Biomod/2015/Berlin15
Video
Check out our YouTube video! [1]
Abstract
- Increased surface area is the key factor for the developement of more efficient nano devices. We are introducing a multifunctional biomolecular nano scaffold with three-dimensional reactive nanostructures. Therefore, the FliC-gene - encoding for flagellin – is modified with variously designed D3-domains to integrate diverse modifications into the protein for our purpose of a multifunctional scaffold. The different D3 domains bear reagions for the incorporation of non-canonical amino acids. These add functionalities to link the designed flagellin with plastic-degrading enzymes to create the scaffold of a molecular filter system for degrading microplastics. However, by incorporating catechol derivatives into the D3-domains of FliC, these modified proteins can be used as ion catchers in nano electronics for different applications. Immobilizing functionalized flagellins on silica monoliths makes our nano device. In addition, immobilization of flagellin coupled with specific antibodies on assay plates display a third dimension which can significantly enhance the sensibility in immunoassays.
Flagella Modification
- Although flagellas are an assembly of multiple proteins, the flagellum filament is mostly made of one single protein called flagellin (FliC-gene). Flagellin proteins consist of four domains (D0-D3). While the D0 and D1 domains are well conserved, the D2 and D3 domains are highly variable in sequence and length. This allows us to modify the D3 domain for our purposes by making use of the Synthetic Biology. We designed different D3-domains with gBLOCKS:
a) MG1655 FliC_D3 (gB3)
b) Salmonella T D3-domain (gB4)
c) Salmonella T D3-domain containing 3x methionine codons and 6 His-tags (gB5)
d) Salmonella T D3-domain containing 3x methionine codons (gB6)
e) Salmonella T D3-domain containing 2x cysteine codons and one methionine codon and one His-tag (gB7)
When the D3-modified FliC-gene is introduced into a methionine auxotrophic strain, the tRNA that used to deliver methionine will deliver L-Homopropargylglycine (HPG; a non-canonical amino acid that is an analoga to methionine) due to the reason that we do not add methionine to the medium, so that the bacteria are basically forced to use HPG. Under mild conditions, HPG then reacts to specifically modified arginine residues that we are going to introduce in our enzymes.
Click Chemistry
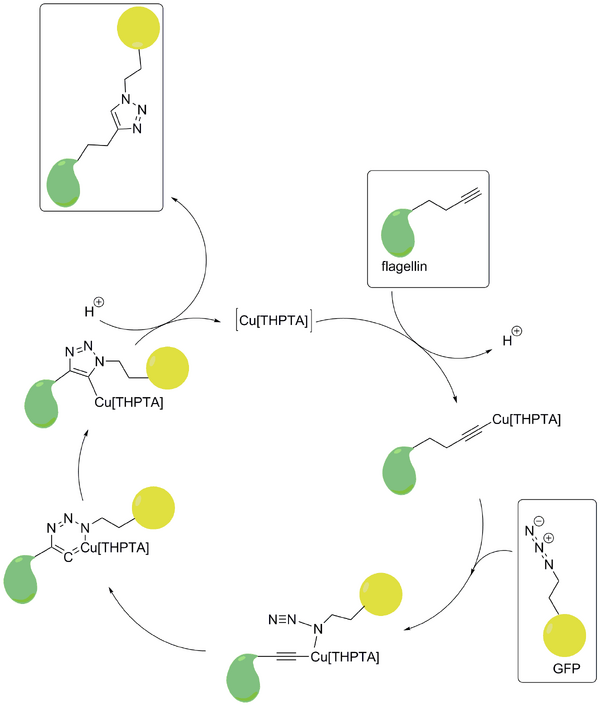
- The main reaction we focused on to link different structures to each other is called Click reaction. Our flagellins contain the L-Homopropargylglycine (HPG) and the structures that are supposed to be linked contain the counter part Azidohomoalanine (AHA). The binding of these compounds is achieved through a cycloaddition.
- Below you will find the reaction scheme of HPG and AHA.
Strategy
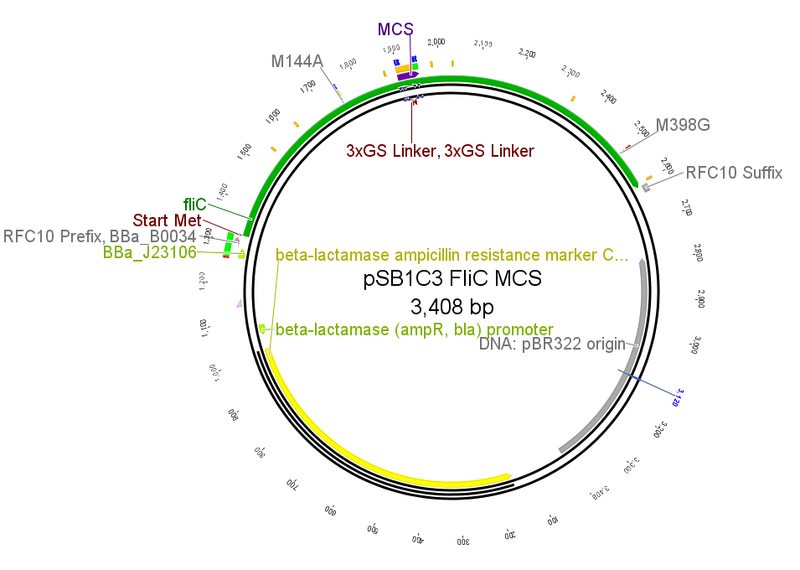
- We modified the FliC-gene by means of the Gibson assembly. We substituted the coding part for the D3-domain with a multiple cloning site (MCS). That way, we had a basic FliC-gene in the pSB1C3 vector, where different D3-domains could be cloned into.
- The several D3-domains were designed with gBLOCKS.
- The next step was cloning the D3-domains into the FliC-MCS-gene.
- After that, the vectors containing the successful FliC-D3-genes were transformed into the E. coli FliC-knockout strain MG1655 Z1. We tested the functionality resulting in the motility of the strain through motility essays. Therefore, we compared our transformants with the knockout strain (MG1655 Z1 ΔFliC) and the highly motile strain MG1655 Z1.
- After proving the functionality of the modified FliC-genes, the vector can be transformed into the methionine auxotrophic E. coli strain B834. The growth medium contains HPG which now will be incorporated instead of methionine. When the flagella have been completely synthesized, they can be harvested chemically or by shearing.
- For the click reaction the counter-part to HPG which is AHA has to be incorporated into the structure, that we want to link to our flagellins. We chose to work with GFP, first.
Results
- We got positive sequencing results for the construct pSB1C3_FliC_gB4 and pSB1C3_FliC_gB5.
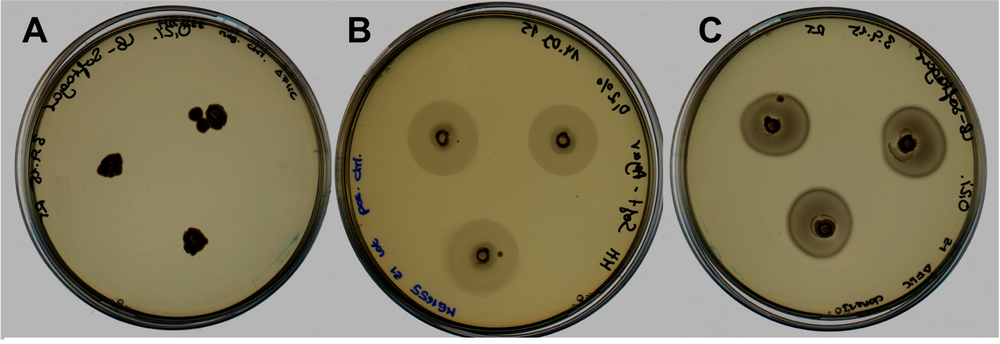
- The figure shows the comparison regarding the motility between our construct FliC_gB4 (right) and a negative (left) and positive (middle) control. Since our construct was transformed into E. coli srtain MG1655 z1 ΔFliC we used the highly motile wildtype strain MG1655 z1 as positive control and as negative control the FliC knockout strain MG1655 z1 ΔFliC.
- The pictures of negative control and construct were taken after 14 hours of incubation at 37 degrees Celsius. The positive control was taken after 5 hours for means of comparison since after 14 hours of incubation the plate of this control was completely overgrown and therefore not evaluable. Our motility assay reveals that our designed construct which was transformed into a FliC deficient strain is able to successfully express flagellins which are folded correctly and integrated into the surface of the cells. Compared to the highly motile wildtype strain the motility of our construct is less pronounced, nevertheless, in contrast to the negative control there is a significant difference in the motility visible. Our BioBrick enables the FliC deficient strain to be motile again.
- We also isolated the flagellas of the pSB1C3_FliC_gB4 construct mechanically and run a 15% SDS gel:
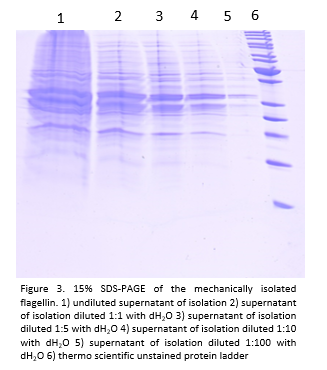
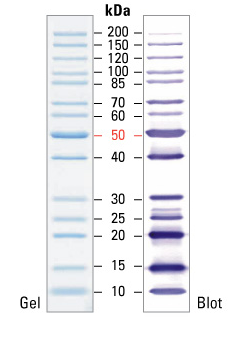
The main bands seem to be 34 kDa big, whereas the flagellins are supposed to be around 54 kDa big.
- Next, we cloned the FliC_gB4 and FliC_gB5 genes into the pET30b vector:
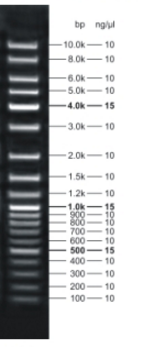
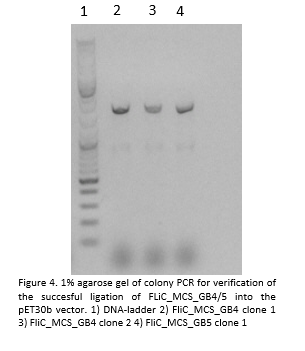
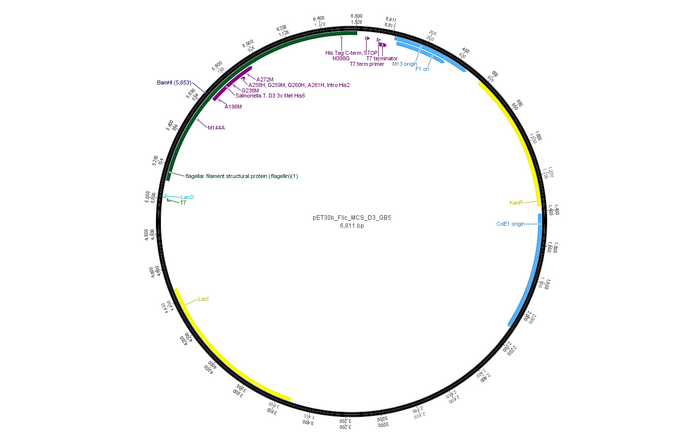
The insert seems to be 1.1 bp long which is approximately the lengths the constructs should have.
- These constructs, then, were transformed into the E. coli strain B834 and were overexpressed.
- Flagellins of the constructs pET30b_FliC_gB4 and pET30b_FliC_gB5 were Ni-NTA purified:
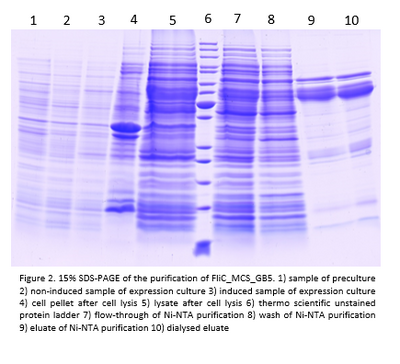
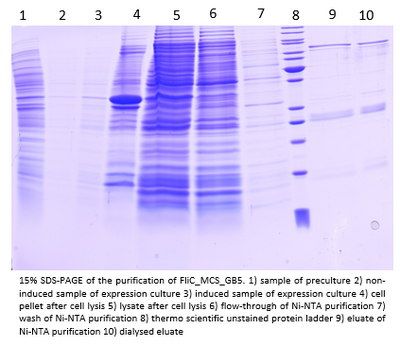
Also here the flagellins seem to be only about 33 kDa big.
- After purification the flagella a mass spectroscopy was performed:
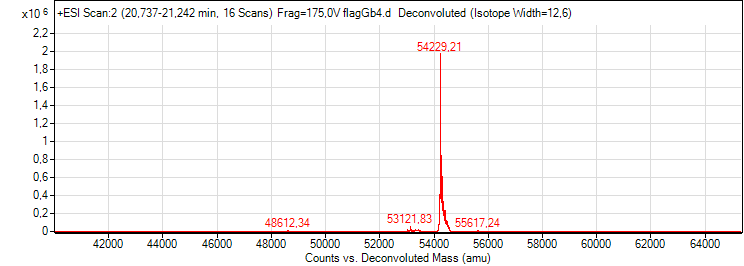
But the mass spectroscopy of the construct FliC_gB4 showed a size of about 54 kDa which is exactly the size of a flagellin.
- Finally, we performed the click reaction of the modified flagellins with GFP:
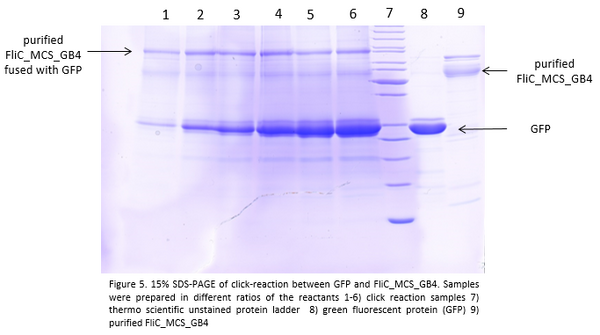
The GFP protein is about 27 kDa big, the pure flagellin is 54 kDa big. The fusion of both proteins through click chemistry should give us a construct that is about 81 kDa big which is exactly the case here. With this result, we can assume that our click reaction worked!
[1] Beatson SA, Minamino T, Pallen MJ. Variation in bacterial flagellins: from sequence to structure. Trends Microbiol. 2006 Apr;14(4):151-5. Epub 2006 Mar 15. PubMed PMID: 16540320