Bryan Hernandez/20.109/Lab notebook/Module 3/Day 3
Colony PCR
module 3 Day 3
Purpose: Confirm the the success of the Spt3 Knockout/ URA3 Knock-in.
Introduction
Based on the numerous applications of PCR, it may seem that the technique has been around forever. In fact it is only a little more than 20 years old. In 1984, Kary Mullis described this technique for amplifying DNA of known or unknown sequence, realizing immediately the significance of his insight.

"Dear Thor!" I exclaimed. I had solved the most annoying problems in DNA chemistry in a single lightening bolt. Abundance and distinction. With two oligonucleotides, DNA polymerase, and the four nucleosidetriphosphates I could make as much of a DNA sequence as I wanted and I could make it on a fragment of a specific size that I could distinguish easily. Somehow, I thought, it had to be an illusion. Otherwise it would change DNA chemistry forever. Otherwise it would make me famous. It was too easy. Someone else would have done it and I would surely have heard of it. We would be doing it all the time. What was I failing to see? "Jennifer, wake up. I've thought of something incredible." --Kary Mullis from his Nobel lecture; December 8, 1983
To begin today’s experiment, you will “pop” some ura+ yeast from your transformation plates and then amplify the relevant portion of the released genomic DNA. The primers you will use are expected to give an ~800 base pair product only if URA3 is inserted into the gene you have tried to delete.
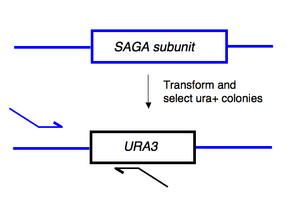
This is accomplished by using a forward primer that anneals upstream of the gene you're deleting and a reverse primer that anneals to a region within the URA3 gene. If the URA3 gene has inserted elsewhere in the genome (giving rise to the ura+ phenotype) but the SAGA-subunit remains in place, then you will see no product from this reaction. Note, however, that a negative result from these reactions (i.e. no PCR product) can just as easily be explained as a failed PCR (bad primers, dead enzyme, wrong reaction conditions, etc), and might incorrectly lead you to toss out perfectly correct samples. Only the positive result is meaningful in this experiment and we will not know the result until we run the gel next lab.
We will remain optimistic and set up overnight cultures of all the candidates you are examining today. In this way you will have cells to examine next time, when you will screen them for phenotypes arising from loss of the SAGA subunit and when you will isolate total RNA from them for microarray analysis.
Protocols
Part 1: Colony PCR
- Begin by counting the colonies that arose from the transformation experiment you performed last time. Choose three candidates (when possible) to pursue, circling the colonies on the back of the petri dish and clearly labeling them "1," and "2." If you do not have enough colonies, don't despair. You can get some additional ones from another group or from the teaching faculty. Note, however, that by doing so, you may be switching SAGA-subunits and you will have to rethink the work you have planned so far. And if applicable, it will be worth noting the "negative" result in your lab report, since it may indicate that despite the subunit being deemed non-essential, cells may fair less well than wild-type cells and so deletions may be difficult to isolate.
- You will use the microwave to release the DNA from the yeast. On the tip of a sterile toothpick, pick-up 1/2 of the colony that is candidate "1" and swirl the cells in 20 μl of sterile water in an eppendorf tube. Be sure to label the tube so you know it belongs to your group and which candidate it contains. Repeat with the second candidates you've selected.
- The teaching faculty will have the parent strain, FY2068, streaked out on a petri dish. You should mix 1/2 of a colony from this plate with 20 μl of sterile water in a fourth eppendorf tube.
- Close the caps to the tubes and microwave them in an eppendorf rack for 15 seconds.
- Move 2.5 μl of the microwaved mixes, yeast debris and all, into a 200 μl PCR tube. Again label your tubes well (write small!).
- In a new eppendorf tube (normal size this time), prepare a PCR cocktail enough for 4 reactions. Each reaction will include:
- 10 μl 2.5X PCR mastermix
- 0.5 μl forward primer specific to your gene of interest
- 0.5 μl reverse primer specific for URA3
- water to a final volume of 22.5 μl.
- Add 22.5 μl of PCR cocktail to each PCR tube with yeast and leave the tubes on ice until everyone is ready.
- Cycle the reactions as:
- 95° 4 minutes
- 95° 1 minute
- 52° 1 minute
- 72° 2 minute
- repeat steps 2-4 35 times
- 72° 10 minutes
- 4° forever
Part 2: Overnight cultures
- Using sterile technique, aliquot 2.5 ml of YPD into four sterile test tubes.
- Label the caps with your team color, and either "parent" "1" or "2."
- Use sterile dowels to move the second 1/2 of each yeast colony from your transformation plates to the media. Swirl the dowel to remove the yeast from the stick and vortex the solution to fully resuspend them.
- Use a sterile dowel to innoculate the "parent" culture, using the FY2068 plate that the teaching faculty have prepared.
- Place the tubes on the roller drum in the 30° incubator. They will grow for 24 hours and then be placed in the 4° fridge until next lab.
- When everyone is ready we will discuss the journal article that was assigned for today.
Part 3: Journal article discussion
As part of your assignment for today, you have read the relevant article by Wu et al, published in Mol Cell in 2004. There is also an associated review article that was written to celebrate and highlight this important work on the structure of the SAGA complex. You and your partner will be randomly assigned a Figure from the text to describe to the class (figures 1-6, not table 1) but be prepared to engage in conversation about other parts of this paper even if you're not "assigned" to the section being discussed.
discussion/interpretation
if we get a PCR product at the expected size we can interpret this as a positive result (out knock out worked). If we dont get a PCR product of the expected size, or a product at all, then this is a negative result which is relatively uninformative. In this case we would need to do further experimentation to confirm the success or failure of the knock-out.