A solution is to use φ29 DNA polymerase to amplify low concentrations of environmental DNA for library construction [8]. While this method has a lot of potential to give access to low concentration metagenomes, the innate biases it creates are not well understood.
References
<biblio>
- BiotechLettReview2010 pmid=20495950
//Recent progress and new challenges in metagenomics for biotechnology.
- EnviroMicroReview2011 pmid=21366818
//Targeted metagenomics: a high-resolution metagenomics approach for specific gene clusters in complex microbial communities.
- PLOSbio2007 pmid=17355171
//The Sorcerer II Global Ocean Sampling expedition: expanding the universe of protein families.
- Science2011 pmid=21273488
//Metagenomic discovery of biomass-degrading genes and genomes from cow rumen.
- ApplEnviroMicro2009 pmid=19717633
//Proteogenomic monitoring of Geobacter physiology during stimulated uranium bioremediation.
- Daniel2004 pmid=15193327
//The soil metagenome--a rich resource for the discovery of novel natural products.
- Hemme2010 pmid=20182523
//Metagenomic insights into evolution of a heavy metal-contaminated groundwater microbial community.
- Abulencia2006 pmid=16672469
//Environmental whole-genome amplification to access microbial populations in contaminated sediments.